Research notes on machine learning approaches to molecular simulation, including neural network potentials, equivariant message-passing architectures, structural representations, and all-atom foundation models.
| Year | Paper | Key Idea |
|---|---|---|
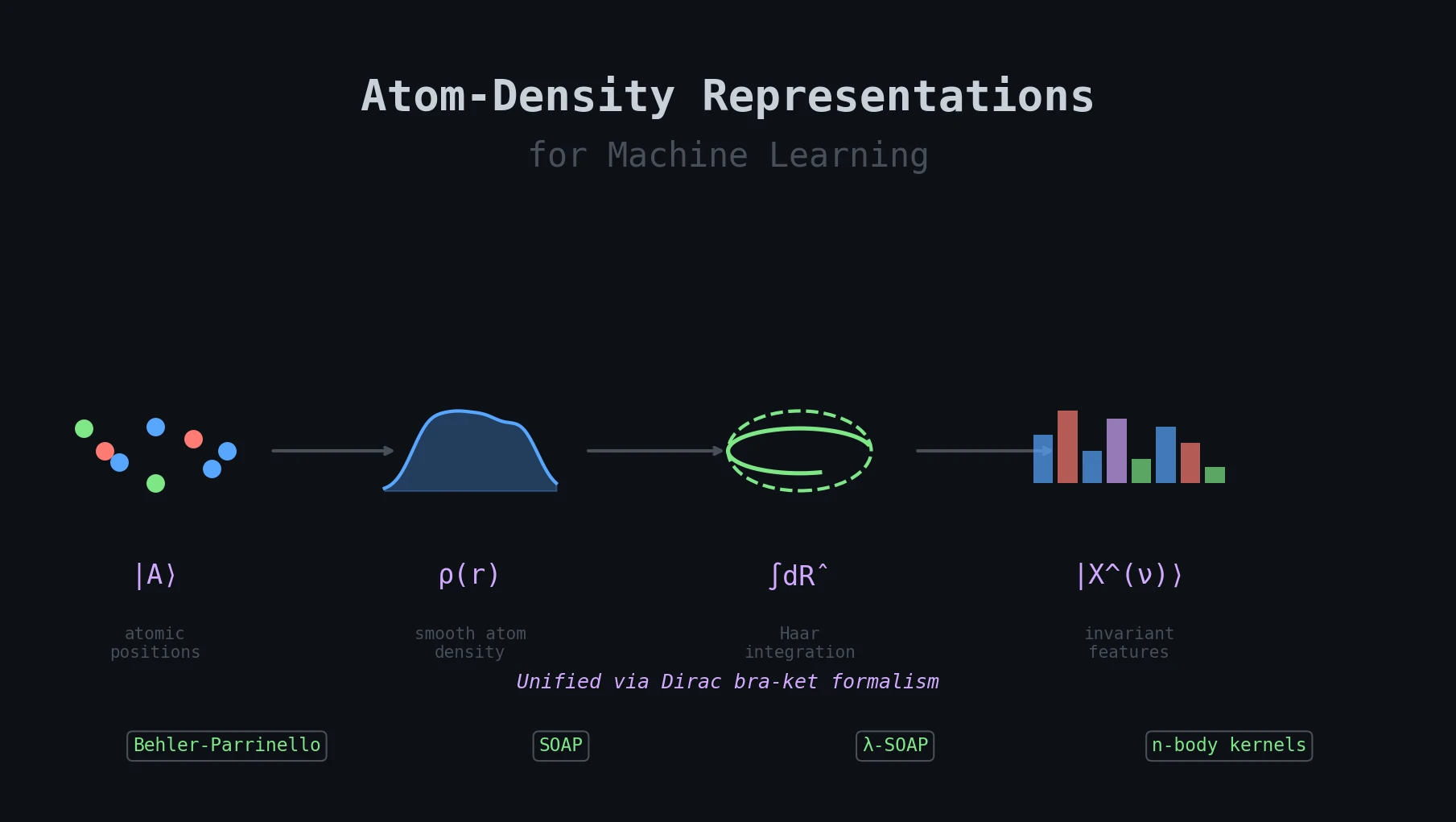
| 2019 | Atom-Density Representations for Machine Learning | Unified bra-ket framework connecting SOAP, Behler-Parrinello, and density-based representations |
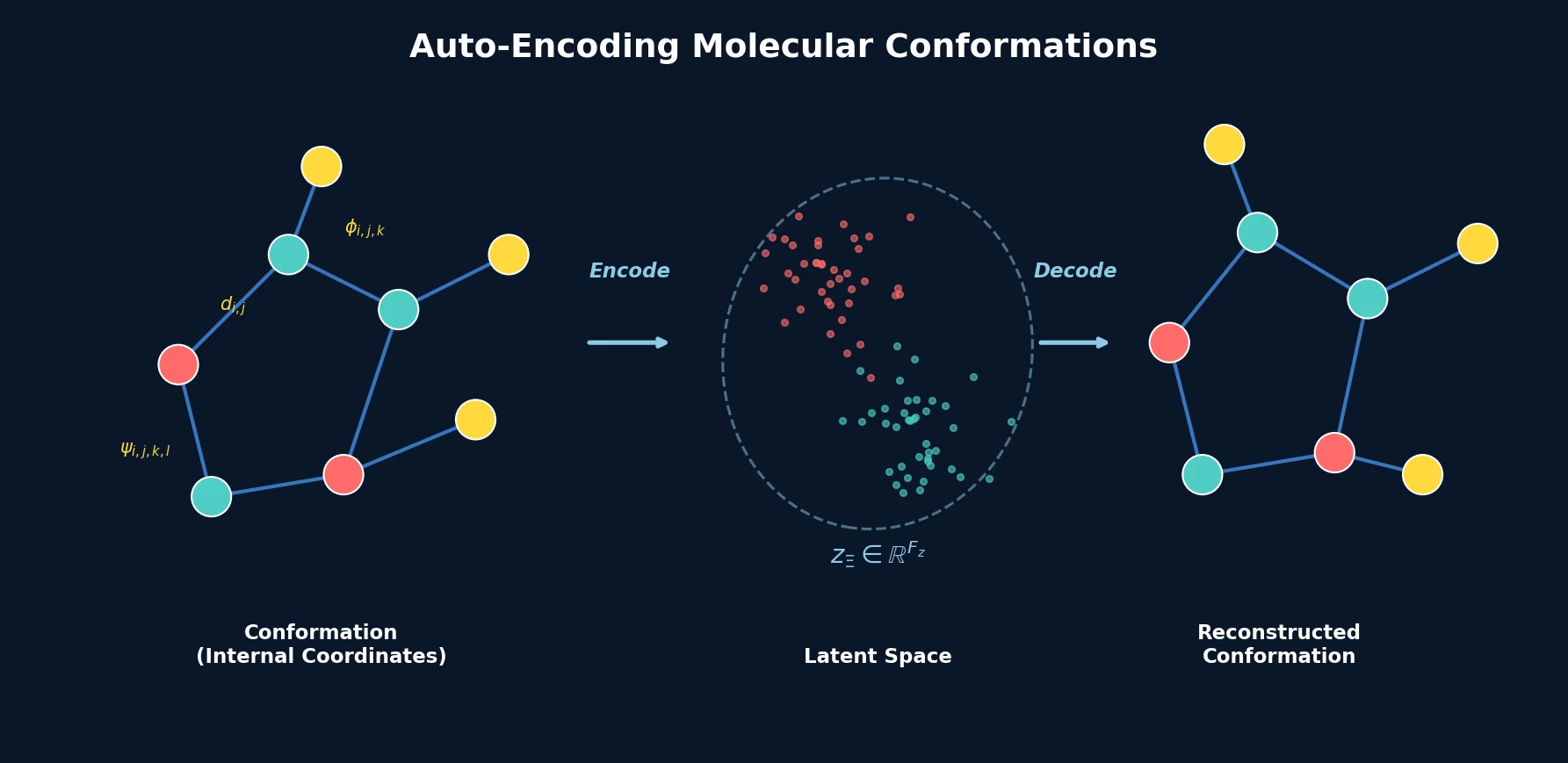
| 2020 | Conformation Autoencoder for 3D Molecules | Autoencoder mapping 3D conformations to fixed-size latent space via internal coordinates |
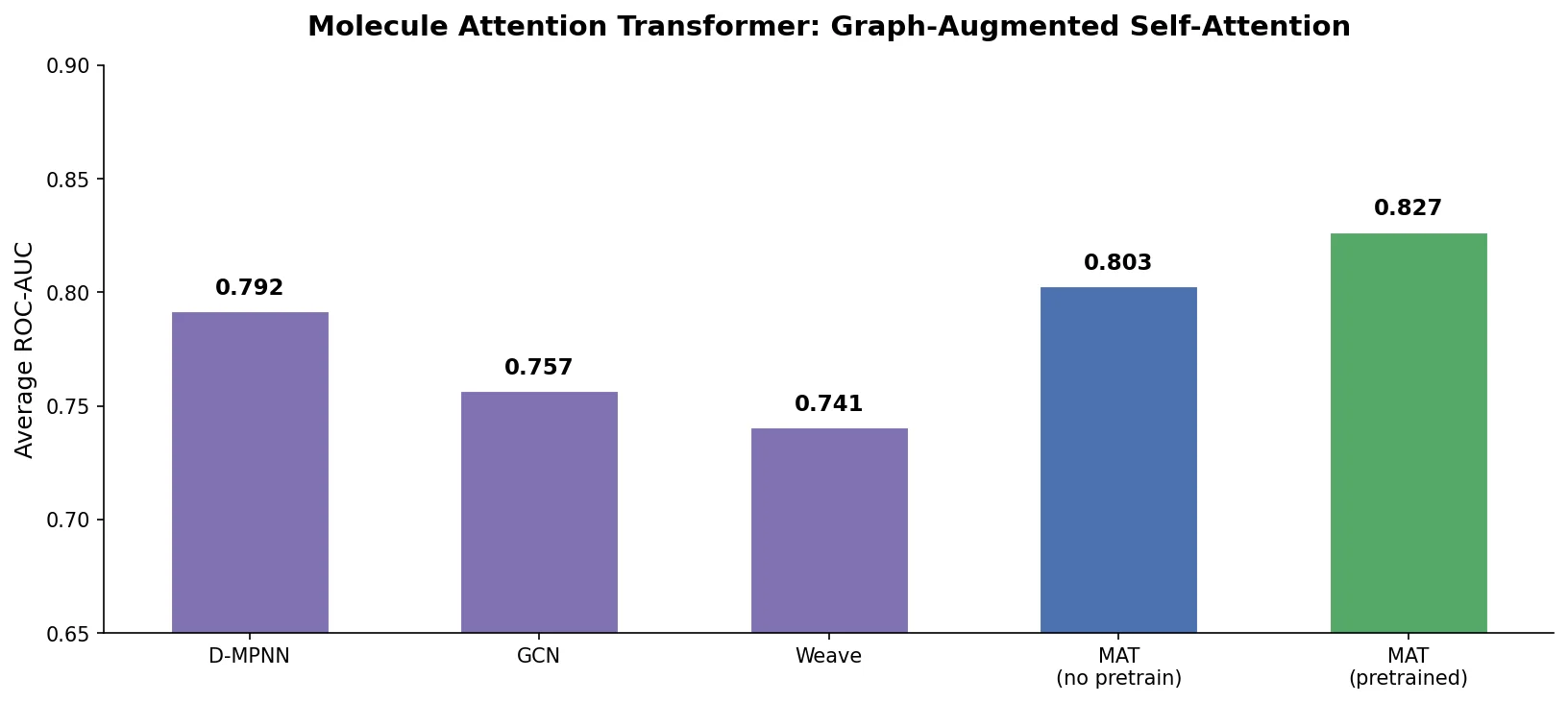
| 2020 | MAT: Graph-Augmented Transformer for Molecules (2020) | Transformer with inter-atomic distances and graph adjacency |
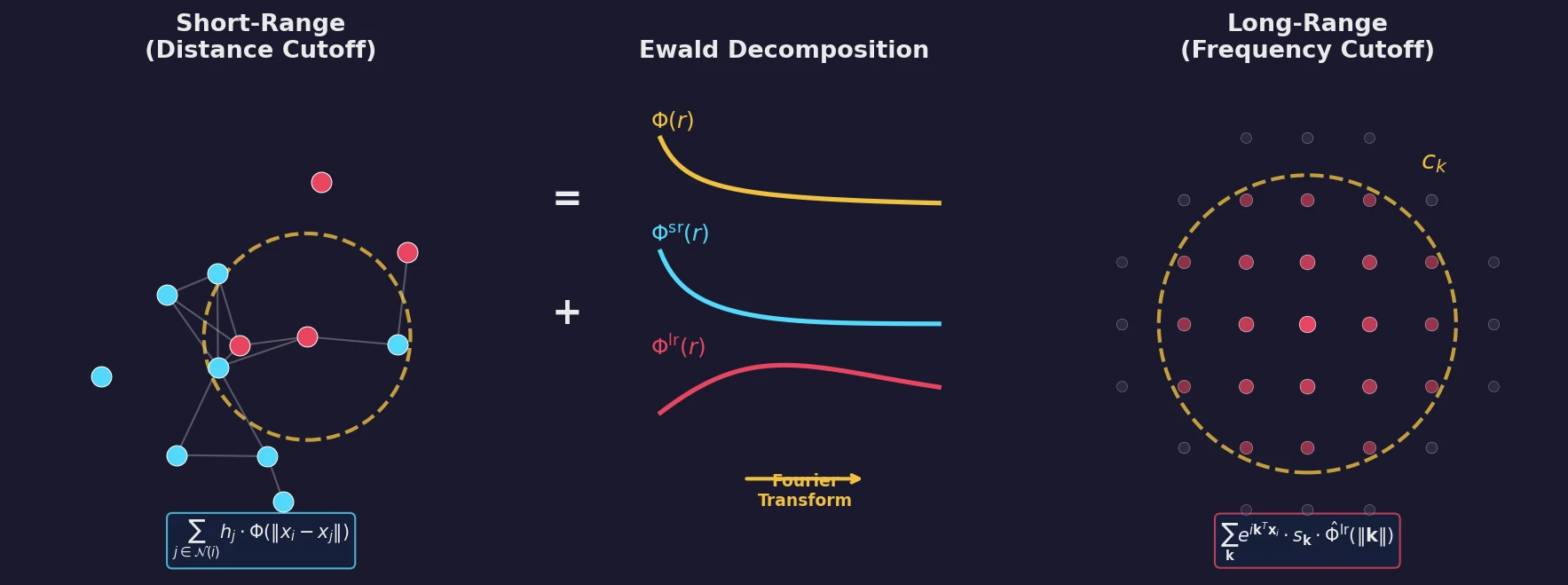
| 2023 | Ewald Message Passing for Molecular Graphs | Fourier-space long-range interactions improve GNN energy predictions |
| 2025 | Beyond Atoms: 3D Space Modeling for Molecular Pretraining | SpaceFormer models entire 3D molecular space for representations |
| 2025 | Dark Side of Forces: Non-Conservative ML Force Models | Critique of non-conservative forces in ML potentials |
| 2025 | DenoiseVAE: Adaptive Noise for Molecular Pre-training | Adaptive atom-specific noise distributions for better force fields |
| 2025 | Efficient DFT Hamiltonian Prediction via Adaptive Sparsity | SPHNet achieves up to 7x speedup in Hamiltonian prediction |
| 2025 | eSEN: Smooth Interatomic Potentials (ICML Spotlight) | Energy conservation as key MLIP diagnostic, introducing eSEN |
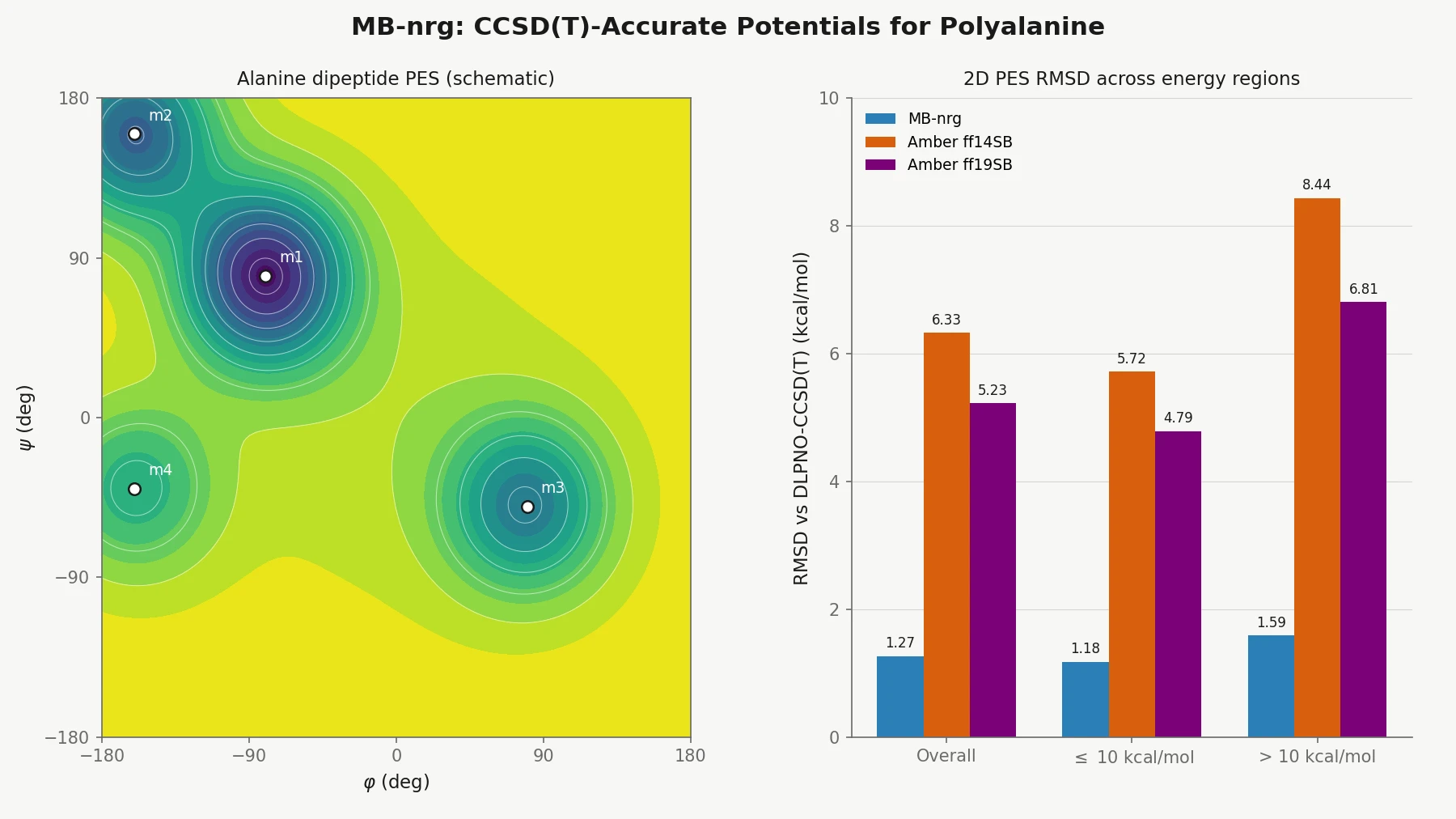
| 2025 | MB-nrg: CCSD(T)-Accurate Potentials for Polyalanine | Functional-group n-mer PIPs trained on DLPNO-CCSD(T) for gas-phase polyalanine |
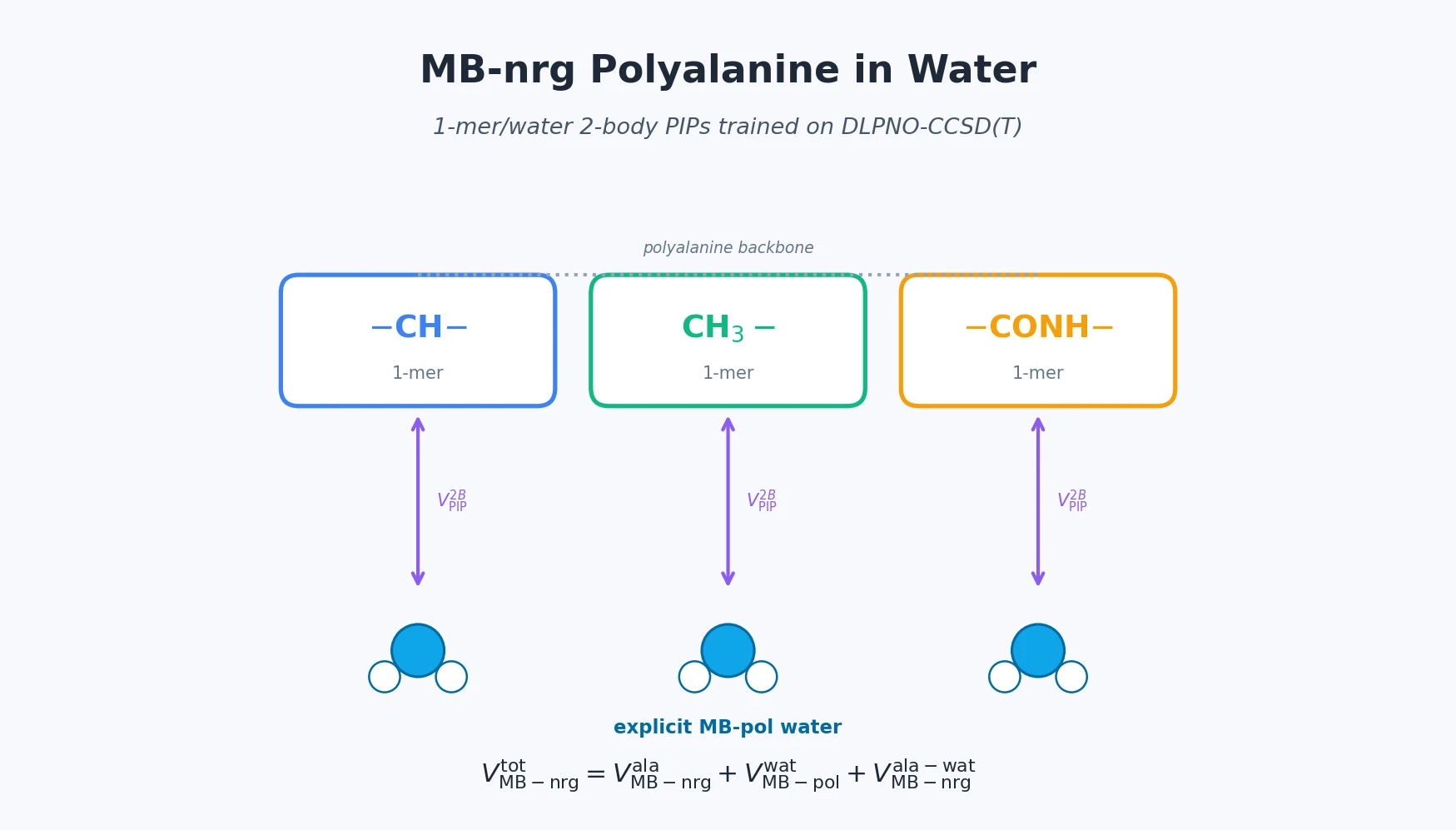
| 2025 | MB-nrg in Solution: Polyalanine in Water with CCSD(T) PEFs | 1-mer/water 2-body PIPs extend MB-nrg to alanine dipeptide solvation |
| 2025 | MOFFlow: Flow Matching for MOF Structure Prediction | Riemannian flow matching for Metal-Organic Framework generation |
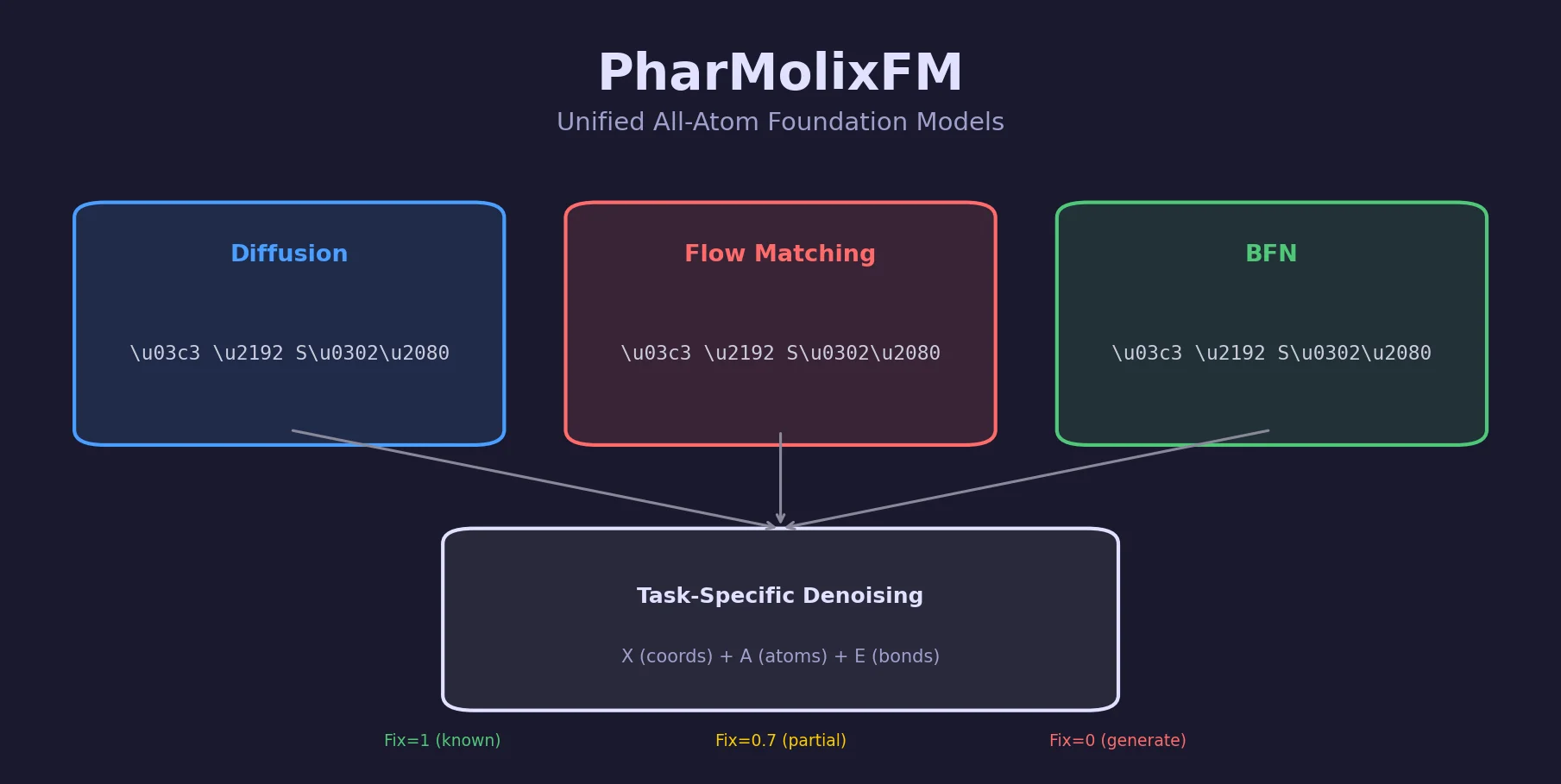
| 2025 | PharMolixFM: Multi-Modal All-Atom Molecular Models | Unified diffusion, flow matching, and BFN for molecular modeling |