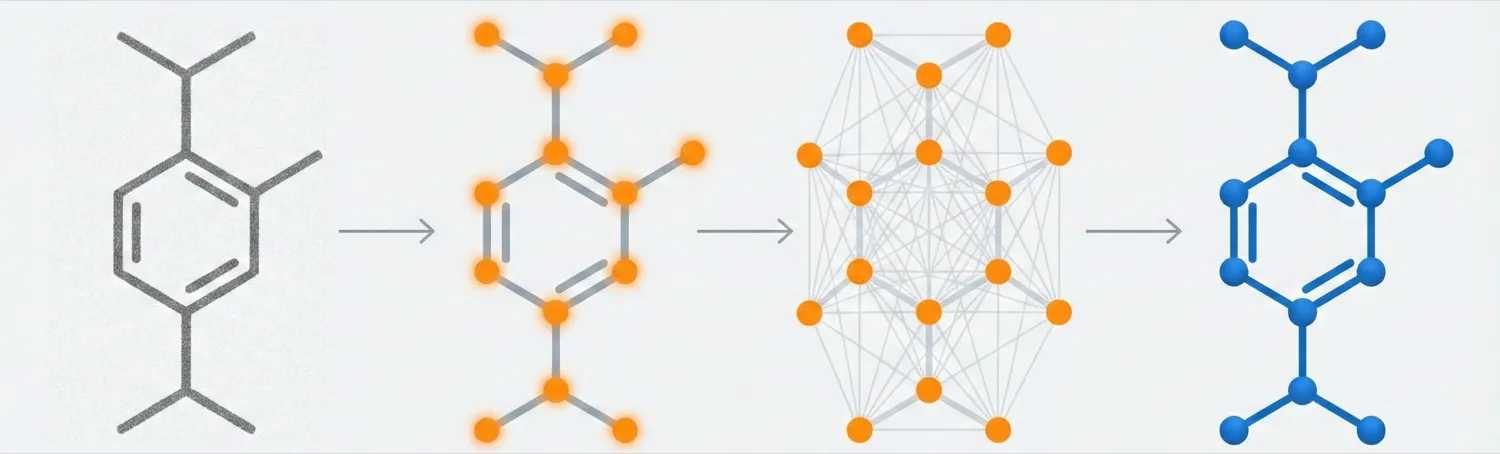
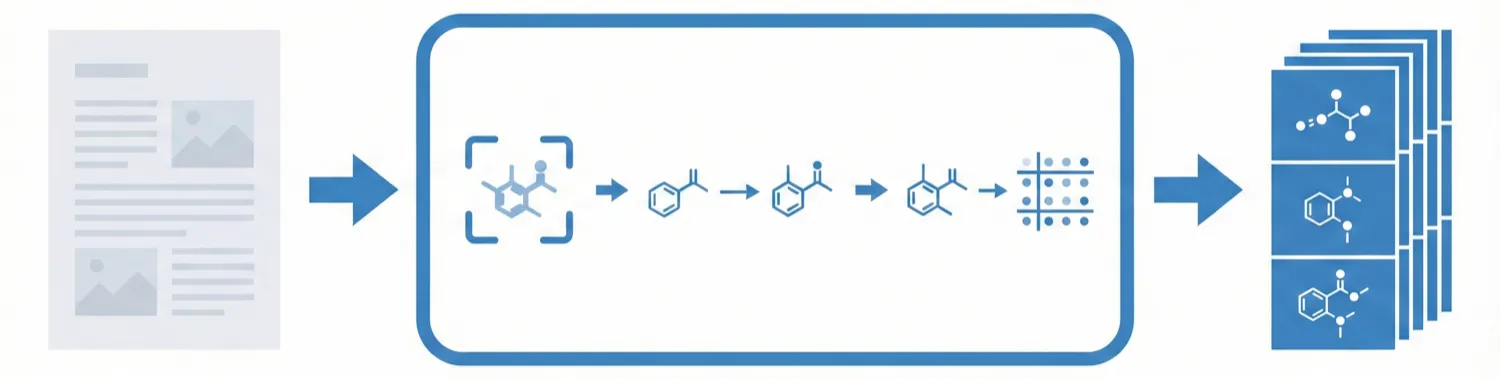
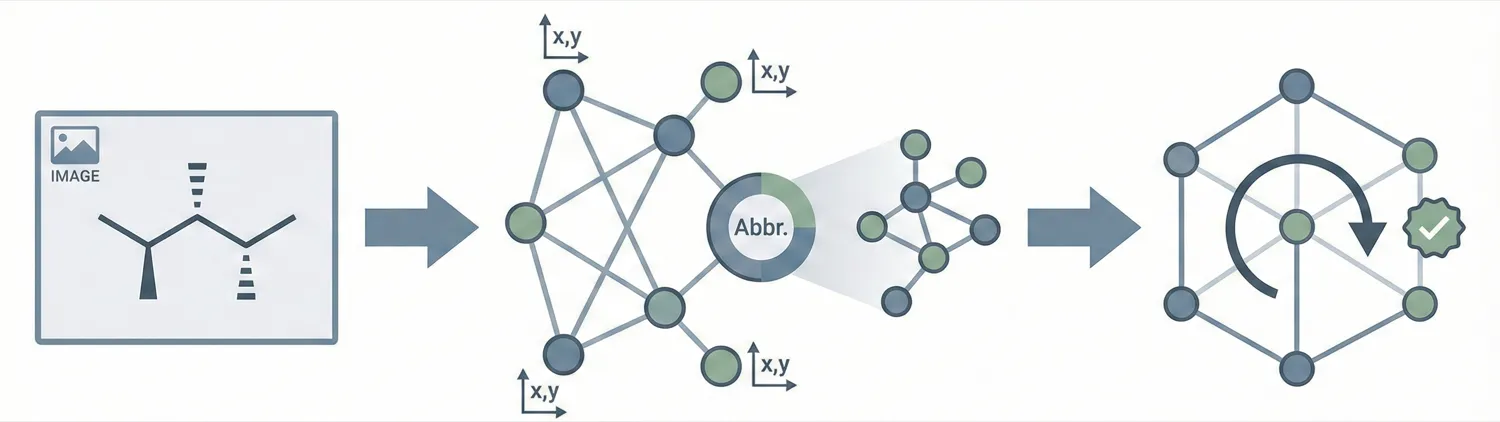
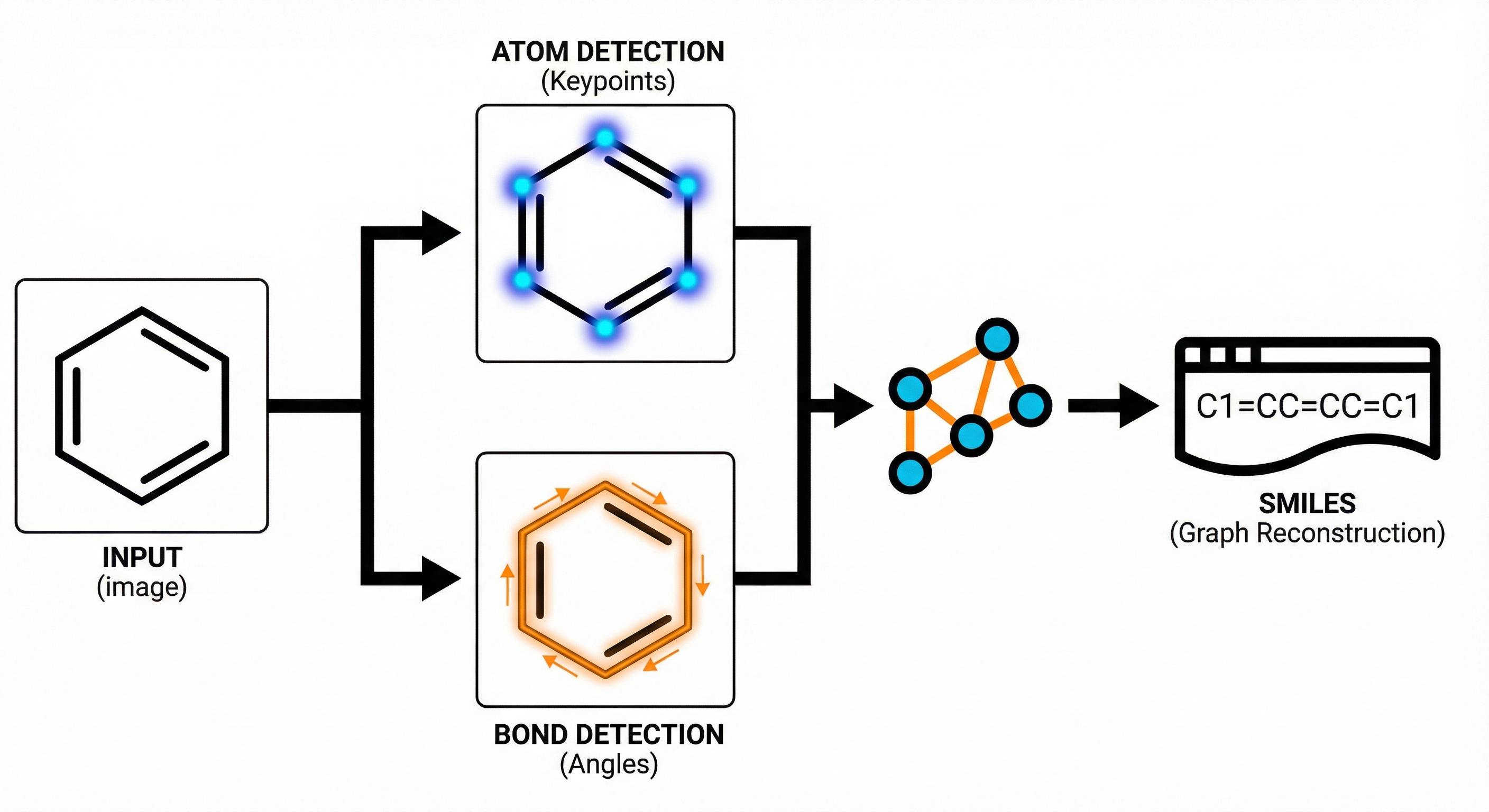
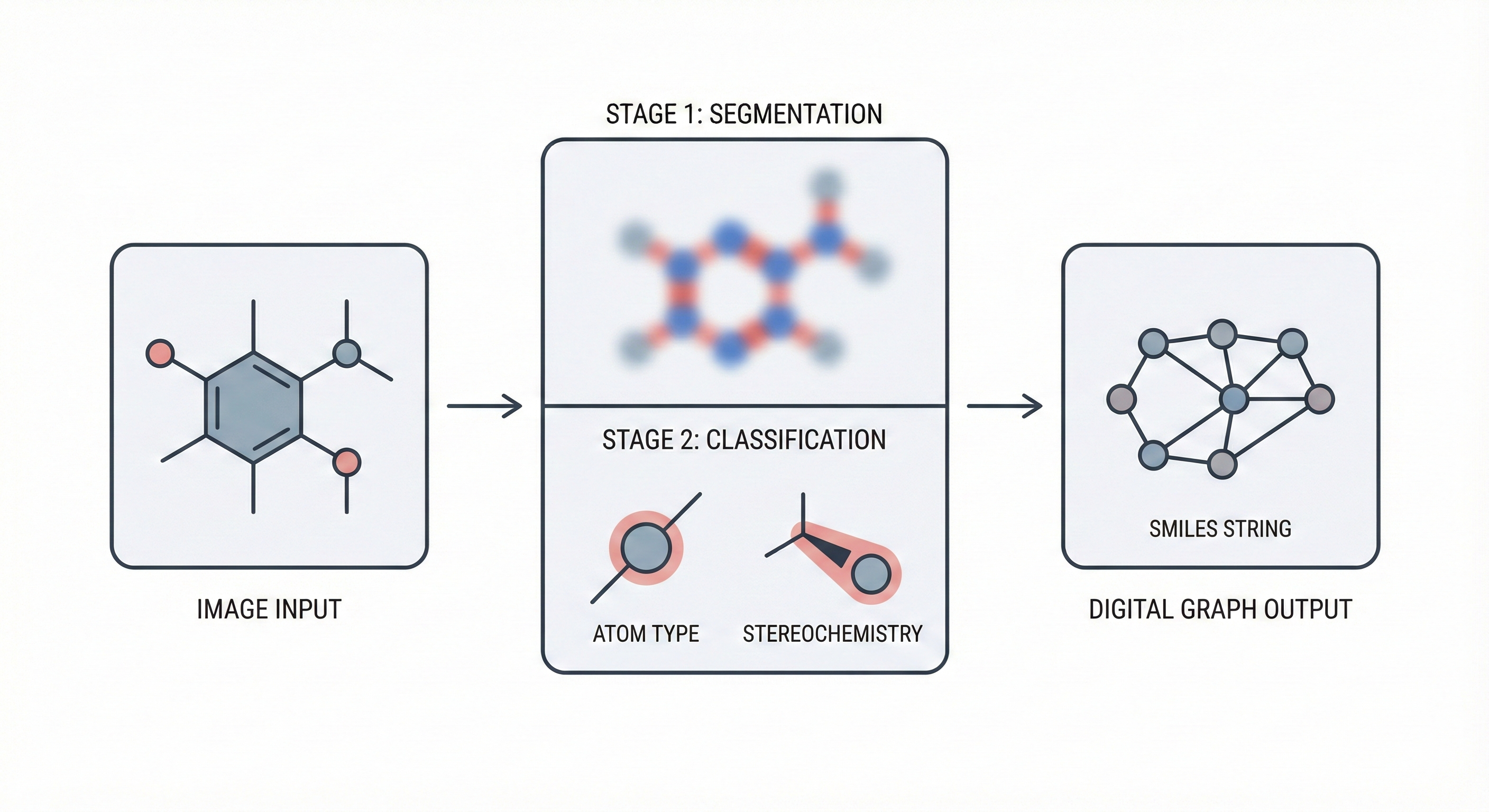
Image-to-graph methods bypass string representations entirely, predicting the molecular graph (atoms as nodes, bonds as edges) directly from the input image. This family includes segmentation-based approaches like ChemGrapher and Staker et al.’s U-Net pipeline, keypoint-detection architectures like ABC-Net, and joint atom-bond-coordinate predictors like MolScribe. By reasoning about spatial structure rather than linearizing it, these models tend to handle stereochemistry and abbreviated groups more naturally than sequence-based alternatives. Full-pipeline systems like MolMiner and MolMole extend the approach to page-level chemical extraction from documents.
| Year | Paper | Key Idea |
|---|---|---|
| 2020 | ChemGrapher: Deep Learning for Chemical Graph OCSR | Semantic segmentation and classification CNNs for chemical graphs |
| 2022 | ABC-Net: Keypoint-Based Molecular Image Recognition | Keypoint estimation to detect atom and bond centers |
| 2022 | Image-to-Graph Transformers for Chemical Structures | Direct image-to-graph conversion with abbreviated symbol support |
| 2022 | MolMiner: Deep Learning OCSR with YOLOv5 Detection | YOLOv5 and MobileNetV2 for document-level molecular extraction |
| 2023 | MolGrapher: Graph-based Chemical Structure Recognition | Graph-based deep learning outperforming image captioning methods |
| 2023 | MolScribe: Robust Image-to-Graph Molecular Recognition | Joint prediction of atoms, bonds, and coordinates for OCSR |
| 2025 | MolMole: Unified Vision Pipeline for Molecule Mining | Unified framework for detection, reaction parsing, and OCSR |
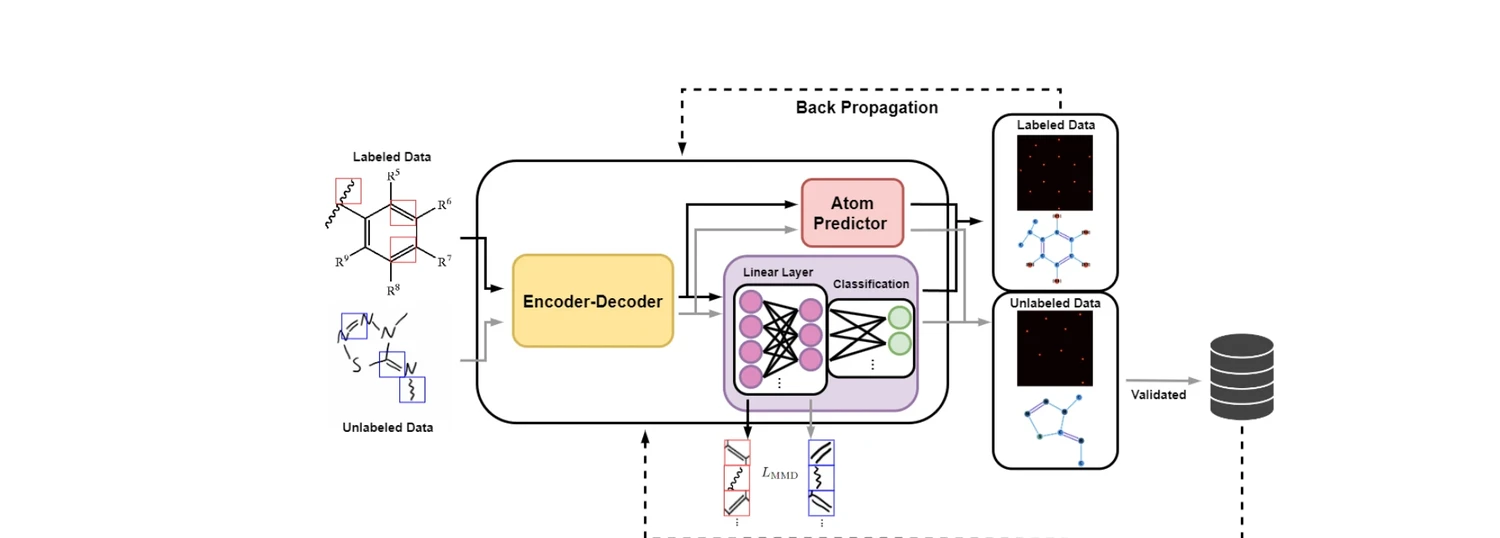
| 2026 | AdaptMol: Domain Adaptation for Molecular OCSR (2026) | MMD-based domain adaptation and self-training for hand-drawn OCSR |
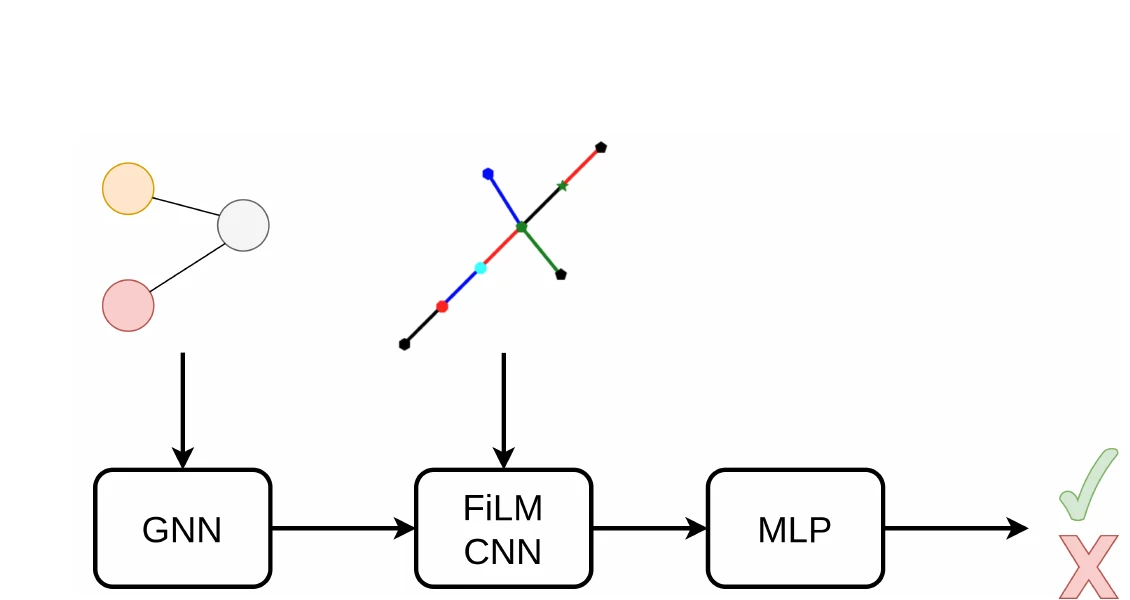
| 2026 | GraSP: Graph Recognition via Subgraph Prediction (2026) | Sequential subgraph prediction framework for image-to-graph OCSR |