Molecular Complexity as Branching in the Molecular Graph
This paper proposes two simple, interpretable measures of molecular complexity grounded in the observation that most GDB-enumerated molecules are synthetically challenging despite containing only standard functional groups and ring systems. The core insight is that branching points (non-divalent nodes) in the molecular graph correspond to synthesis difficulty: each additional branching point implies a new ring or substituent requiring extra synthetic steps, possible protecting groups, potential stereogenic centers, and increased steric hindrance.
Motivation: Why Most GDB Molecules Are Hard to Make
The Generated DataBases (GDBs) enumerate billions of hypothetical small organic molecules by exhaustively substituting atoms and bonds in mathematical graphs. Despite applying filters for ring strain, functional group diversity, fragment-likeness, drug-likeness, and ChEMBL-likeness, most enumerated molecules remain daunting to synthesize. Even in the most restrictive subset (GDB-13s, 99.4 million molecules from the 977 million in GDB-13), practical synthesis remains challenging for most entries. This motivated the search for a complexity measure that captures why these molecules are hard, without relying on reaction databases or machine learning.
MC1 and MC2: Two Graph-Based Complexity Measures
The two proposed measures are:
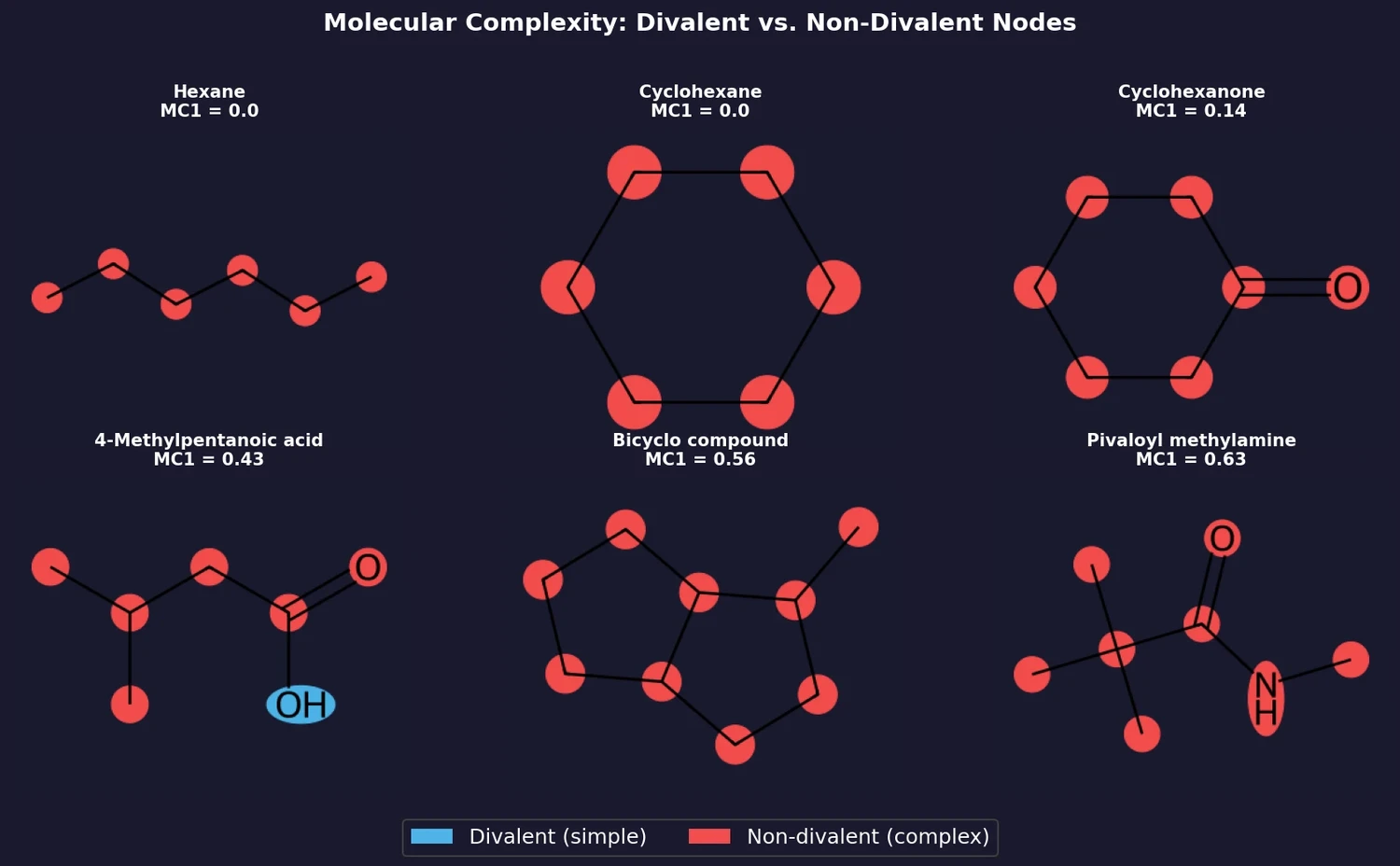
MC1 (size-independent): the fraction of non-divalent nodes in the molecular graph.
$$ \text{MC1} = 1 - \text{FDV} $$
where FDV is the fraction of divalent nodes (e.g., $-\text{CH}_2-$, $=\text{CH}-$, $=\text{C}=$, $-\text{O}-$, $-\text{NH}-$, $=\text{N}-$, $-\text{S}-$) in the molecular graph. The graph is computed by treating the molecule as if all bonds were single and all heavy atoms were carbon. MC1 is independent of molecule size, making it useful for comparing molecules of different sizes.
MC2 (size-dependent): the count of non-divalent nodes, excluding carbonyl carbons in standard carboxyl derivatives.
$$ \text{MC2} = \text{NDV} $$
where NDV is the number of non-divalent nodes, not counting $\text{C}{=}\text{O}$ in $(\text{X}-\text{C}{=}\text{O})$ for $\text{X} = \text{N}$ or $\text{O}$ (acids, esters, amides, carbonates, carbamates, ureas). MC2 grows with molecule size only when branching increases. Linear extensions (adding divalent atoms to chains or enlarging rings) do not increase MC2.
The rationale for excluding carboxyl groups from MC2 is that their chemistry (amide bond formation, esterification) is well-established and straightforward. Functional groups like amidines, guanidines, thioesters, thiones, sulfoxides, sulfinates, sulfones, and sulfonamides, as well as phosphorus-containing groups, are still counted because their synthesis is less routine.
Design Choices and Limitations
MC1 and MC2 deliberately do not distinguish between $\text{sp}^2$ and $\text{sp}^3$ branching points or count chiral centers. This choice is motivated by the observation that unusual substitution patterns on aromatic rings in GDB molecules are also synthetically difficult, and that functionalization of aromatic/heteroaromatic rings and control of atropisomerism in biaryls are both challenging. A consequence is that carbohydrates and polyphenols receive high complexity scores despite being abundant in biomass.
MC1 gives uninformative values for very small molecules (trifluoroacetic acid and tert-butanol both score $\text{MC1} = 1$) and for polymers (where the repeating unit dominates). MC2 similarly cannot give useful values for polymers due to its size dependence.
Comparison with Existing Complexity Measures
The authors compare MC1 and MC2 against six molecular complexity scores and two synthetic accessibility scores across four databases: GDB-13s, ZINC, ChEMBL, and COCONUT.
| Measure | Category | Description |
|---|---|---|
| FCFP4 | Complexity | Number of on-bits in a binary 2048-bit FCFP4 fingerprint |
| DataWarrior | Complexity | Fractal complexity via Minkowski-Bouligand (box-counting) dimension of distinct substructures up to 7 bonds |
| Böttcher | Complexity | Shannon entropy using additive atom contributions (valence electrons, atom environment, chirality, symmetry) |
| Proudfoot | Complexity | Shannon entropy using additive atom contributions (atomic number, connections, paths up to length 2) |
| SPS/nSPS | Complexity | Spacial score summing heavy atom contributions (hybridization, stereochemistry, nonaromaticity, neighbor count); nSPS normalizes by HAC |
| SAscore | Synthesizability | Fragment frequency from PubChem combined with complexity penalty (ring types, stereochemistry, size) |
| SCS | Synthesizability | Machine-learned score from 12 million Reaxys reactions predicting synthesis steps from ECFP4 fingerprint (max value 5) |
Key findings from the correlation analysis:
- For GDB-13s (where nearly all molecules have HAC = 13), complexity measures generally do not correlate with each other ($r^2 < 0.6$), except MC1 with MC2 and SPS with nSPS (expected, since each pair differs only in size normalization).
- For ZINC, ChEMBL, and COCONUT (spanning a broad range of molecular sizes), several complexity measures correlate with heavy atom count (HAC) and therefore with each other.
- Size-independent measures (DataWarrior, nSPS, SCS, SAscore, MC1) are unaffected by molecule size across datasets, while Böttcher and Proudfoot scores are strongly size-dependent. FCFP4 and SPS show partial size dependence.
- SPS and nSPS also correlate with SAscore.
The analysis is supported by interactive TMAP visualizations (tree-maps organized by MAP4C molecular fingerprint similarity) for 30,000 random molecules from each database, color-coded by each complexity measure. The interactive TMAPs are available online for GDB-13s, ZINC, ChEMBL, and COCONUT.
Reproducibility Details
| Artifact | Type | License | Notes |
|---|---|---|---|
| Molecular_Complexity | Code | MIT | Python implementation of MC1, MC2, and eight comparison metrics with Jupyter notebooks |
The paper is open access (hybrid). The GitHub repository provides Python code for computing MC1 and MC2 along with Jupyter notebooks demonstrating all ten complexity and synthesizability measures from Table 1. The four databases used (GDB-13s, ZINC, ChEMBL, COCONUT) are all publicly available. No model training or specialized hardware is involved, as MC1 and MC2 are deterministic graph computations.
Reproducibility status: Highly Reproducible.
Paper Information
- Journal: Journal of Chemical Information and Modeling, Vol. 65, No. 16, pp. 8405-8410
- Published: May 15, 2025
- Part of: Special issue “Chemical Compound Space Exploration by Multiscale High-Throughput Screening and Machine Learning”
@article{buehler2025view,
title={A View on Molecular Complexity from the GDB Chemical Space},
author={Buehler, Ye and Reymond, Jean-Louis},
journal={Journal of Chemical Information and Modeling},
volume={65},
number={16},
pages={8405--8410},
year={2025},
publisher={American Chemical Society},
doi={10.1021/acs.jcim.5c00334}
}