A Standardized Benchmark for Molecular Optimization
This is a Resource paper that introduces PMO (Practical Molecular Optimization), an open-source benchmark for evaluating molecular optimization algorithms with a focus on sample efficiency. The primary contribution is not a new algorithm but a comprehensive evaluation framework that exposes blind spots in how the field measures progress. By benchmarking 25 methods across 23 oracle functions under a fixed budget of 10,000 oracle calls, the authors provide a standardized protocol for transparent and reproducible comparison of molecular design methods.
The Missing Dimension: Oracle Budget in Molecular Design
Molecular optimization is central to drug and materials discovery, and the field has seen rapid growth in computational methods. Despite this progress, the authors identify three persistent problems with how methods are evaluated:
Lack of oracle budget control: Most papers do not report how many candidate molecules were evaluated by the oracle to achieve their results, despite this number spanning orders of magnitude. In practice, the most valuable oracles (wet-lab experiments, high-accuracy simulations) are expensive, making sample efficiency critical.
Trivial or self-designed oracles: Many papers only report on easy objectives like QED or penalized LogP, or introduce custom tasks that make cross-method comparison impossible.
Insufficient handling of randomness: Many algorithms are stochastic, yet existing benchmarks examined no more than five methods and rarely reported variance across independent runs.
Prior benchmarks such as GuacaMol, Therapeutics Data Commons (TDC), and Tripp et al.’s analysis each suffer from at least one of these issues. PMO addresses all three simultaneously.
The PMO Benchmark Design
The core innovation of PMO is its evaluation protocol rather than any single algorithmic contribution. The benchmark enforces three design principles:
Oracle budget constraint: All methods are limited to 10,000 oracle calls. This is deliberately much smaller than the unconstrained budgets typical in the literature, reflecting the practical reality that experimental evaluations are costly.
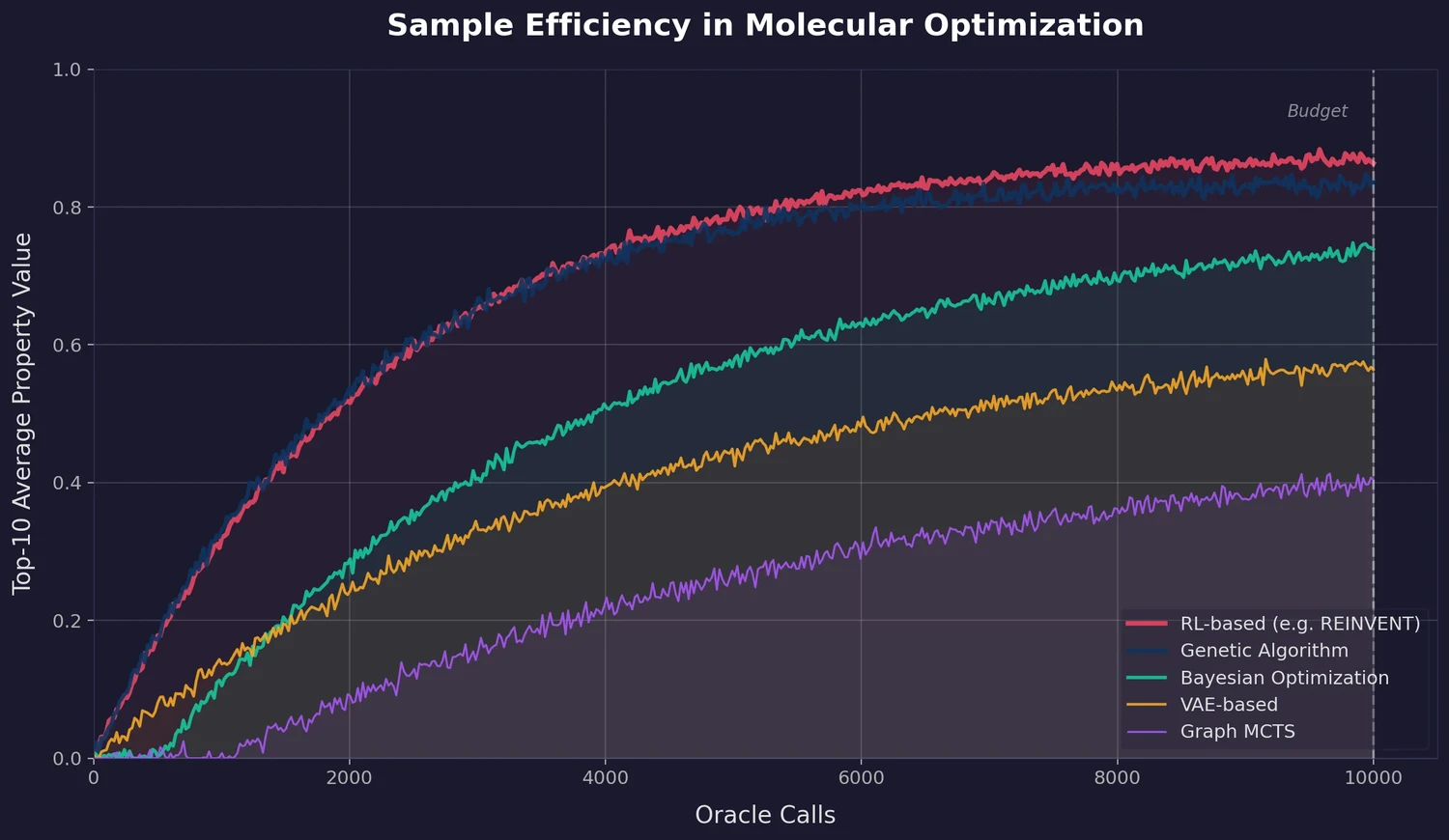
AUC-based metric: Instead of reporting only the final top-K score, PMO uses the area under the curve (AUC) of top-K average property value versus oracle calls:
$$ \text{AUC Top-}K = \int_{0}^{N} \bar{f}_{K}(n) , dn $$
where $\bar{f}_{K}(n)$ is the average property value of the top $K$ molecules found after $n$ oracle calls, and $N = 10{,}000$. The paper uses $K = 10$. This metric rewards methods that reach high property values quickly, not just those that eventually converge given enough budget. All AUC values are min-max scaled to [0, 1].
Standardized data: All methods use only the ZINC 250K dataset (approximately 250,000 molecules) whenever a database is required, ensuring a level playing field.
The benchmark includes 23 oracle functions: QED, DRD2, GSK3-beta, JNK3, and 19 oracles from GuacaMol covering multi-property objectives (MPOs) based on similarity, molecular weight, CLogP, and other pharmaceutically relevant criteria. All oracle scores are normalized to [0, 1].
25 Methods Across Nine Algorithm Families
The benchmark evaluates 25 molecular optimization methods organized along two dimensions: molecular assembly strategy (SMILES, SELFIES, atom-level graphs, fragment-level graphs, synthesis-based) and optimization algorithm (GA, MCTS, BO, VAE, GAN, score-based modeling, hill climbing, RL, gradient ascent). Each method was hyperparameter-tuned on two held-out tasks (zaleplon_mpo and perindopril_mpo) and then evaluated across all 23 oracles for 5 independent runs.
The following table summarizes the top 10 methods by sum of mean AUC Top-10 across all 23 tasks:
| Rank | Method | Assembly | Sum AUC Top-10 |
|---|---|---|---|
| 1 | REINVENT | SMILES | 14.196 |
| 2 | Graph GA | Fragments | 13.751 |
| 3 | SELFIES-REINVENT | SELFIES | 13.471 |
| 4 | GP BO | Fragments | 13.156 |
| 5 | STONED | SELFIES | 13.024 |
| 6 | LSTM HC | SMILES | 12.223 |
| 7 | SMILES GA | SMILES | 12.054 |
| 8 | SynNet | Synthesis | 11.498 |
| 9 | DoG-Gen | Synthesis | 11.456 |
| 10 | DST | Fragments | 10.989 |
The bottom five methods by overall ranking were GFlowNet-AL, Pasithea, JT-VAE, Graph MCTS, and MolDQN.
REINVENT is ranked first across all six metrics considered (AUC Top-1, AUC Top-10, AUC Top-100, Top-1, Top-10, Top-100). Graph GA is consistently second. Both methods were released several years before many of the methods they outperform, yet they are rarely used as baselines in newer work.
Key Findings: Older Methods Win and SELFIES Offers Limited Advantage
The benchmark yields several findings with practical implications:
No method solves optimization within realistic budgets. None of the 25 methods can optimize the included objectives within hundreds of oracle calls (the scale at which experimental evaluations would be feasible), except for trivially easy oracles like QED, DRD2, and osimertinib_mpo.
Older algorithms remain competitive. REINVENT (2017) and Graph GA (2019) outperform all newer methods tested, including those published at top AI conferences. The absence of standardized benchmarking had obscured this fact.
SMILES versus SELFIES. SELFIES was designed to guarantee syntactically valid molecular strings, but head-to-head comparisons show that SELFIES-based variants of language model methods (REINVENT, LSTM HC, VAE) generally do not outperform their SMILES counterparts. Modern language models learn SMILES grammar well enough that syntactic invalidity is no longer a practical issue. The one exception is genetic algorithms, where SELFIES-based GAs (STONED) outperform SMILES-based GAs, likely because SELFIES provides more intuitive mutation operations.
Model-based methods need careful design. Model-based variants (GP BO relative to Graph GA, GFlowNet-AL relative to GFlowNet) do not consistently outperform their model-free counterparts. GP BO outperformed Graph GA in 12 of 23 tasks but underperformed on sum, and GFlowNet-AL underperformed GFlowNet in nearly every task. The bottleneck is the quality of the predictive surrogate model, and naive surrogate integration can actually hurt performance.
Oracle landscape determines method suitability. Clustering analysis of relative AUC Top-10 scores reveals clear patterns. String-based GAs excel on isomer-type oracles (which are sums of atomic contributions), while RL-based and fragment-based methods perform better on similarity-based MPOs. This suggests there is no single best algorithm, and method selection should be informed by the optimization landscape.
Hyperparameter tuning and multiple runs are essential. Optimal hyperparameters differed substantially from default values in original papers. For example, REINVENT’s performance is highly sensitive to its sigma parameter, and the best value under the constrained-budget setting is much larger than originally suggested. Methods like Graph GA and GP BO also show high variance across runs, underscoring the importance of reporting distributional outcomes rather than single-run results.
Limitations
The authors acknowledge several limitations: they cannot exhaustively tune every hyperparameter or include every variant of each method; the conclusion may be biased toward similarity-based oracles (which dominate the 23 tasks); important quantities like synthesizability and diversity are not thoroughly evaluated; and oracle calls from pre-training data in model-based methods are counted against the budget, which may disadvantage methods that could leverage prior data collection. For a follow-up study that adds property filters and diversity requirements to the PMO evaluation, see Re-evaluating Sample Efficiency.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Molecule library | ZINC 250K | ~250,000 molecules | Used for screening, pre-training generative models, and fragment extraction |
| Oracle functions | TDC / GuacaMol | 23 tasks | All scores normalized to [0, 1] |
Algorithms
25 molecular optimization methods spanning 9 algorithm families and 5 molecular assembly strategies. Each method was hyperparameter-tuned on 2 held-out tasks (zaleplon_mpo, perindopril_mpo) using 3 independent runs, then evaluated on all 23 tasks with 5 independent runs each.
Evaluation
| Metric | Description | Notes |
|---|---|---|
| AUC Top-K | Area under curve of top-K average vs. oracle calls | Primary metric; K=10; min-max scaled to [0, 1] |
| Top-K | Final top-K average property value at 10K calls | Secondary metric |
| Sum rank | Sum of AUC Top-10 across all 23 tasks | Used for overall ranking |
Hardware
The paper states hardware details are in Appendix C.2. The benchmark runs on standard compute infrastructure and does not require GPUs for most methods. Specific compute requirements vary by method.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| mol_opt | Code | MIT | Full benchmark implementation with all 25 methods |
| Benchmark results | Dataset | Unknown | All experimental results from the paper |
| TDC | Dataset | MIT | Oracle functions and evaluation infrastructure |
Citation
@inproceedings{gao2022sample,
title={Sample Efficiency Matters: A Benchmark for Practical Molecular Optimization},
author={Gao, Wenhao and Fu, Tianfan and Sun, Jimeng and Coley, Connor W.},
booktitle={Advances in Neural Information Processing Systems},
volume={35},
pages={21342--21357},
year={2022}
}
Paper Information
Citation: Gao, W., Fu, T., Sun, J., & Coley, C. W. (2022). Sample Efficiency Matters: A Benchmark for Practical Molecular Optimization. Advances in Neural Information Processing Systems, 35, 21342-21357. https://arxiv.org/abs/2206.12411
Publication: NeurIPS 2022
Additional Resources: