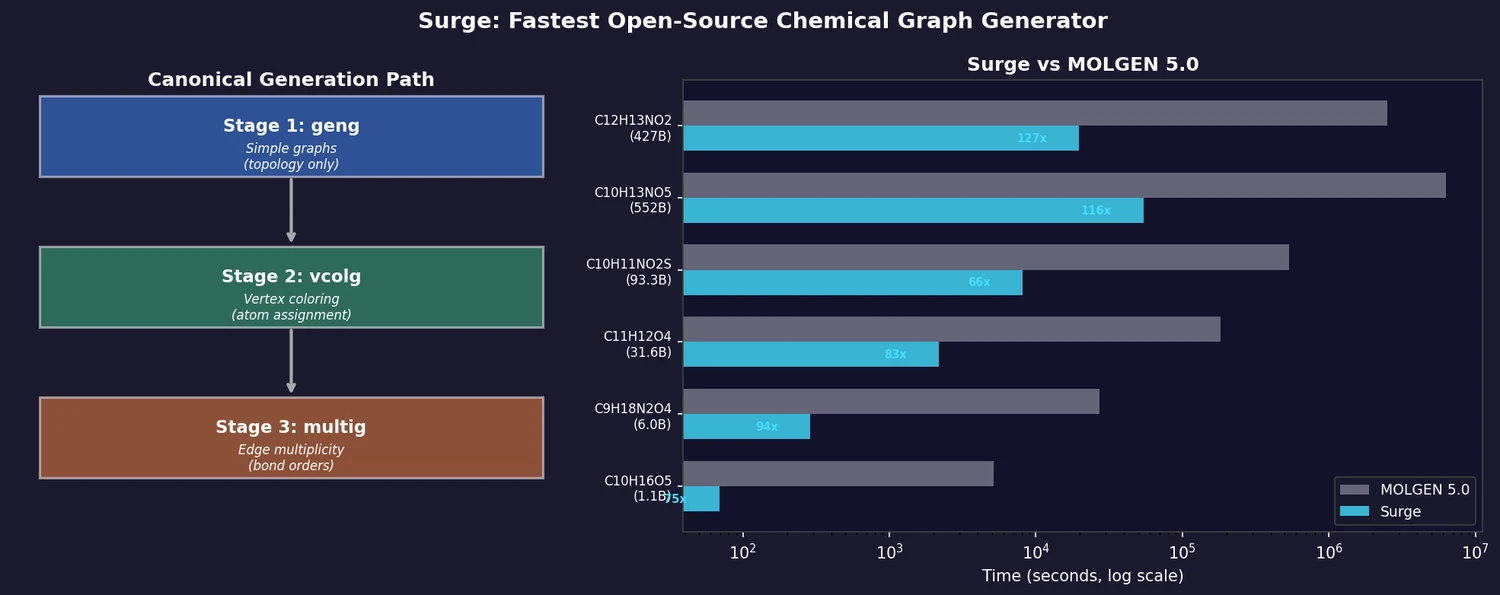
A Three-Stage Canonical Generation Path
Surge is an open-source constitutional isomer generator that enumerates all possible molecular structures for a given molecular formula. It is built on the nauty package for graph automorphism computation and uses a three-stage canonical generation path method that decomposes the enumeration problem into progressively refined graph operations. Surge outperforms the previous state-of-the-art (MOLGEN 5.0) by orders of magnitude in speed while running in under 5 MB of RAM regardless of molecule size.
Motivation: The Need for Fast, Open Structure Generators
Chemical structure generators are essential for computer-assisted structure elucidation (CASE), virtual library creation, and chemical space enumeration (e.g., GDB-17’s 166.4 billion molecules). MOLGEN had been the gold standard for decades but is closed-source. The previous best open-source alternative, MAYGEN, was roughly 3x slower than MOLGEN. Reymond’s lab used an in-house nauty-based generator for GDB-17 but did not release it publicly. Surge fills this gap as a fast, open-source, and extensible alternative.
The Three-Stage Algorithm
Given a molecular formula (e.g., $\text{C}_9\text{H}_{18}\text{N}_2\text{O}_4$), Surge proceeds through three stages:
Stage 1 (geng): Simple graph generation. Computes all connected simple graphs with the appropriate number of non-hydrogen atoms and edges, subject to maximum degree constraints from the molecular formula. These graphs represent molecular topologies without atom types or bond orders. For Lysopine ($\text{C}_9\text{H}_{18}\text{N}_2\text{O}_4$), this produces 534,493 graphs in 1.3 seconds.
Stage 2 (vcolg): Vertex coloring (atom assignment). Assigns element types (C, N, O, S, etc.) to vertices in all distinct ways, using the automorphism group of each simple graph to avoid generating equivalent assignments. Given a fixed ordering of elements (e.g., $\text{C} < \text{O} < \text{S}$), element assignments are represented as lists $L$ and compared lexicographically. Exactly one representative from each equivalence class is selected by computing the canonical (lexicographically maximal) list:
$$ \text{canon}(L) = \max\{\gamma(L) \mid \gamma \in \text{Aut}(G)\} $$
A list $L$ is accepted if and only if $\text{canon}(L) = L$, i.e., no automorphism produces a lexicographically larger list. For Lysopine, this expands to 3.0 billion vertex-labeled graphs in 90 seconds.
Stage 3 (multig): Edge multiplicity (bond orders). Assigns bond multiplicities (single, double, triple) to edges, again using automorphism group factorization to avoid duplicates. For Lysopine, this produces 6.0 billion completed molecules in an additional 100 seconds.
Efficient Automorphism Handling via Group Factorization
The key algorithmic innovation is the factorization of the automorphism group:
$$ \text{Aut}(G) = NM = \{\gamma\delta \mid \gamma \in N,; \delta \in M\} $$
where $M$ is the “minor subgroup” generated by transpositions of leaves sharing a common neighbor (“flowers”), and $N$ is a complete set of coset representatives computed by nauty. A flower is a maximal set of degree-1 vertices (leaves) with the same neighbor. The minor subgroup $M$ is normal in $\text{Aut}(G)$, making the factorization well-defined.
Theorem. A list $L$ satisfies $L = \text{canon}(L)$ if and only if $L = \max\{\delta(L) \mid \delta \in M\}$ and $L = \max\{\gamma(L) \mid \gamma \in N\}$.
This factorization enables efficient canonicity testing. Maximality under $M$ reduces to enforcing decreasing element order within each flower (simple inequality constraints during recursive assignment). Maximality under $N$ requires explicit testing against the $N$ generators, but $N$ is trivial (identity only) 58% of the time in Stage 2 and 98% of the time in Stage 3.
Benchmark Results
Benchmarked against MOLGEN 5.0 on 30 natural product molecular formulas from the COCONUT database on a compute-optimized c2-standard-4 Google Cloud VM, Surge achieves 7-22 million molecules per second with a memory footprint of at most 5 MB regardless of molecule size. Representative results:
| Formula | Isomers | Surge (s) | MOLGEN (s) | Speedup |
|---|---|---|---|---|
| $\text{C}_{10}\text{H}_{16}\text{O}_5$ | 1.1B | 69 | 5,146 | 75x |
| $\text{C}_9\text{H}_{18}\text{N}_2\text{O}_4$ | 6.0B | 289 | 27,250 | 94x |
| $\text{C}_{11}\text{H}_{12}\text{O}_4$ | 31.6B | 2,179 | 181,725 | 83x |
| $\text{C}_{10}\text{H}_{13}\text{NO}_5$ | 552B | 54,372 | 6,325,646 | 116x |
| $\text{C}_{10}\text{H}_{10}\text{N}_2\text{O}_3$ | 1.5T | 83,186 | 8,292,585 | 100x |
| $\text{C}_9\text{H}_{12}\text{N}_2\text{O}_5$ | 1.8T | 180,727 | 13,983,652 | 77x |
MOLGEN hit its built-in limit of $2^{31} - 1$ structures for most formulas; reported times were linearly extrapolated. Both generators were instructed to generate but not output structures. MOLGEN was run with -noaromaticity for fair comparison since Surge v1.0 lacks aromaticity detection.
Surge supports output in both SDfile and SMILES formats. SMILES output is produced efficiently by constructing a template for each simple graph at Stage 1, so that only atom types and bond multiplicities must be filled in before output.
Surge also supports built-in filters applied during generation (more efficient than post-hoc filtering):
-p0:1: at most one cycle of length 5-P: the molecule must be planar-B5: no atom has two double bonds and otherwise only hydrogen neighbors-B9: no atom lies on more than one cycle of length 3 or 4
These filter options are inspired by corresponding features in MOLGEN. Surge’s open-source design also supports a plugin mechanism: users can write small code snippets to insert custom filters into any of the three stages, enabling efficient pruning of the generation tree.
Limitations
- Version 1.0 does not perform Hückel aromaticity detection, so it generates duplicate Kekulé structures for aromatic rings that are graph-theoretically distinct
- Benchmarking against MOLGEN required disabling MOLGEN’s aromaticity detection (
-noaromaticity) for fair comparison - Written in C (from the nauty suite), which limits accessibility compared to Python-based tools, though this is also the source of its speed
Reproducibility Details
| Artifact | Type | License | Notes |
|---|---|---|---|
| Surge on GitHub | Code | Apache 2.0 | Official C implementation from the nauty suite |
| Surge project page | Other | Apache 2.0 | Project homepage with documentation and binaries |
- Status: Highly Reproducible. Source code, build instructions, and benchmark formulas are all publicly available.
- Hardware: Benchmarks used a compute-optimized c2-standard-4 Google Cloud VM. Surge runs in at most 5 MB of RAM regardless of molecule size.
- Build: Standard Unix Configure/Make scheme producing a standalone command-line executable. Written in portable C from the nauty suite.
- Dependencies: Requires the nauty package (bundled).
Paper Information
- Published: Journal of Cheminformatics, Volume 14, Article 24, April 23, 2022
- Preprint: ChemRxiv, December 7, 2021
- License: Apache 2.0 (software), Open Access (paper)
@article{mckay2022surge,
title={Surge: a fast open-source chemical graph generator},
author={McKay, Brendan D. and Yirik, Mehmet Aziz and Steinbeck, Christoph},
journal={Journal of Cheminformatics},
volume={14},
number={1},
pages={24},
year={2022},
publisher={BioMed Central},
doi={10.1186/s13321-022-00604-9}
}