Overview
SMILES (Simplified Molecular Input Line Entry System), originally developed by David Weininger in the late 1980s, is a one-dimensional string format for representing chemical molecular structures. It linearizes 3D molecular structures by performing a depth-first traversal of the molecular graph, recording the atoms and bonds along the way.
For example, the simple molecule ethanol ($\text{C}_2\text{H}_6\text{O}$) can be represented as CCO, while the more complex caffeine molecule becomes CN1C=NC2=C1C(=O)N(C(=O)N2C)C.
Key Characteristics
- Human-readable: Designed primarily for human readability. Compare with InChI, a hierarchical representation optimized for machine parsing.
- Compact: More compact than other representations (3D coordinates, connectivity tables)
- Simple syntax: A language with simple syntax and structure, making it relatively easy to learn and use for chemists and researchers
- Flexible: Both linear and cyclic structures can be represented in many different valid ways
For a hands-on tutorial on visualizing SMILES strings as 2D molecular images, see Converting SMILES Strings to 2D Molecular Images.
Basic Syntax
Atomic Symbols
SMILES uses standard atomic symbols with implied hydrogen atoms:
C(methane, $\text{CH}_4$)N(ammonia, $\text{NH}_3$)O(water, $\text{H}_2\text{O}$)P(phosphine, $\text{PH}_3$)S(hydrogen sulfide, $\text{H}_2\text{S}$)Cl(hydrogen chloride, $\text{HCl}$)
Bracket notation: Elements outside the organic subset must be shown in brackets, e.g., [Pt] for elemental platinum. The organic subset (B, C, N, O, P, S, F, Cl, Br, and I) can omit brackets.
Bond Representation
Bonds are represented by symbols:
- Single bond:
-(usually omitted)

CC- Double bond:
=
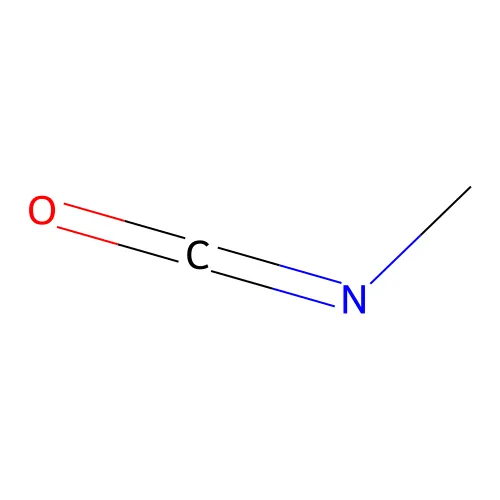
CN=C=O- Triple bond:
#
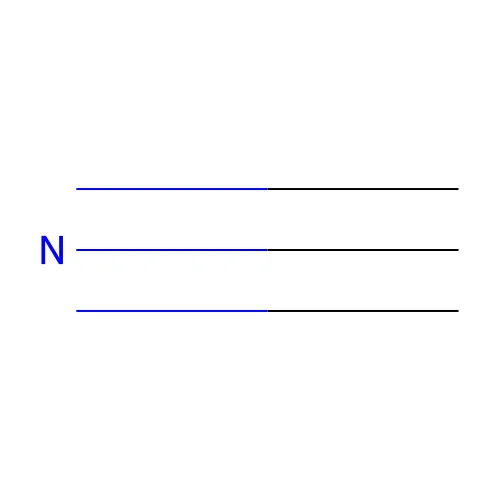
C#N- Aromatic bond:
:(usually omitted when lowercase atom symbols indicate aromaticity)
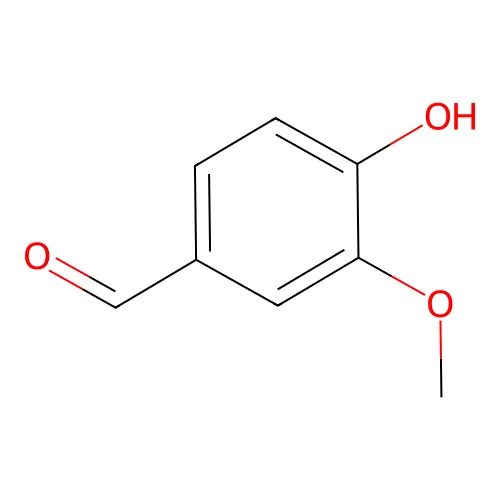
O=Cc1ccc(O)c(OC)c1- Disconnected structures:
.(separates disconnected components such as salts and ionic compounds)
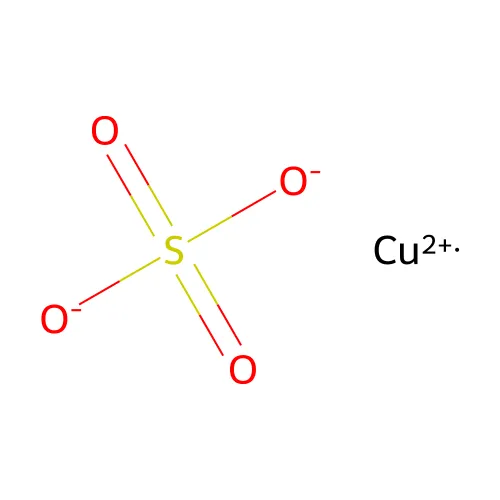
[Cu+2].[O-]S(=O)(=O)[O-]Structural Features
- Branches: Enclosed in parentheses and can be nested. For example,
CC(C)C(=O)Orepresents isobutyric acid, where(C)and(=O)are branches off the main chain.
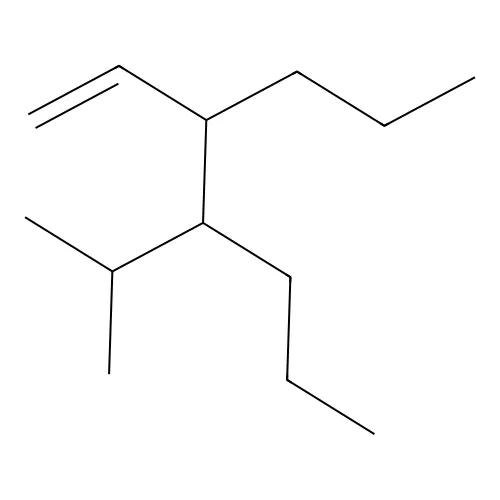
C=CC(CCC)C(C(C)C)CCC- Cyclic structures: Written by breaking bonds and using numbers to indicate bond connections. For example,
C1CCCCC1represents cyclohexane (the1connects the first and last carbon). - Aromaticity: Lower case letters are used for atoms in aromatic rings. For example, benzene is written as
c1ccccc1. - Formal charges: Indicated by placing the charge in brackets after the atom symbol, e.g.,
[C+],[C-], or[C-2]
Stereochemistry and Isomers
Isotope Notation
Isotope notation specifies the exact isotope of an element and comes before the element within square brackets, e.g., [13C] for carbon-13.
Double Bond Stereochemistry
Directional bonds can be specified using \ and / symbols to indicate the stereochemistry of double bonds:
C/C=C\Crepresents (E)-2-butene (trans configuration)C/C=C/Crepresents (Z)-2-butene (cis configuration)
The direction of the slashes indicates which side of the double bond each substituent is on.
Tetrahedral Chirality
Chirality around tetrahedral centers uses @ and @@ symbols:
N[C@](C)(F)C(=O)OvsN[C@@](F)(C)C(=O)O- Anti-clockwise counting vs clockwise counting
@and@@are shorthand for@TH1and@TH2, respectively
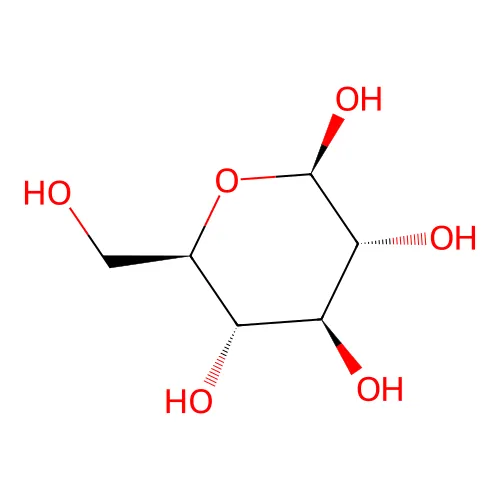
OC[C@@H](O1)[C@@H](O)[C@H](O)[C@@H](O)[C@H](O)1Advanced Stereochemistry
More general notation for other stereocenters:
@AL1,@AL2for allene-type stereocenters@SP1,@SP2,@SP3for square-planar stereocenters@TB1…@TB20for trigonal bipyramidal stereocenters@OH1…@OH30for octahedral stereocenters
SMILES allows partial specification since it relies on local chirality.
SMILES in Machine Learning
Beyond its original role as a compact notation, SMILES has become the dominant molecular input format for deep learning in chemistry. Its adoption has revealed both strengths and challenges specific to neural architectures.
Canonical vs. Randomized SMILES
Canonical SMILES algorithms produce a single unique string per molecule, which is valuable for database deduplication. In generative modeling, however, canonical representations introduce training bias: the canonicalization algorithm constrains how the molecular graph is traversed (e.g., prioritizing sidechains over ring atoms), forcing models to learn both valid SMILES syntax and the specific ordering rules. Structurally similar molecules can have substantially different canonical strings, making complex topologies harder to sample.
Randomized SMILES address this by generating non-unique representations through random atom orderings. Training RNN-based generative models on randomized SMILES acts as data augmentation, improving chemical space coverage, sampling uniformity, and completeness compared to canonical SMILES (Arus-Pous et al., 2019). In one benchmark, randomized SMILES recovered significantly more of GDB-13 chemical space than canonical SMILES across all training set sizes.
RDKit makes it straightforward to enumerate randomized SMILES for a given molecule:
from rdkit import Chem
mol = Chem.MolFromSmiles("c1ccc(C(=O)O)cc1") # benzoic acid
# Canonical form (deterministic)
print(Chem.MolToSmiles(mol))
# -> O=C(O)c1ccccc1
# Randomized forms (different each call)
for _ in range(5):
print(Chem.MolToSmiles(mol, doRandom=True))
# -> OC(=O)c1ccccc1
# -> O=C(c1ccccc1)O
# -> OC(c1ccccc1)=O
# -> C(O)(c1ccccc1)=O
# -> c1c(C(=O)O)cccc1
Each of these strings encodes the same molecule but presents a different traversal of the molecular graph, giving a generative model more diverse training signal per molecule.
Validity and the Role of Invalid SMILES
A large fraction of SMILES strings generated by neural models are syntactically or semantically invalid. Early efforts aimed to eliminate invalid outputs entirely, either through constrained representations like SELFIES (which guarantee 100% validity) or modified syntax like DeepSMILES (which removes paired syntax; see Variants below for syntax details).
More recent work has complicated this picture. Skinnider (2024) demonstrated that invalid SMILES generation actually benefits chemical language models. Invalid strings tend to be low-likelihood samples from the model’s probability distribution. Filtering them out is equivalent to removing the model’s least confident predictions, acting as implicit quality control. Meanwhile, enforcing absolute validity (as SELFIES does) can introduce systematic structural biases that impair distribution learning. This reframes SMILES’ non-robustness as potentially advantageous in certain ML contexts.
Tokenization Challenges
Converting SMILES strings into token sequences for neural models is non-trivial. The two baseline approaches illustrate the problem using chloramphenicol (O=C(NC([C@@H](O)c1ccc([N+](=O)[O-])cc1)CO)C(Cl)Cl):
import re
smiles = "O=C(NC([C@@H](O)c1ccc([N+](=O)[O-])cc1)CO)C(Cl)Cl"
# Character-level: splits every character individually
char_tokens = list(smiles)
# ['O', '=', 'C', '(', 'N', 'C', '(', '[', 'C', '@', '@', 'H', ']',
# '(', 'O', ')', 'c', '1', 'c', 'c', 'c', '(', '[', 'N', '+', ']',
# '(', '=', 'O', ')', '[', 'O', '-', ']', ')', 'c', 'c', '1', ')',
# 'C', 'O', ')', 'C', '(', 'C', 'l', ')', 'C', 'l']
# -> 49 tokens
# Atom-level: regex groups brackets, two-char elements, and bond symbols
atom_pattern = (
r"(\[[^\]]+\]|Br?|Cl?|N|O|S|P|F|I|"
r"b|c|n|o|s|p|\(|\)|\.|=|#|-|\+|"
r"\\|\/|:|~|@|\?|>>?|\*|%[0-9]{2}|[0-9])"
)
atom_tokens = re.findall(atom_pattern, smiles)
# ['O', '=', 'C', '(', 'N', 'C', '(', '[C@@H]', '(', 'O', ')', 'c',
# '1', 'c', 'c', 'c', '(', '[N+]', '(', '=', 'O', ')', '[O-]', ')',
# 'c', 'c', '1', ')', 'C', 'O', ')', 'C', '(', 'Cl', ')', 'Cl']
# -> 36 tokens
Character-level tokenization splits Cl (chlorine) into C + l, making the chlorine indistinguishable from carbon. It also fragments [C@@H] (a chiral carbon) into six meaningless tokens: [, C, @, @, H, ]. Atom-level tokenization preserves these as single tokens but still produces long sequences (~40 tokens per molecule on average in ChEMBL).
Several chemistry-aware tokenizers go further:
- SMILES Pair Encoding (SPE) adapts byte pair encoding to learn high-frequency SMILES substrings from large chemical datasets, compressing average sequence length from ~40 to ~6 tokens while preserving chemically meaningful substructures.
- Atom Pair Encoding (APE) preserves atomic identity during subword merging, preventing chemically meaningless token splits.
- Atom-in-SMILES (AIS) encodes each atom’s local chemical environment into the token itself (e.g., distinguishing a carbonyl carbon from a methyl carbon), reducing token degeneration and improving translation accuracy.
- Smirk achieves full OpenSMILES coverage with only 165 tokens by decomposing bracketed atoms into glyphs.
SMILES-Based Foundation Models
SMILES serves as the primary input format for molecular encoder models, including SMILES-BERT, SMILES-Transformer, BARTSmiles, SMI-TED, and MolBERT. These models learn molecular representations from large SMILES corpora through pre-training objectives like masked language modeling.
A key open challenge is robustness to SMILES variants. The AMORE framework revealed that current chemical language models struggle to recognize chemically equivalent SMILES representations (such as hydrogen-explicit vs. implicit forms, or different atom orderings) as encoding the same molecule.
Molecular Generation
SMILES is the dominant representation for de novo molecular generation. The typical pipeline trains a language model on SMILES corpora, then steers sampling toward molecules with desired properties. Major architecture families include:
- Variational autoencoders: The Automatic Chemical Design VAE (Gomez-Bombarelli et al., 2018) encodes SMILES into a continuous latent space, enabling gradient-based optimization toward target properties.
- RL-tuned generators: REINVENT and its successors fine-tune a pre-trained SMILES language model using reinforcement learning, rewarding molecules that satisfy multi-objective scoring functions. DrugEx extends this with Pareto-based multi-objective optimization.
- Adversarial approaches: ORGAN and LatentGAN apply GAN-based training to SMILES generation, using domain-specific rewards alongside the discriminator signal.
The challenges of canonical vs. randomized SMILES and invalid outputs discussed above are particularly relevant in this generation context.
Property Prediction
SMILES strings serve as the primary input for quantitative structure-activity relationship (QSAR) models. SMILES2Vec learns fixed-length molecular embeddings directly from SMILES for property regression and classification. MaxSMI demonstrates that SMILES augmentation (training on multiple randomized SMILES per molecule) improves property prediction accuracy, connecting the data augmentation benefits observed in generative settings to discriminative tasks.
Optical Chemical Structure Recognition
SMILES is also the standard output format for optical chemical structure recognition (OCSR) systems, which extract molecular structures from images in scientific literature. Deep learning approaches like DECIMER and Image2SMILES frame this as an image-to-SMILES translation problem, using encoder-decoder architectures to generate SMILES strings directly from molecular diagrams. For a taxonomy of OCSR approaches, see the OCSR methods overview.
Limitations
Classical Limitations
- Non-uniqueness: Different SMILES strings can represent the same molecule (e.g., ethanol can be written as
CCOorOCC). Canonical SMILES algorithms address this by producing a single unique representation. - Non-robustness: SMILES strings can be written that do not correspond to any valid molecular structure.
- Strings that cannot represent a molecular structure.
- Strings that violate basic rules (more bonds than is physically possible).
- Information loss: If 3D structural information exists, a SMILES string cannot encode it.
Machine Learning Limitations
The challenges described above (canonical ordering bias motivating randomized SMILES, validity constraints motivating DeepSMILES and SELFIES, and tokenization ambiguity motivating chemistry-aware tokenizers) remain active areas of research. See the linked sections for details on each.
Variants and Standards
Canonical SMILES
For how canonical vs. randomized SMILES affects generative modeling, see Canonical vs. Randomized SMILES above.
Canonical SMILES algorithms produce a single unique string per molecule by assigning a deterministic rank to each atom and then traversing the molecular graph in that rank order. Most implementations build on the Morgan algorithm (extended connectivity): each atom starts with an initial invariant based on its properties (atomic number, degree, charge, hydrogen count), then iteratively updates its invariant by incorporating its neighbors’ invariants until the ranking stabilizes. The final atom ranks determine the traversal order, which determines the canonical string.
In practice, the Morgan algorithm alone does not fully resolve all ties. Implementations must also make choices about tie-breaking heuristics, aromaticity perception (Kekulé vs. aromatic form), and stereochemistry encoding. Because these choices differ across toolkits (RDKit, OpenBabel, Daylight, ChemAxon), the same molecule can produce different “canonical” SMILES depending on the software. A canonical SMILES is only guaranteed unique within a single implementation, not across implementations.
from rdkit import Chem
# RDKit's canonical SMILES for caffeine
mol = Chem.MolFromSmiles("CN1C=NC2=C1C(=O)N(C(=O)N2C)C")
print(Chem.MolToSmiles(mol))
# -> Cn1c(=O)c2c(ncn2C)n(C)c1=O
Isomeric SMILES
Isomeric SMILES incorporates isotopes and stereochemistry information, providing more detailed molecular representations than generic SMILES. Non-isomeric SMILES strip this information, collapsing stereoisomers and isotopologues into the same string:
from rdkit import Chem
# L-alanine (chiral center)
mol = Chem.MolFromSmiles("N[C@@H](C)C(=O)O")
print(Chem.MolToSmiles(mol, isomericSmiles=True))
# -> C[C@H](N)C(=O)O (preserves chirality)
print(Chem.MolToSmiles(mol, isomericSmiles=False))
# -> CC(N)C(=O)O (chirality lost)
# Deuterated water (isotope labels)
mol2 = Chem.MolFromSmiles("[2H]O[2H]")
print(Chem.MolToSmiles(mol2, isomericSmiles=True))
# -> [2H]O[2H] (preserves isotopes)
print(Chem.MolToSmiles(mol2, isomericSmiles=False))
# -> [H]O[H] (isotope info lost)
OpenSMILES vs. Proprietary
- Proprietary: The original SMILES specification was proprietary (Daylight Chemical Information Systems), which led to compatibility issues between different implementations.
- OpenSMILES: An open-source alternative standardization effort to address compatibility concerns and provide a freely available specification.
Extensions and Related Notations
DeepSMILES
DeepSMILES modifies two aspects of SMILES syntax that cause most invalid strings in generative models, while remaining interconvertible with standard SMILES without information loss.
Ring closures: Standard SMILES uses paired digits (c1ccccc1 for benzene). A model must remember which digits are “open” and close them correctly. DeepSMILES replaces this with a single ring-size indicator at the closing position: cccccc6 means “connect to the atom 6 positions back.”
Branches: Standard SMILES uses matched parentheses (C(OC)(SC)F). DeepSMILES uses a postfix notation with only closing parentheses, where consecutive ) symbols indicate how far to pop back on the atom stack: COC))SC))F.
SMILES: c1ccccc1 C(OC)(SC)F
DeepSMILES: cccccc6 COC))SC))F
↑ ↑
single digit = no opening parens,
ring size )) pops back to C
A single unpaired symbol cannot be “unmatched,” eliminating the two main sources of syntactically invalid strings from generative models.
Reaction SMILES
Reaction SMILES extends the notation to represent chemical reactions by separating reactants, reagents, and products with > symbols. The general format is reactants>reagents>products, where each group can contain multiple molecules separated by .:
CC(=O)O.CCO>>CC(=O)OCC.O
│ │ │ │
│ │ │ └─ water
│ │ └─ ethyl acetate
│ └─ ethanol
└─ acetic acid
(Fischer esterification: acetic acid + ethanol → ethyl acetate + water)
The Molecular Transformer treats this as a machine translation problem, translating reactant SMILES to product SMILES with a Transformer encoder-decoder architecture.
SMARTS and SMIRKS
SMARTS (SMILES Arbitrary Target Specification) is a pattern language built on SMILES syntax for substructure searching. It extends SMILES with query primitives like atom environments ([CX3] for a carbon with three connections) and logical operators, enabling precise structural pattern matching:
from rdkit import Chem
# SMARTS pattern for a carboxylic acid group: C(=O)OH
pattern = Chem.MolFromSmarts("[CX3](=O)[OX2H1]")
for name, smi in [("acetic acid", "CC(=O)O"),
("benzoic acid", "c1ccc(C(=O)O)cc1"),
("ethanol", "CCO"),
("acetone", "CC(=O)C")]:
mol = Chem.MolFromSmiles(smi)
print(f" {name:15s} -> {'match' if mol.HasSubstructMatch(pattern) else 'no match'}")
# -> acetic acid -> match
# -> benzoic acid -> match
# -> ethanol -> no match
# -> acetone -> no match
SMIRKS extends SMARTS to describe reaction transforms, using atom maps (:1, :2, …) to track which atoms in the reactants correspond to which atoms in the products:
from rdkit.Chem import AllChem, MolFromSmiles, MolToSmiles
# SMIRKS for ester hydrolysis: break the C-O ester bond
smirks = "[C:1](=[O:2])[O:3][C:4]>>[C:1](=[O:2])[OH:3].[C:4][OH]"
rxn = AllChem.ReactionFromSmarts(smirks)
reactant = MolFromSmiles("CC(=O)OCC") # ethyl acetate
products = rxn.RunReactants((reactant,))
print(" + ".join(MolToSmiles(p) for p in products[0]))
# -> CC(=O)O + CCO (acetic acid + ethanol)
See the Smirk tokenizer for a recent approach to tokenizing these extensions for molecular foundation models.
t-SMILES
t-SMILES encodes molecules as fragment-based strings by decomposing a molecule into chemically meaningful substructures, arranging them into a full binary tree, and traversing it breadth-first. This dramatically reduces nesting depth compared to standard SMILES (99.3% of tokens at depth 0-2 vs. 68.0% for SMILES on ChEMBL).
Standard SMILES (depth-first, atom-level):
CC(=O)Oc1ccccc1C(=O)O (aspirin)
t-SMILES pipeline:
1. Fragment: [CC(=O)O*] [*c1ccccc1*] [*C(=O)O]
2. Binary tree:
[*c1ccccc1*]
/ \
[CC(=O)O*] [*C(=O)O]
3. BFS string: [*c1ccccc1*] ^ [CC(=O)O*] ^ [*C(=O)O]
The framework introduces two symbols beyond standard SMILES: ^ separates adjacent fragments (analogous to spaces between words), and & marks empty tree nodes. Only single closure symbols are needed per fragment, eliminating the deep nesting that makes standard SMILES difficult for generative models on small datasets.
Further Reading
For a more robust alternative that guarantees 100% valid molecules, see SELFIES (Self-Referencing Embedded Strings). For the historical context and design philosophy behind SMILES, see SMILES: The Original Paper (Weininger 1988).