String notations for encoding molecular structure, from the original SMILES format through SELFIES and InChI, plus tokenization methods and representation surveys.
Concept References
| Paper | Key Idea |
|---|---|
| SMILES: A Compact Notation for Chemical Structures | Comprehensive overview of SMILES ASCII string notation |
| SELFIES: A Robust Molecular String Representation | Overview of SELFIES, where every string decodes to a valid molecule |
| InChI: The International Chemical Identifier | IUPAC’s layered chemical identifier for database interoperability |
SMILES Family
| Year | Paper | Key Idea |
|---|---|---|
| 1988 | SMILES Notation: The Original Paper by Weininger (1988) | Introduced SMILES encoding rules for molecular strings |
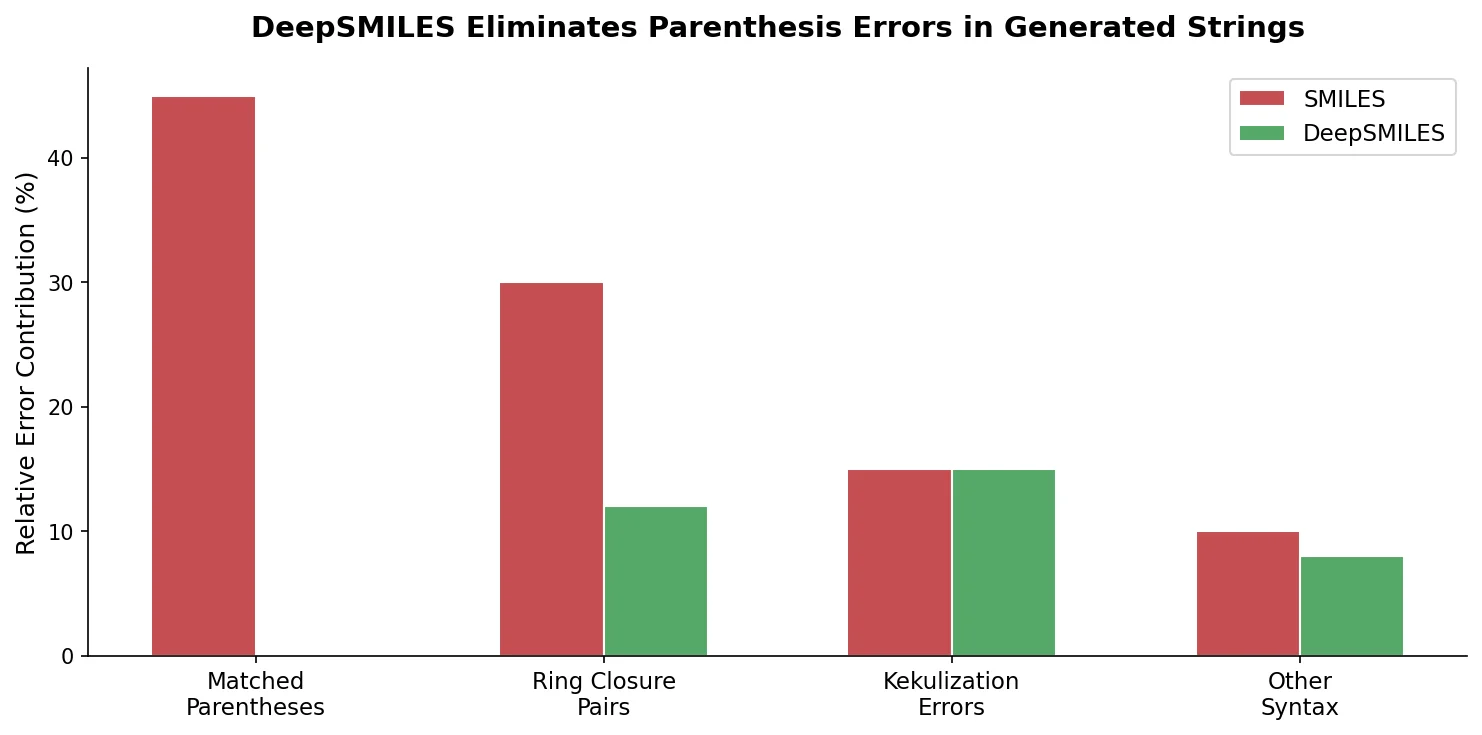
| 2018 | DeepSMILES: Adapting SMILES Syntax for Machine Learning | Postfix notation eliminating unbalanced parentheses and ring closures |
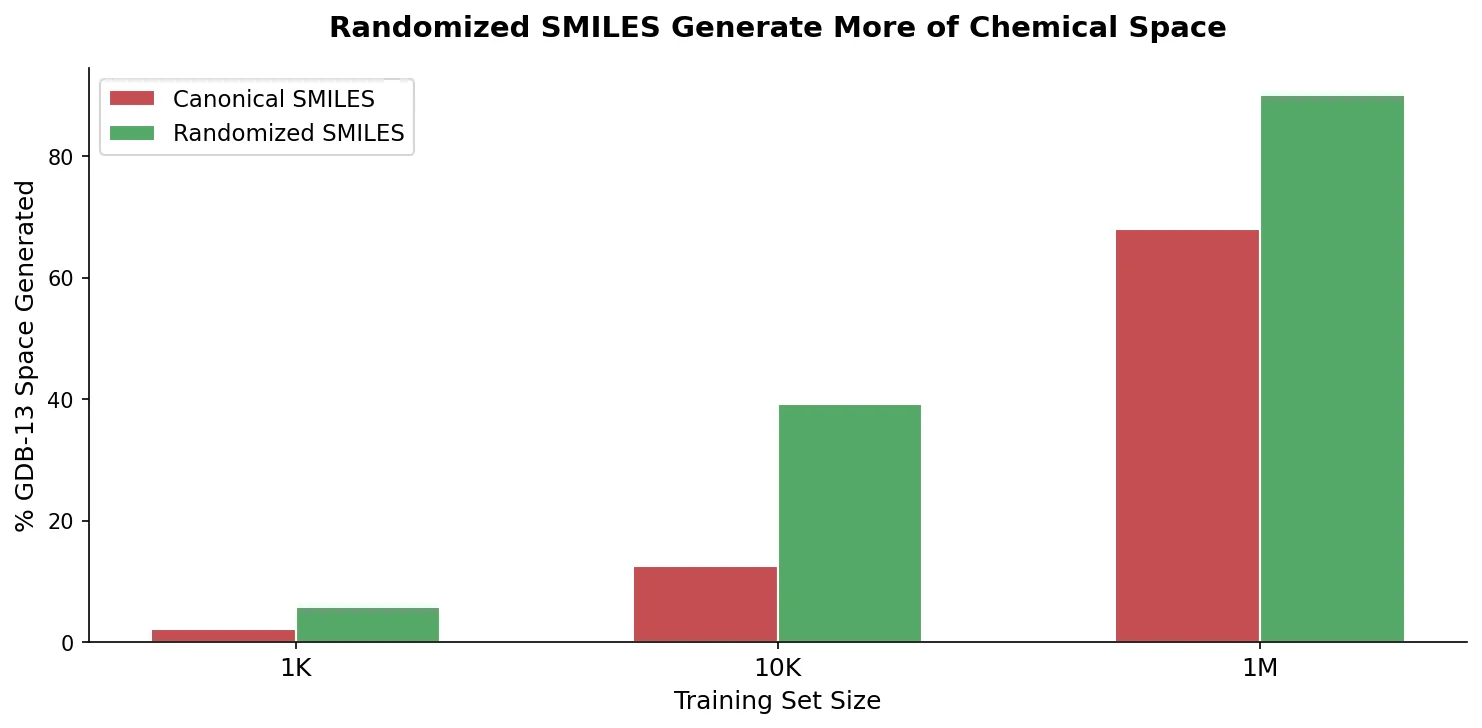
| 2019 | Randomized SMILES Improve Molecular Generative Models | Non-canonical SMILES augmentation improves RNN generation |
| 2024 | Invalid SMILES Benefit Chemical Language Models: A Study | Invalid SMILES generation improves CLMs via quality filtering |
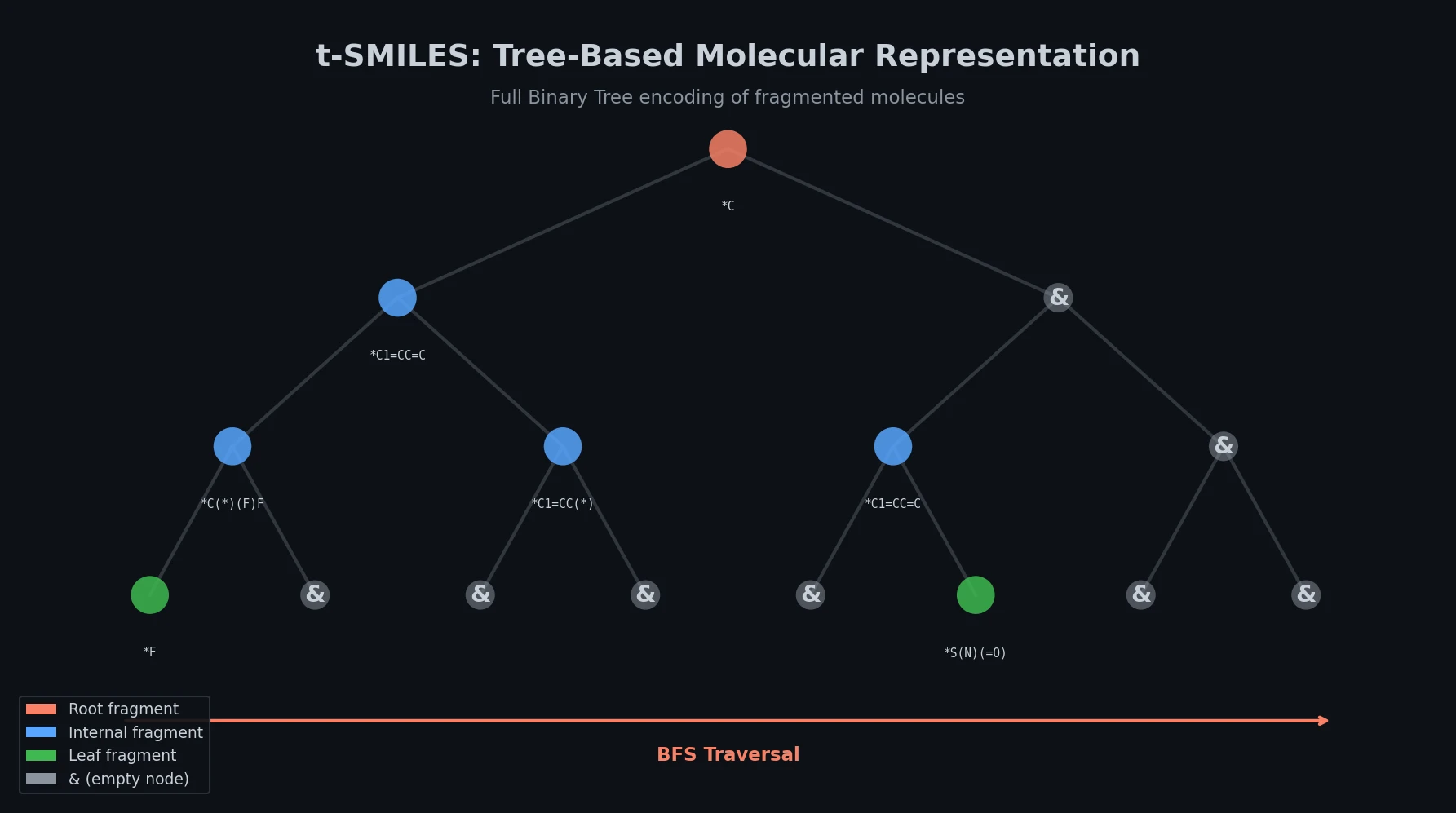
| 2024 | t-SMILES: Tree-Based Fragment Molecular Encoding | Fragment-based binary tree traversal reduces nesting depth |
SELFIES Family
| Year | Paper | Key Idea |
|---|---|---|
| 2020 | SELFIES: The Original Paper on Robust Molecular Strings | 100% robust molecular representation for generative ML |
| 2022 | SELFIES and the Future of Molecular String Representations | Perspective proposing 16 research directions for SELFIES |
| 2023 | Recent Advances in the SELFIES Library: 2023 Update | Streamlined grammar, aromatic support, performance upgrades |
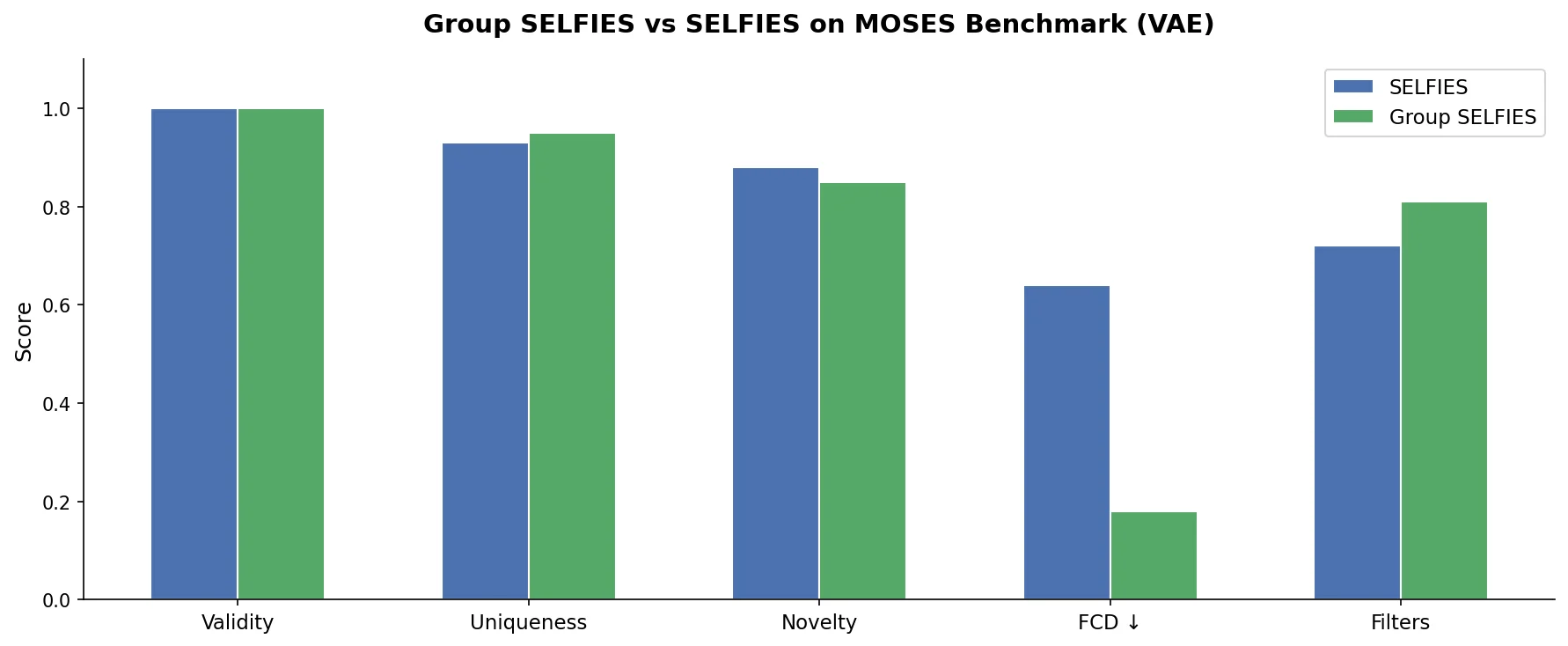
| 2023 | Group SELFIES: Fragment-Based Molecular Strings | Fragment group tokens extending SELFIES for distribution learning |
InChI Family
| Year | Paper | Key Idea |
|---|---|---|
| 2013 | InChI: The Worldwide Chemical Structure Identifier Standard | How InChI became the global standard for chemical identifiers |
| 2018 | RInChI: The Reaction International Chemical Identifier | Extends InChI to uniquely identify chemical reactions |
| 2019 | Mixfile & MInChI: Machine-Readable Mixture Formats | First standardized machine-readable formats for mixtures |
| 2020 | NInChI: Toward a Chemical Identifier for Nanomaterials | Extending InChI to represent multi-component nanomaterials |
| 2020 | InChI and Tautomerism: Toward Comprehensive Treatment | 86 tautomeric rules validated across 400M+ structures for InChI V2 |
| 2025 | Making InChI FAIR and Sustainable for Inorganic Chemistry | InChI v1.07 adds inorganic support and FAIR compliance |
Tokenization
| Year | Paper | Key Idea |
|---|---|---|
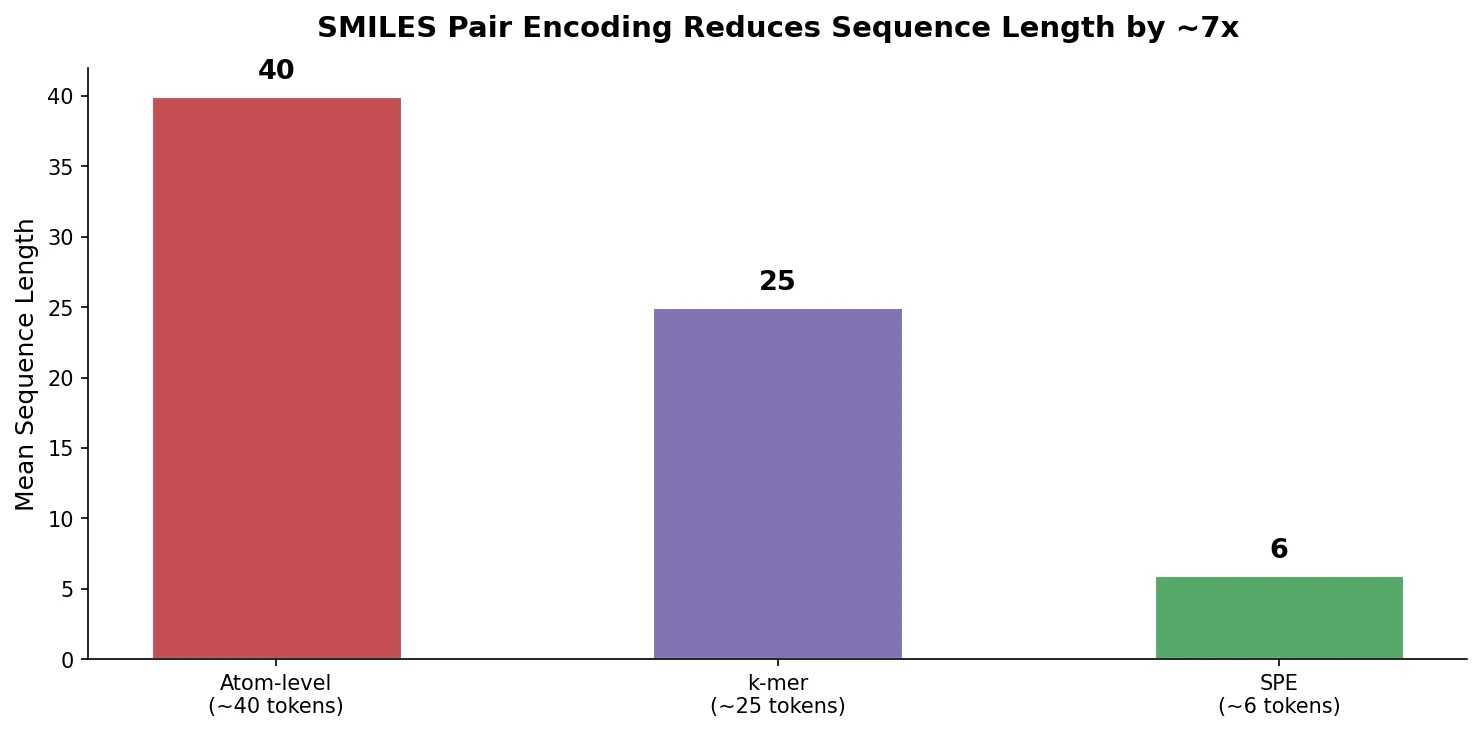
| 2021 | SPE: Data-Driven SMILES Substructure Tokenization | BPE-style algorithm learning chemically meaningful SMILES tokens |
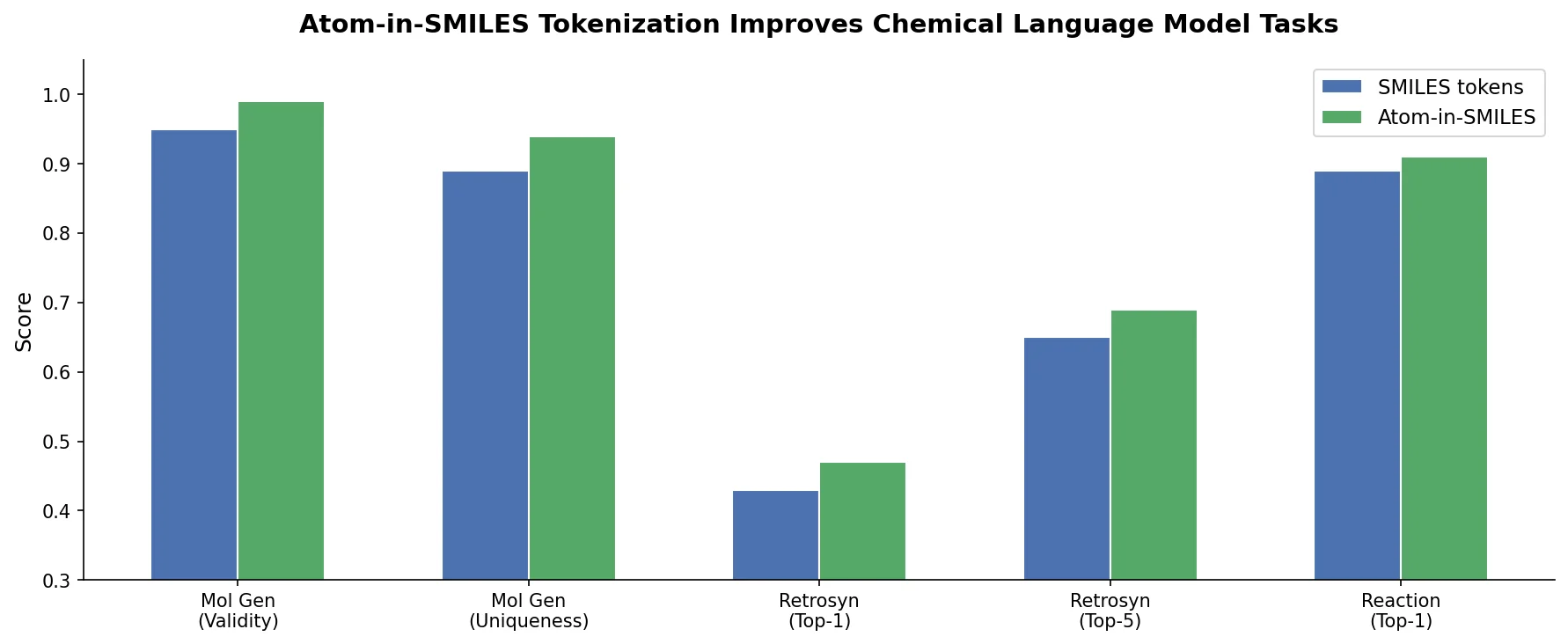
| 2023 | Atom-in-SMILES: Better Tokens for Chemical Models | Environment-aware atomic tokens reduce token degeneration |
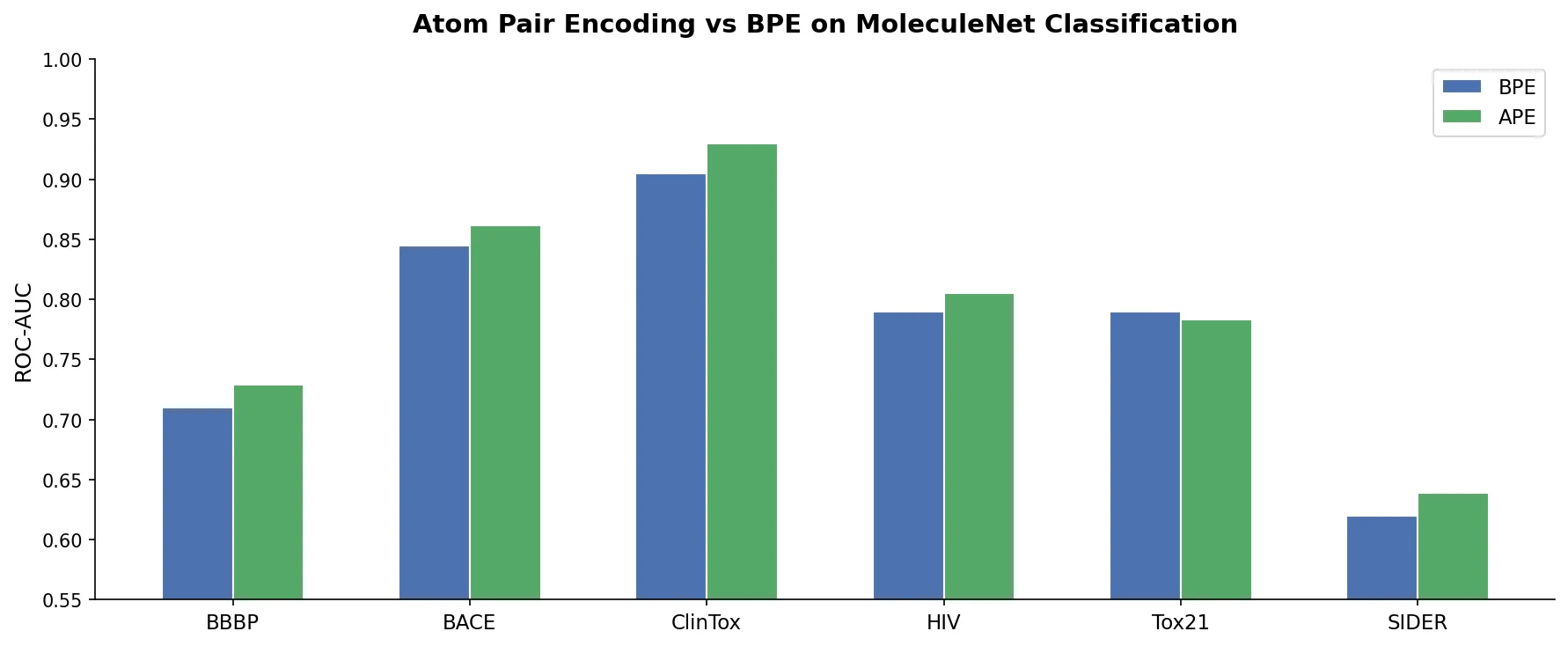
| 2024 | SMILES vs SELFIES Tokenization for Chemical LMs | Atom Pair Encoding tokenizer outperforms BPE on both formats |
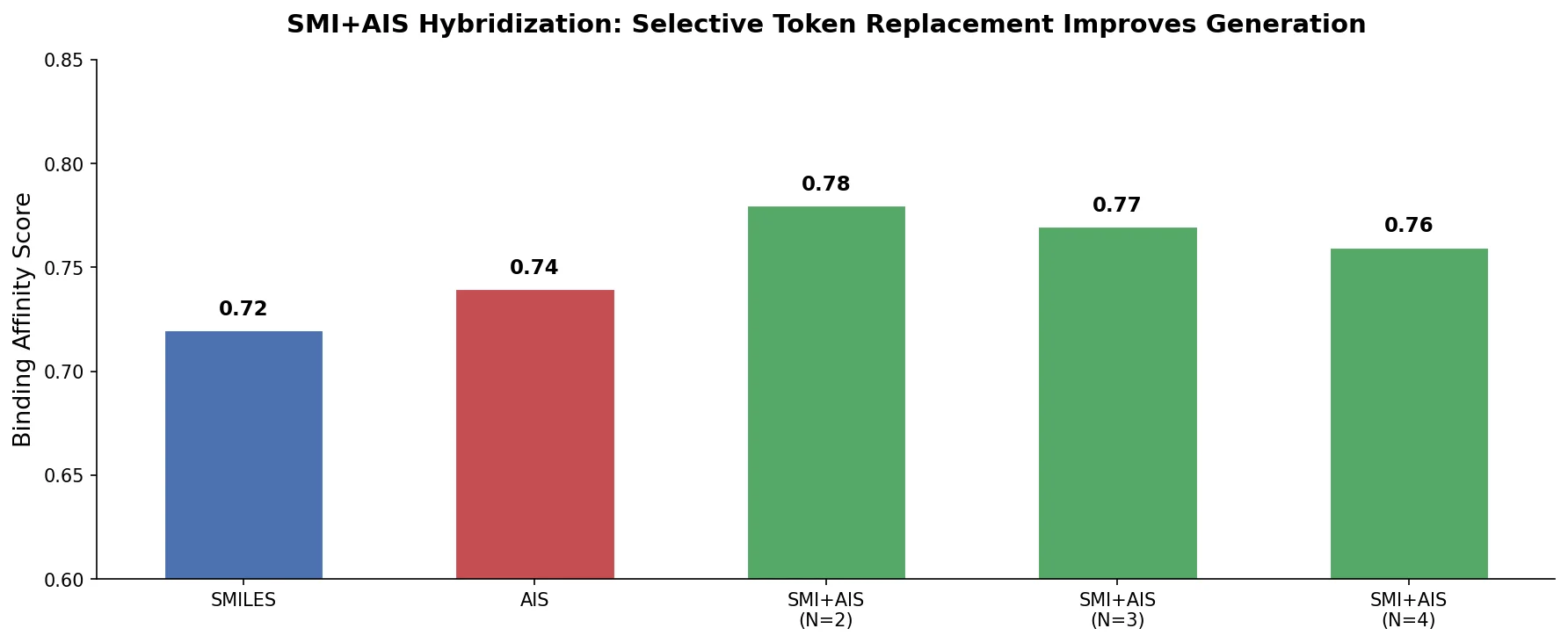
| 2025 | SMI+AIS: Hybridizing SMILES with Environment Tokens | Hybrid SMILES + Atom-In-SMILES tokens improve generation |
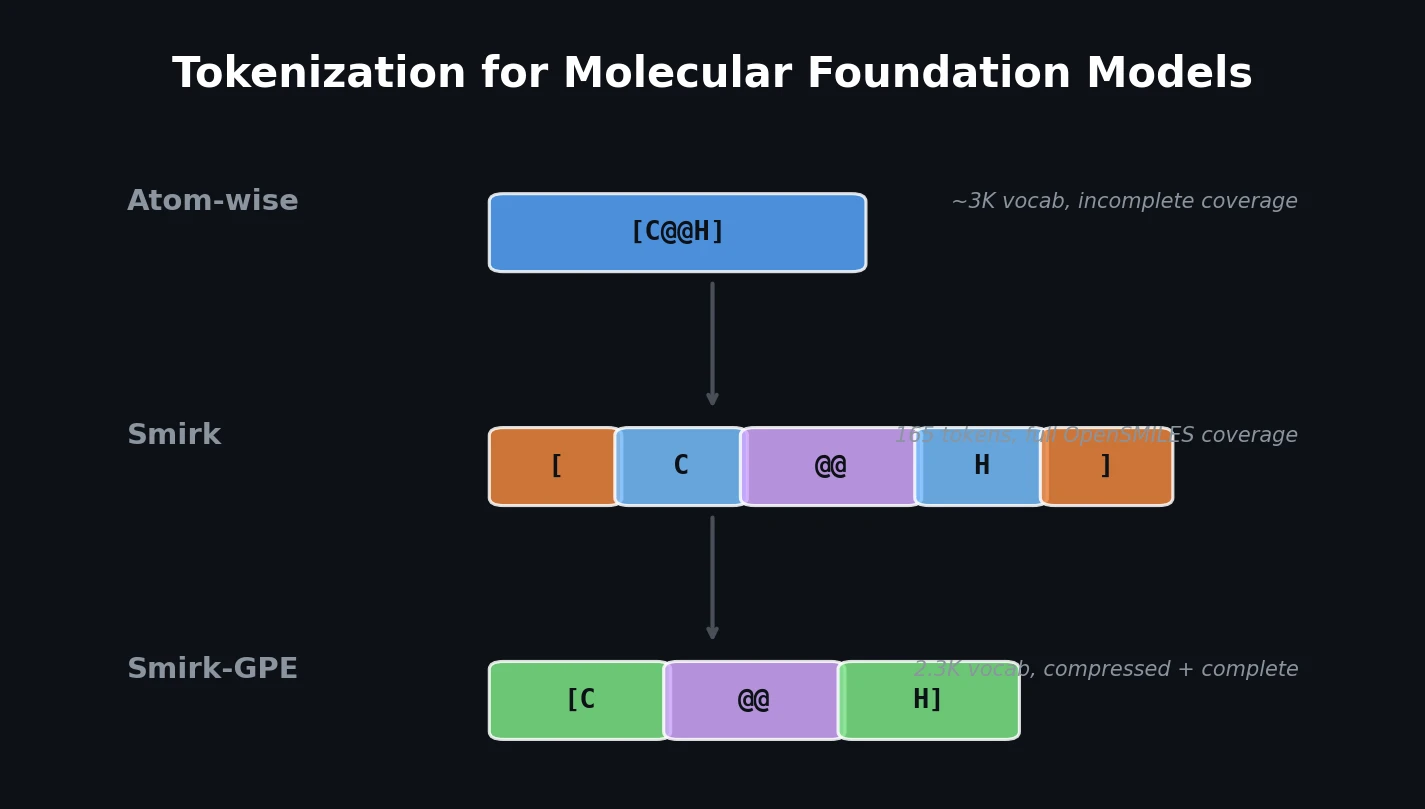
| 2026 | Smirk: Complete Tokenization for Molecular Models | 165-token set achieving full OpenSMILES specification coverage |
Surveys and Foundations
| Year | Paper | Key Idea |
|---|---|---|
| 1931 | The Number of Isomeric Hydrocarbons of the Methane Series | Recursive formulas for counting alkane structural isomers |
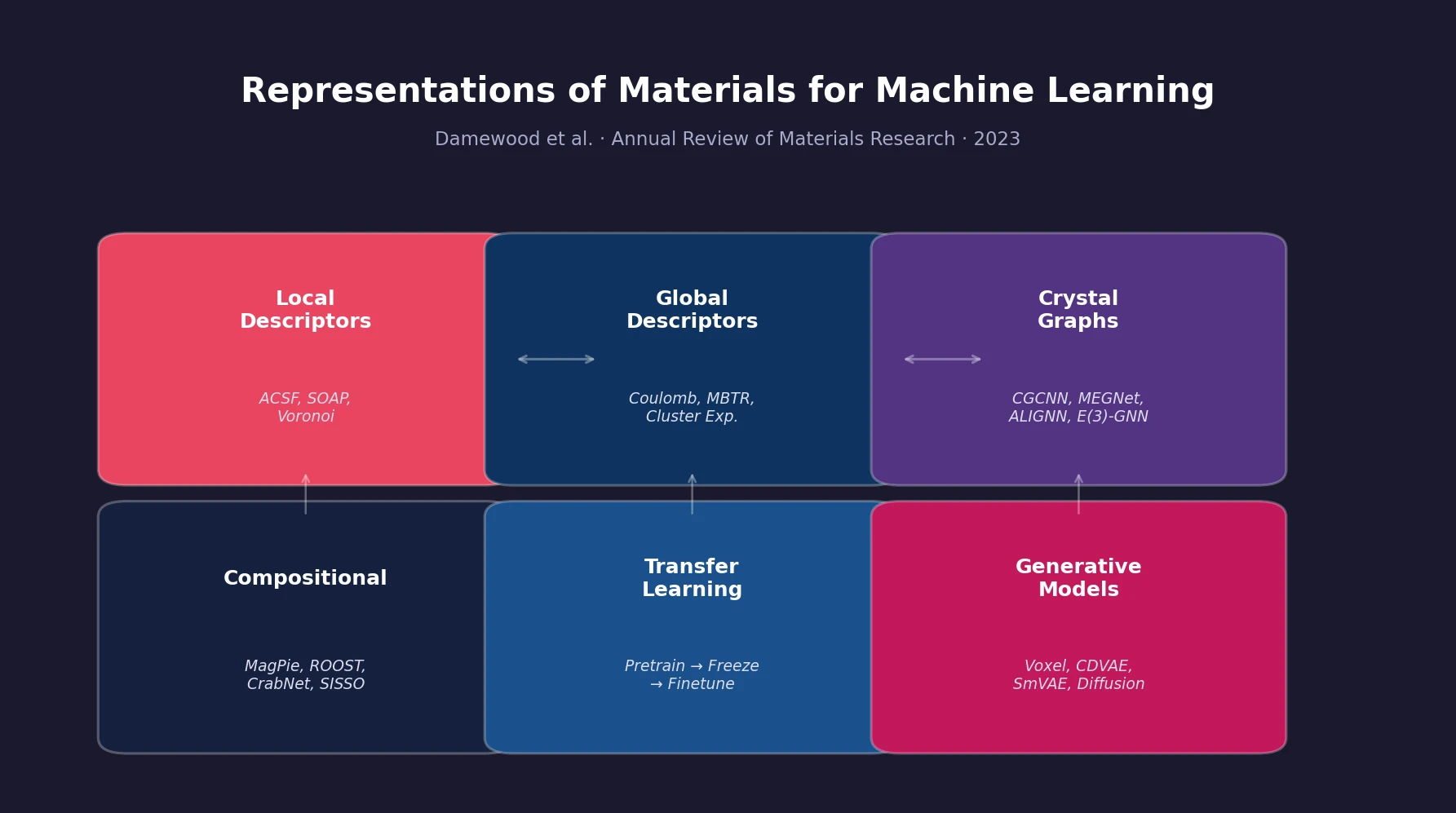
| 2023 | Materials Representations for ML Review | Review of solid-state material representations for ML |
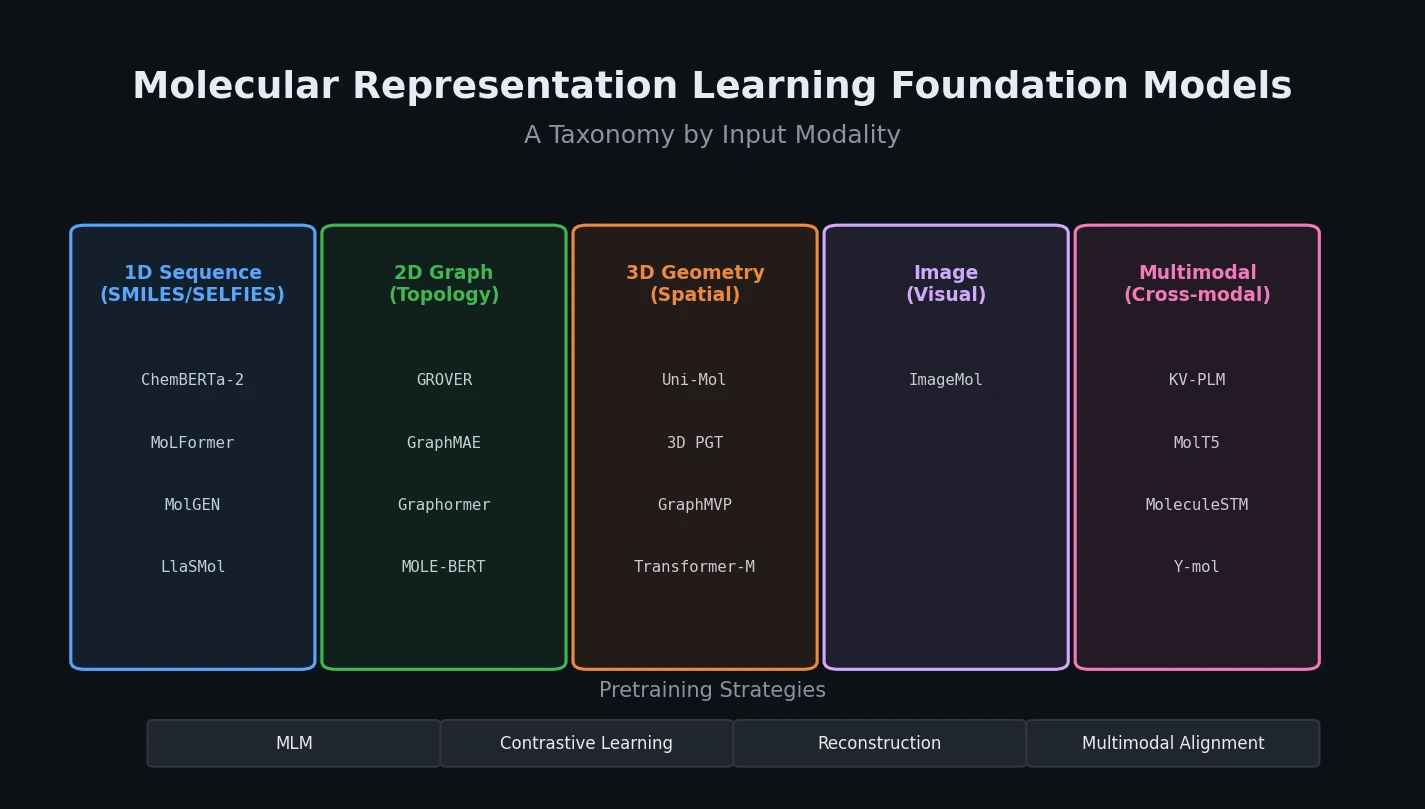
| 2025 | Review of Molecular Representation Learning Models | Survey of foundation models across five molecular modalities |