Models for predicting molecular or crystal properties from chemical string representations, plus benchmark suites and evaluation studies for assessing prediction quality.
Prediction Methods
| Paper | Year | Approach | Key Idea |
|---|---|---|---|
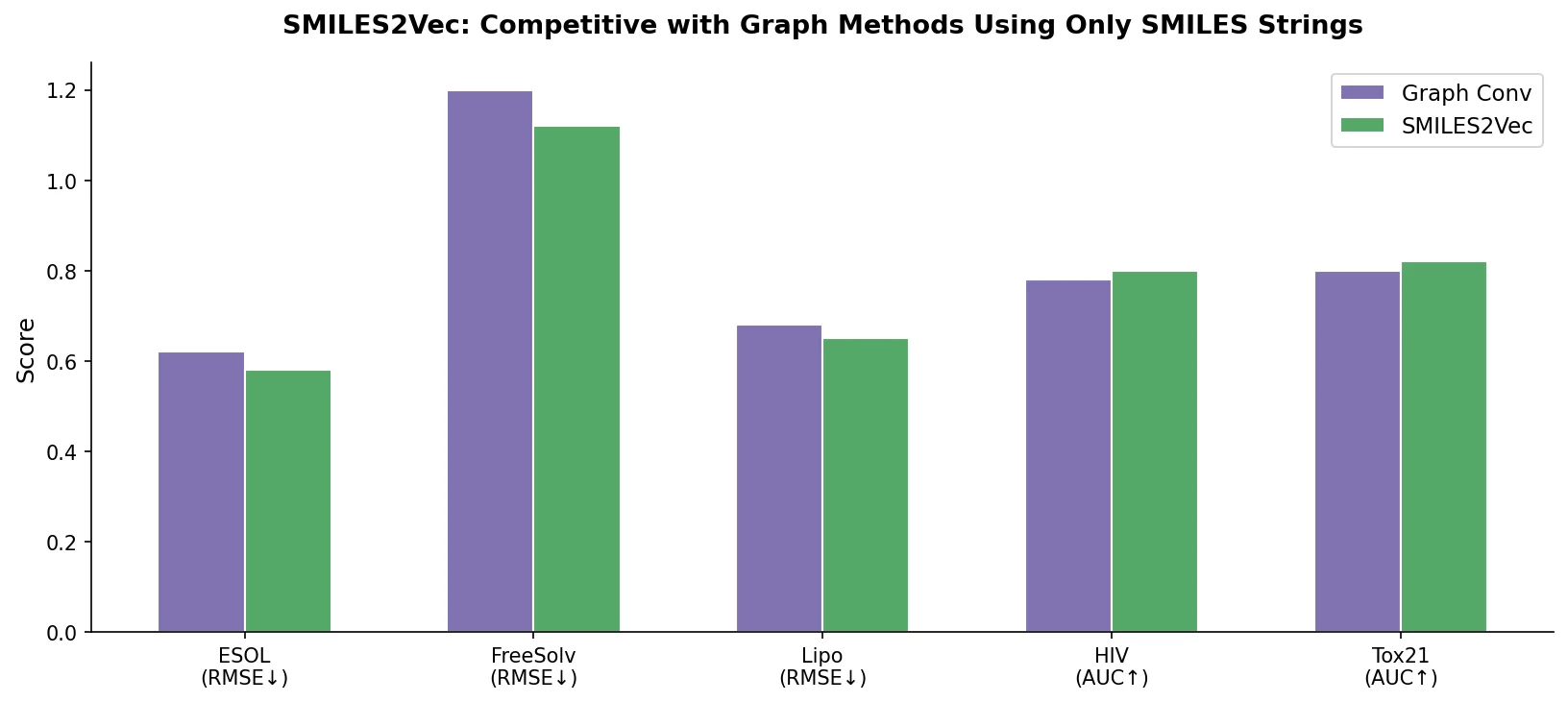
| SMILES2Vec | 2017 | CNN-GRU | Interpretable property prediction from raw SMILES embeddings |
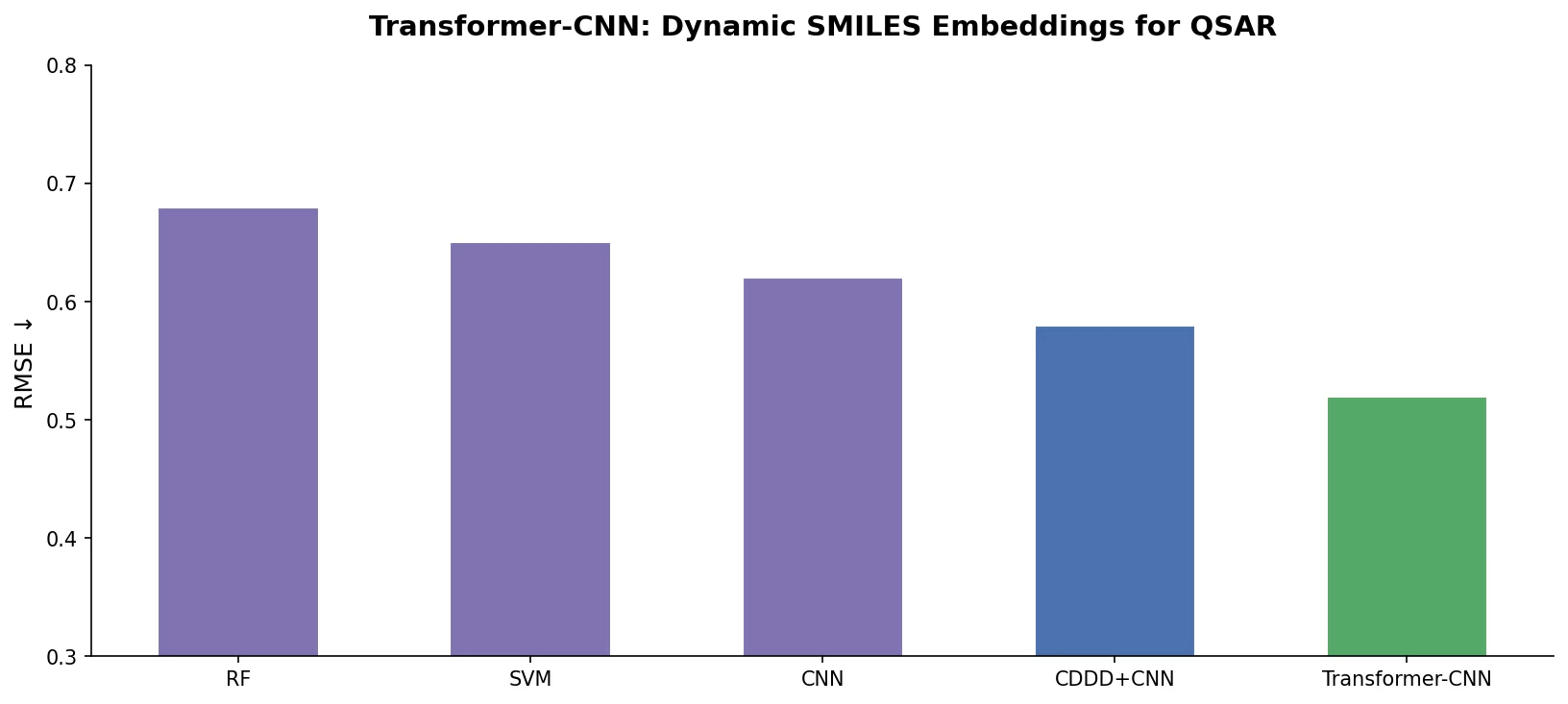
| Transformer-CNN | 2020 | Transformer + CNN | Transformer SMILES embeddings with CNN for interpretable QSAR |
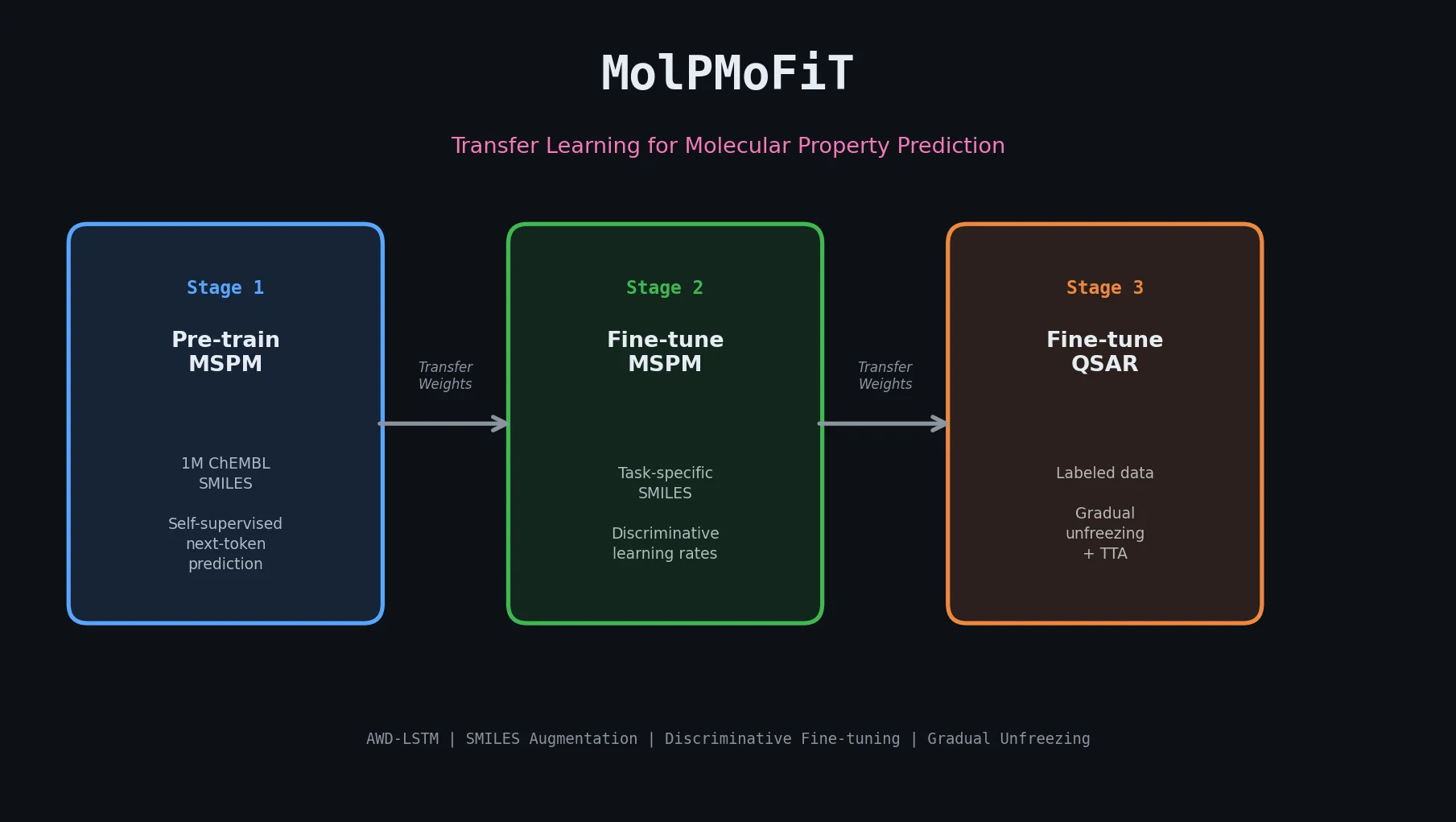
| MolPMoFiT | 2020 | Transfer learning | ULMFiT-style inductive transfer for QSAR on small datasets |
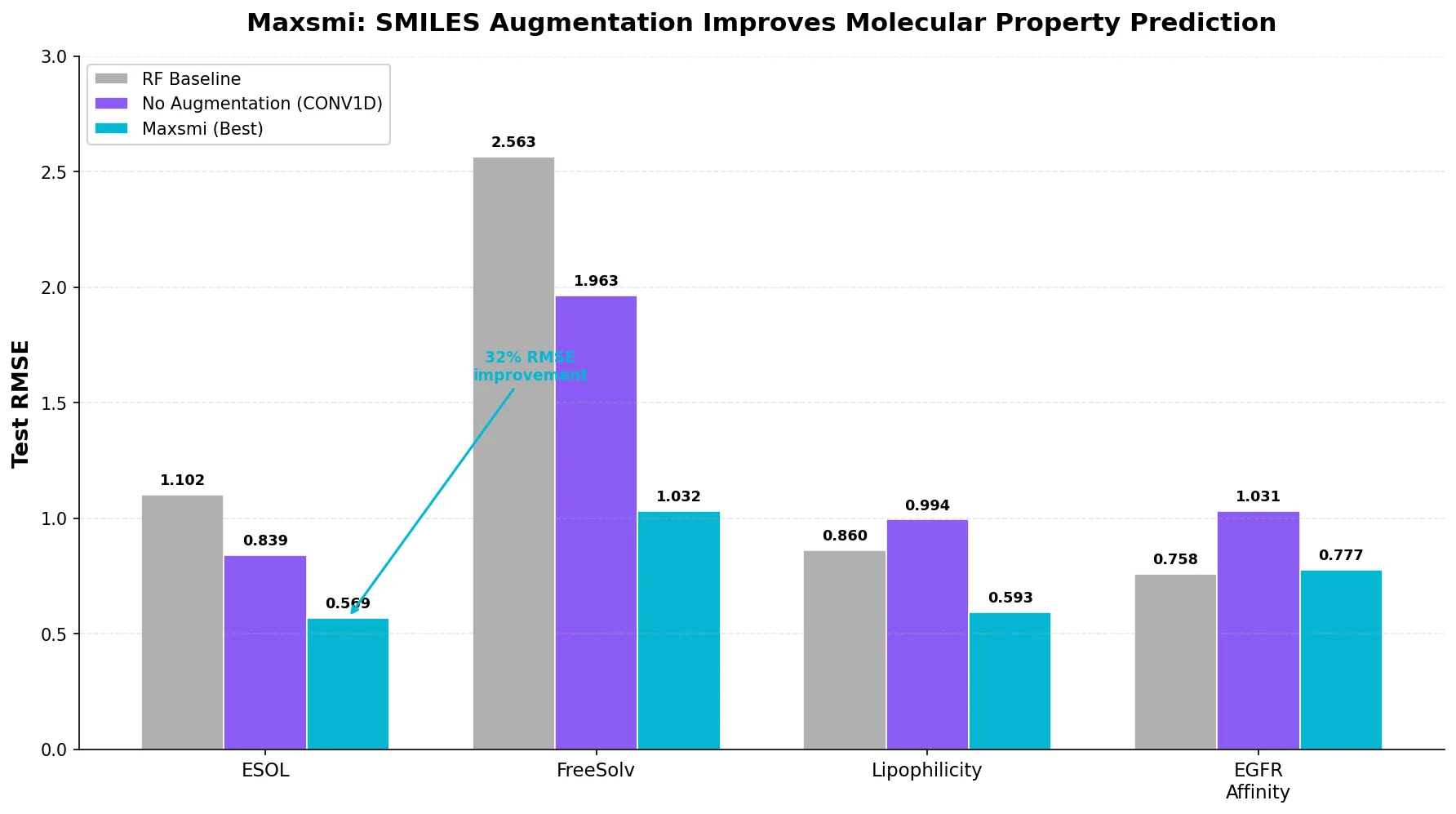
| Maxsmi | 2021 | CNN/RNN | SMILES augmentation improves CNN and RNN property prediction |
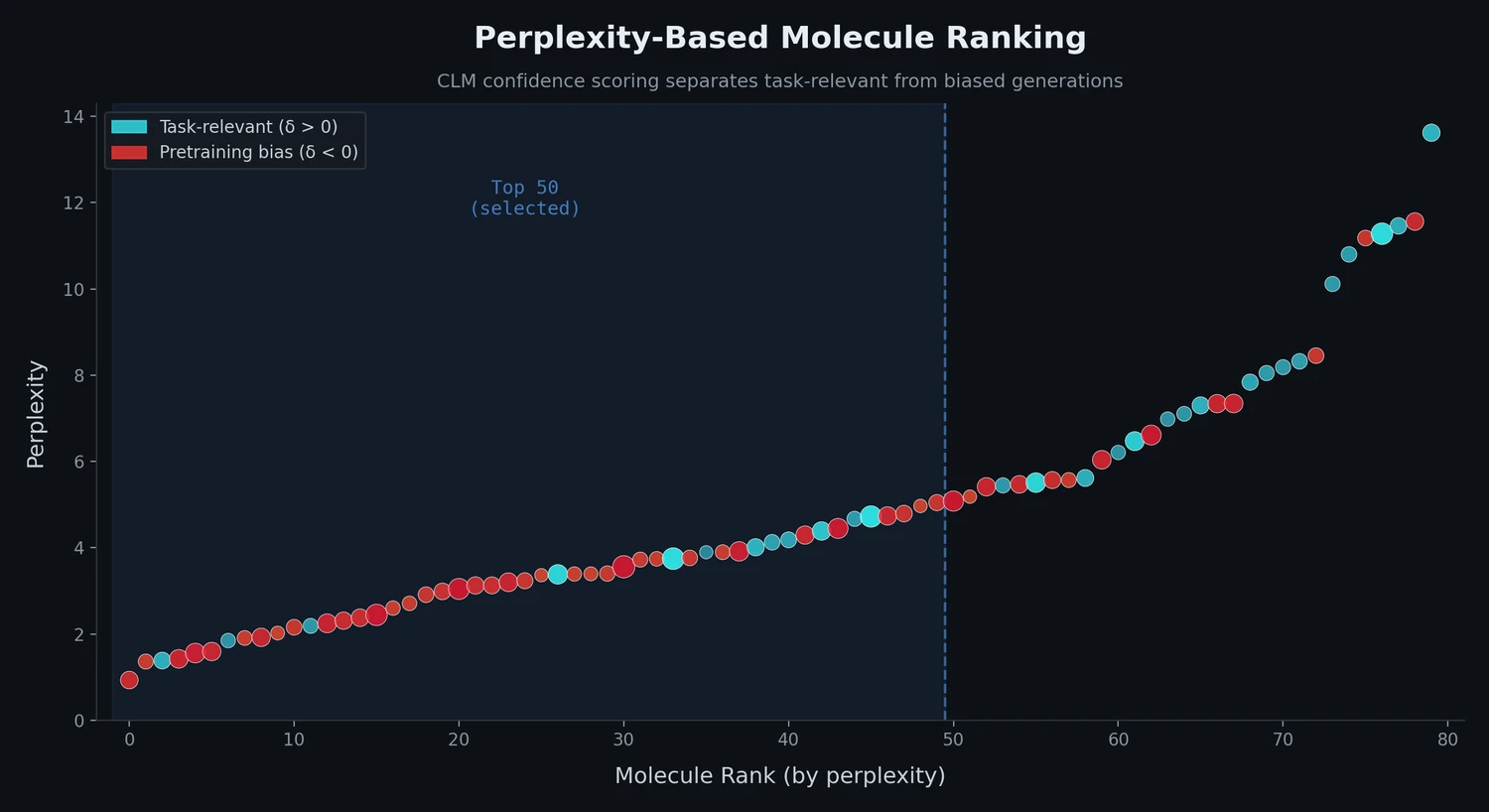
| Perplexity Ranking | 2022 | LM scoring | Perplexity scores rank molecules and detect pretraining bias |
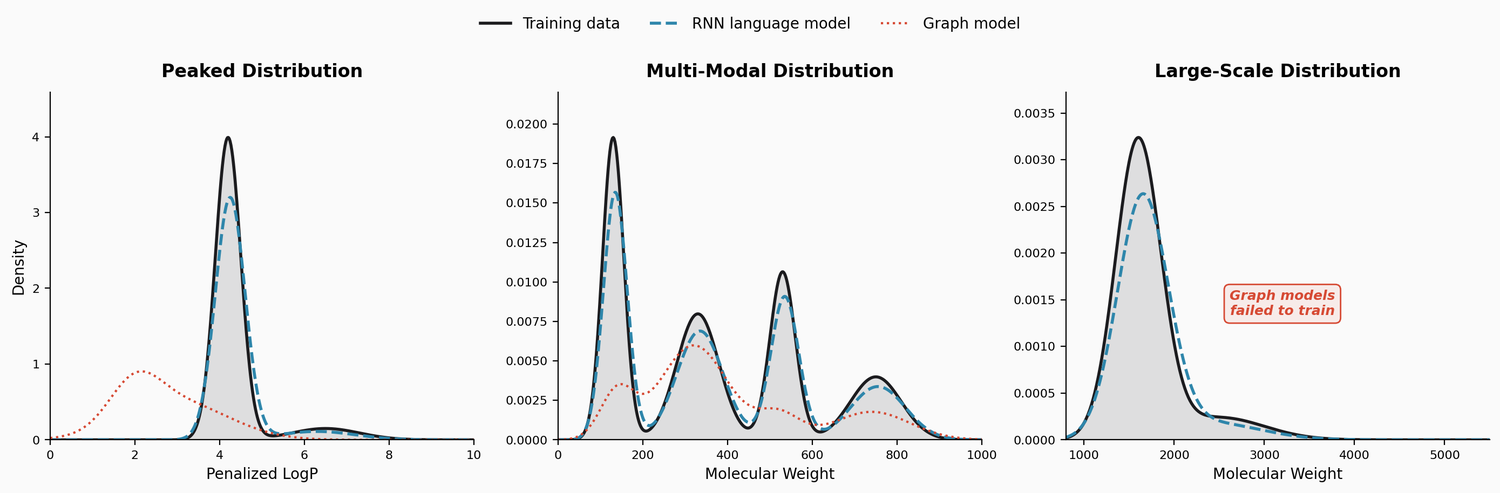
| LM Distributions | 2022 | RNN LM | RNN language models capture complex molecular distributions |
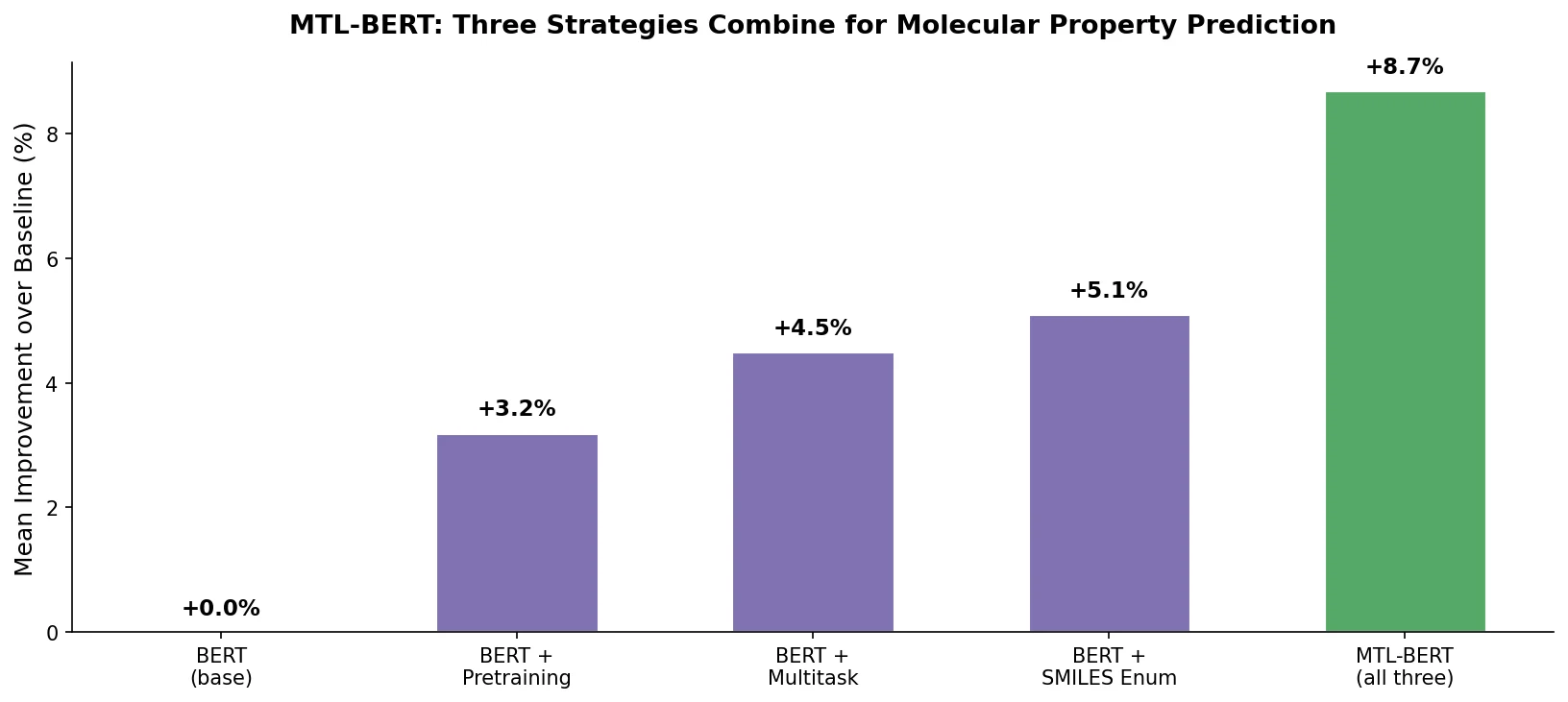
| MTL-BERT | 2022 | BERT | Multitask pretraining with SMILES enumeration augmentation |
| Regression Transformer | 2023 | Transformer | Unifies property prediction and conditional generation in one model |
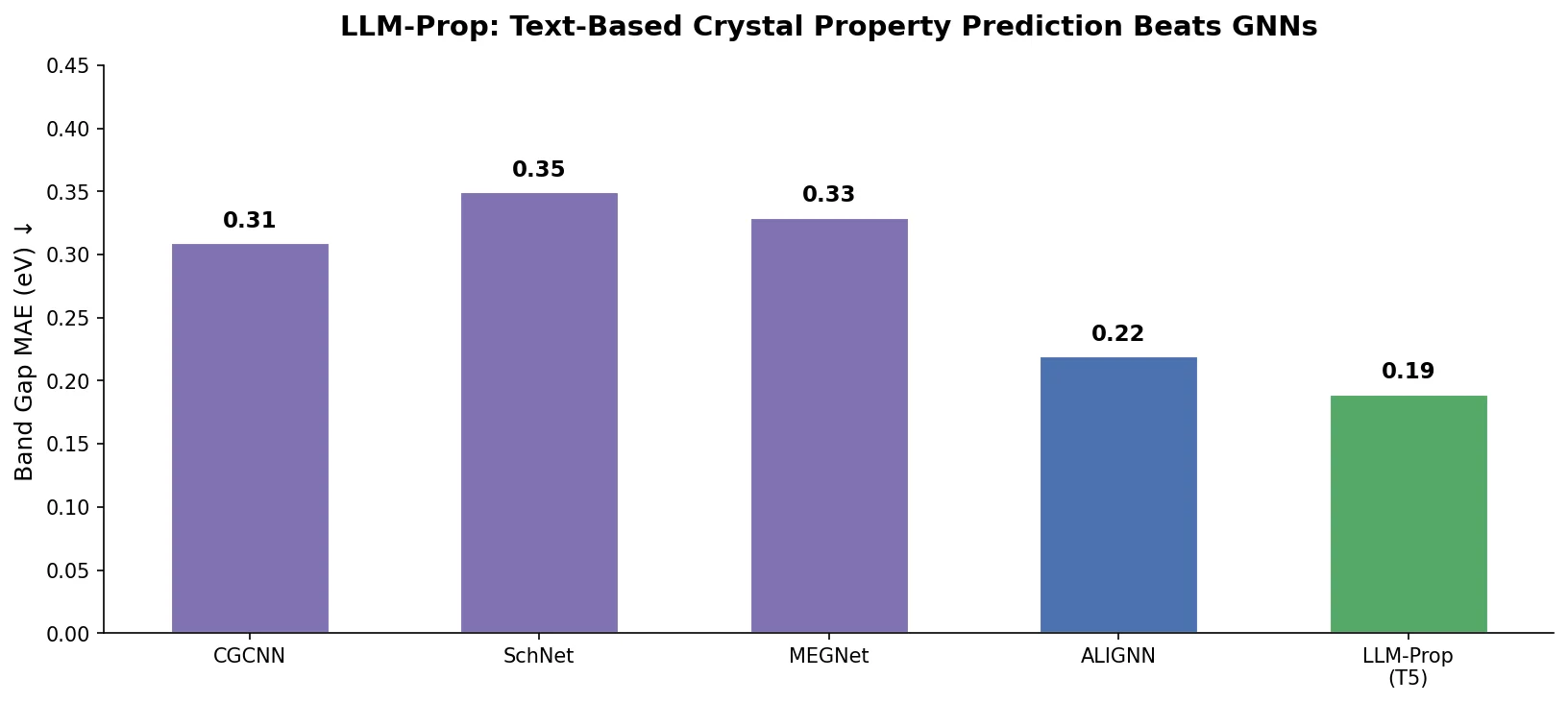
| LLM-Prop | 2025 | T5 | Crystal property prediction from text descriptions |
Benchmarks, Evaluation & Surveys
| Paper | Year | Key Idea |
|---|---|---|
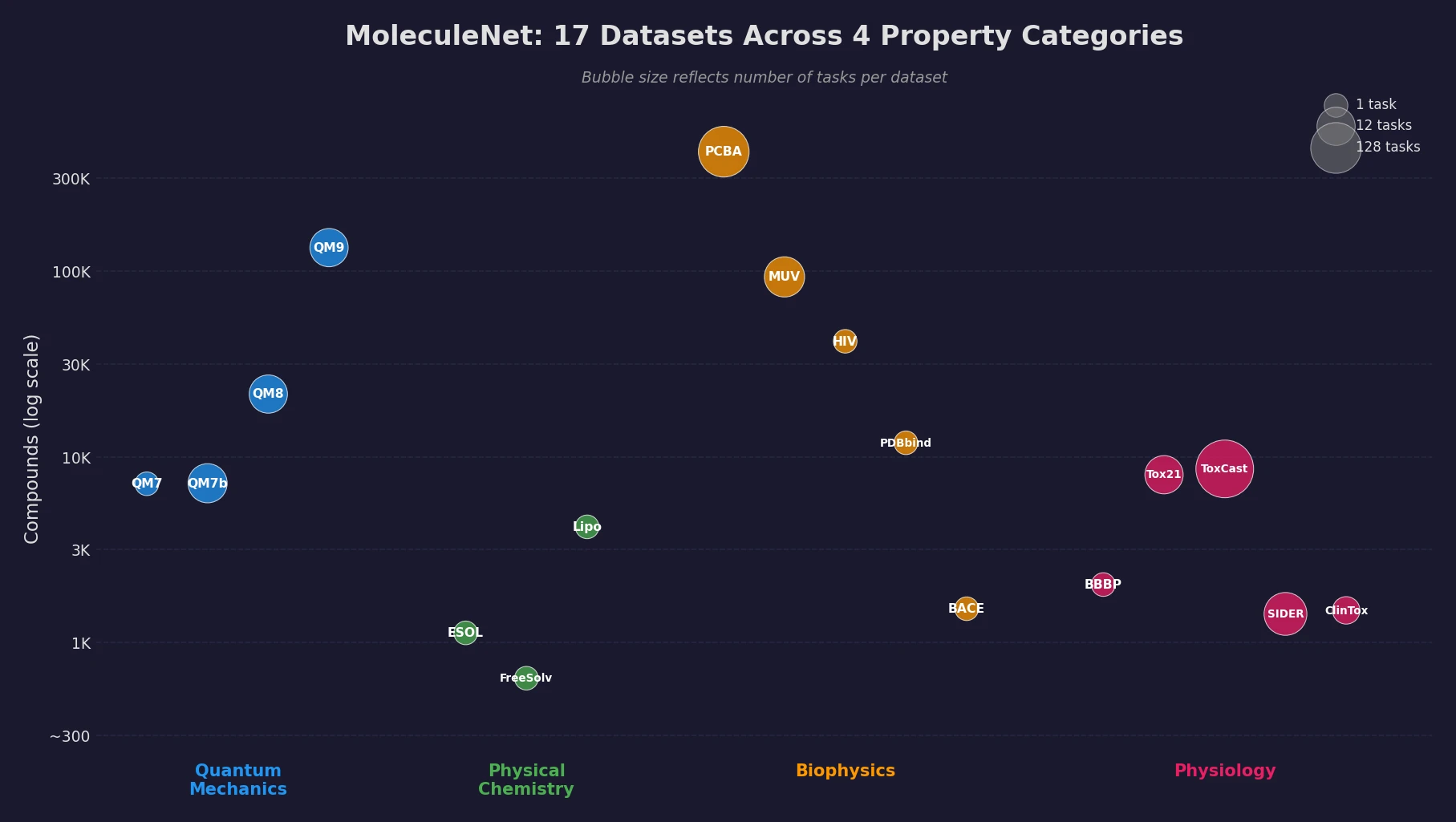
| MoleculeNet | 2018 | Benchmark suite across quantum mechanics, physical chemistry, biophysics, and physiology tasks |
| Activity Cliffs | 2022 | Exposes ML limitations where structurally similar molecules have very different activities |
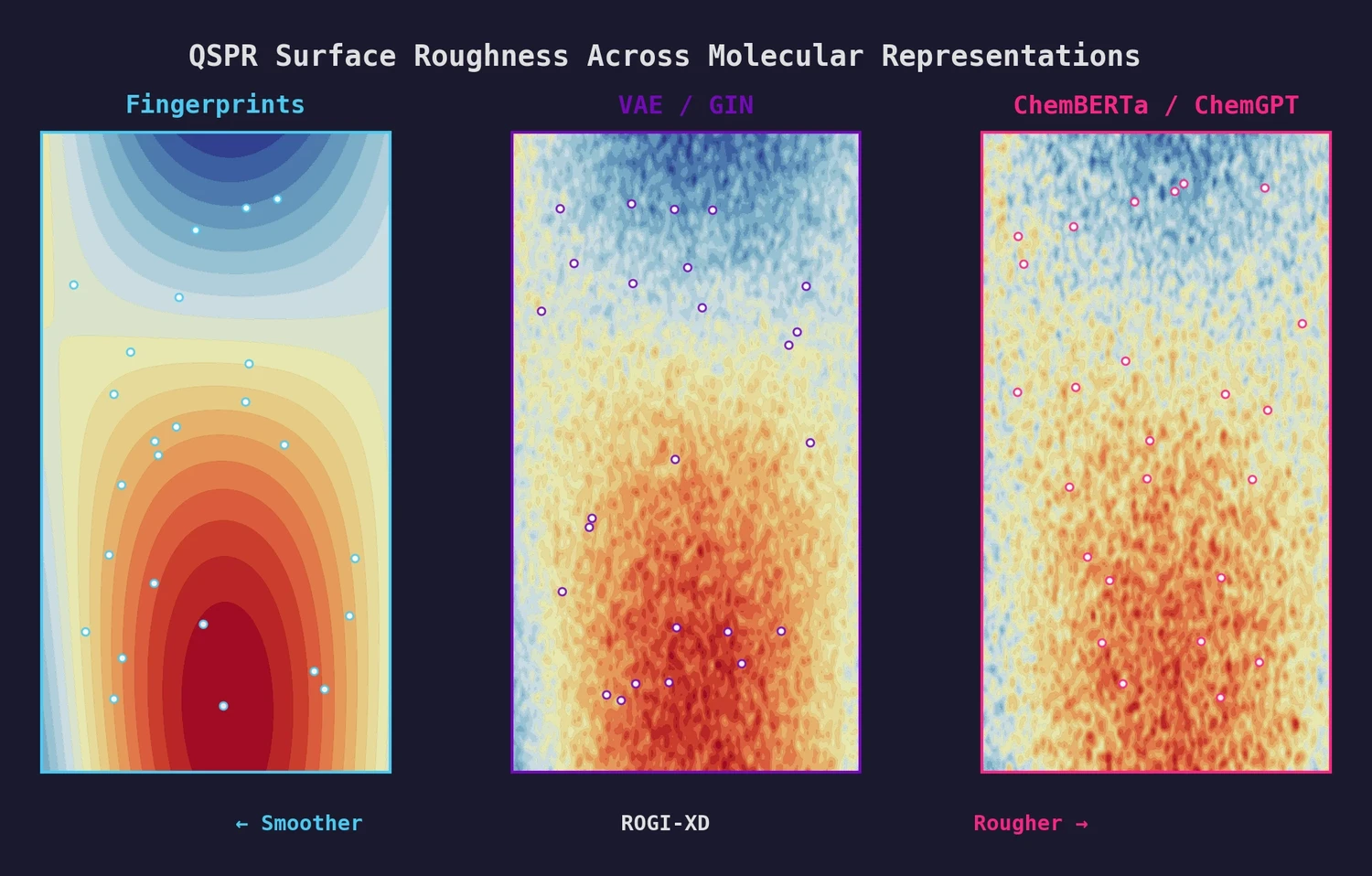
| ROGI-XD | 2023 | Task-independent measure of representation quality via structure-activity landscape roughness |
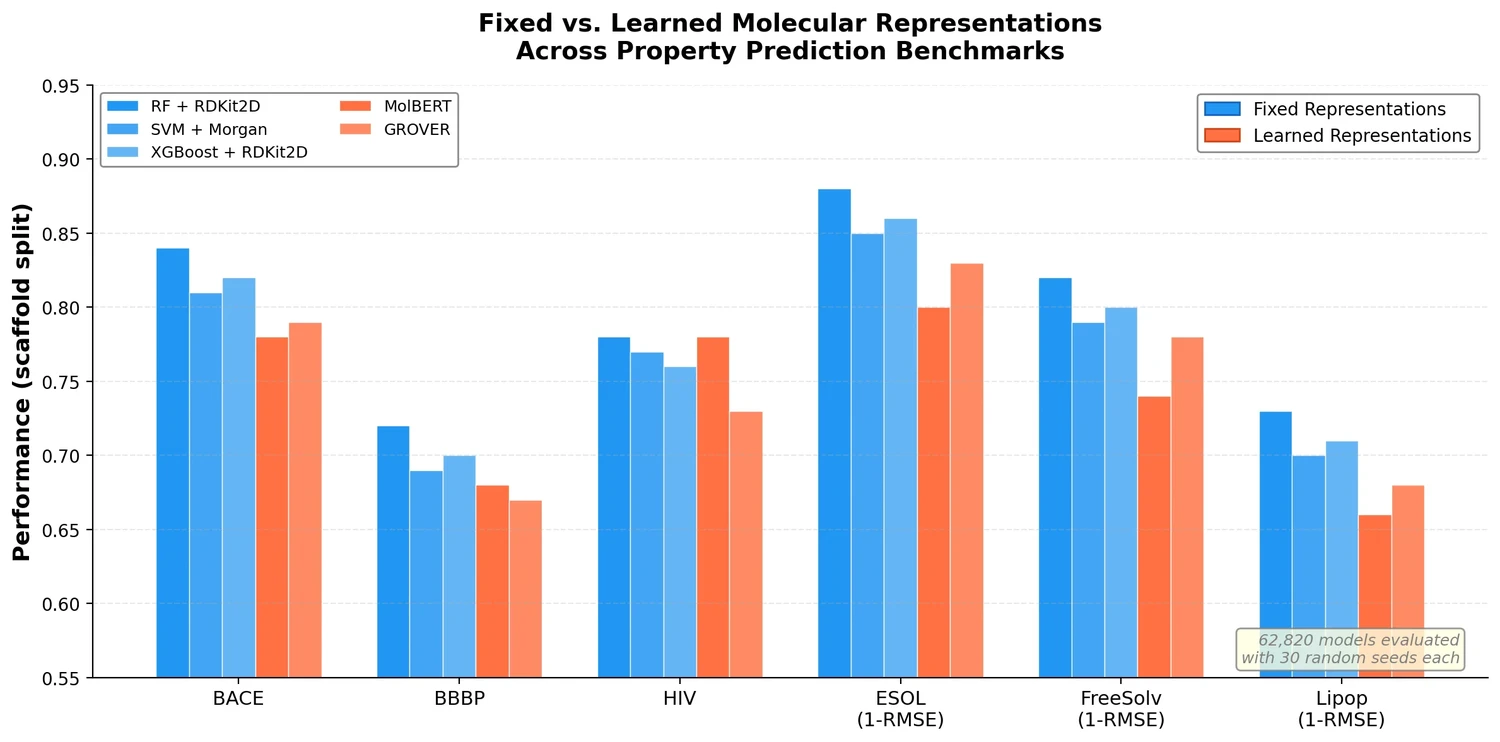
| Benchmarking at Scale | 2023 | Large-scale systematic comparison of molecular property prediction approaches |
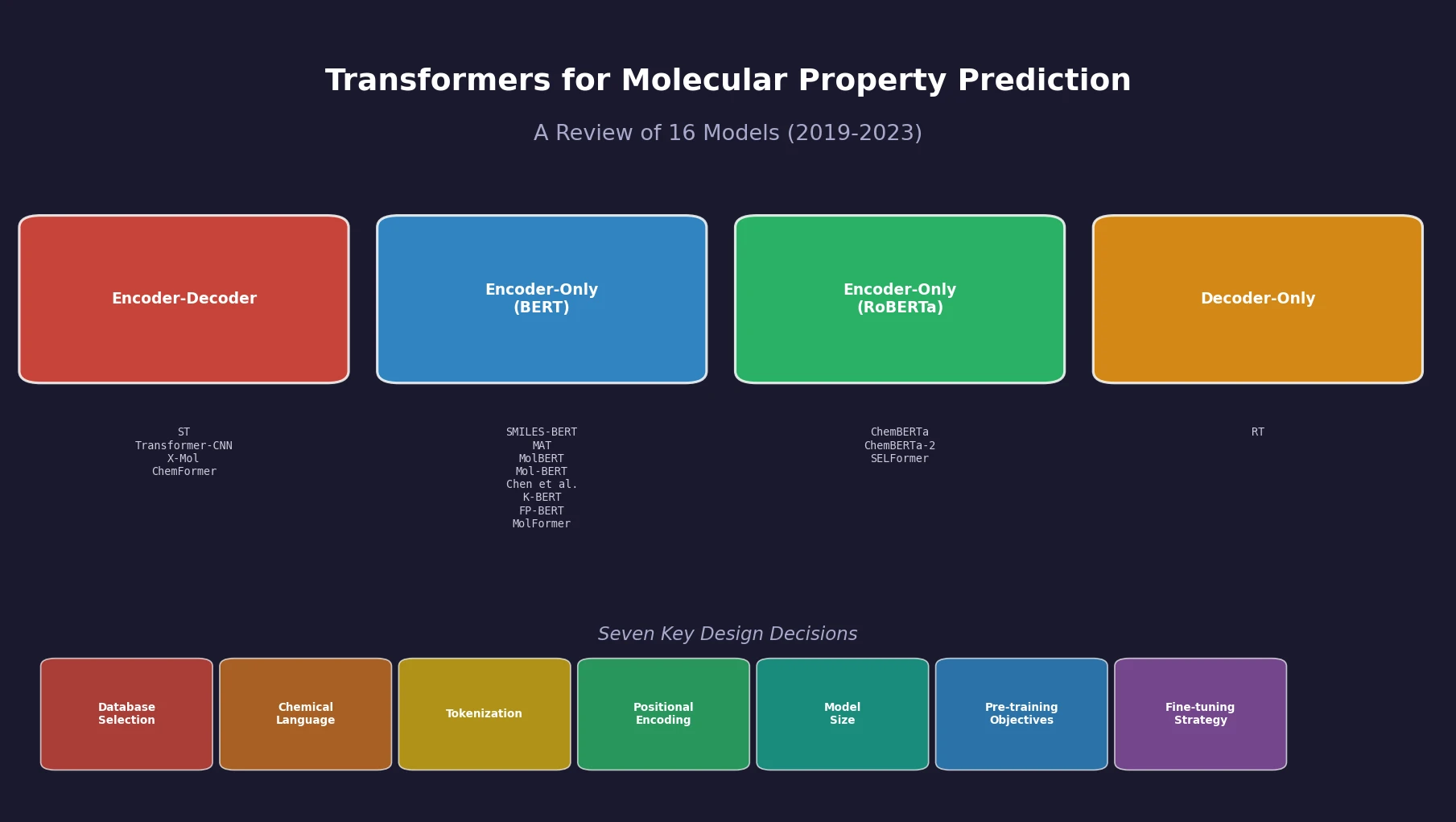
| Transformers for Property Prediction | 2024 | Review of transformer architectures applied to molecular property prediction |