Exhaustive Hydrocarbon Enumeration Without Exclusion Filters
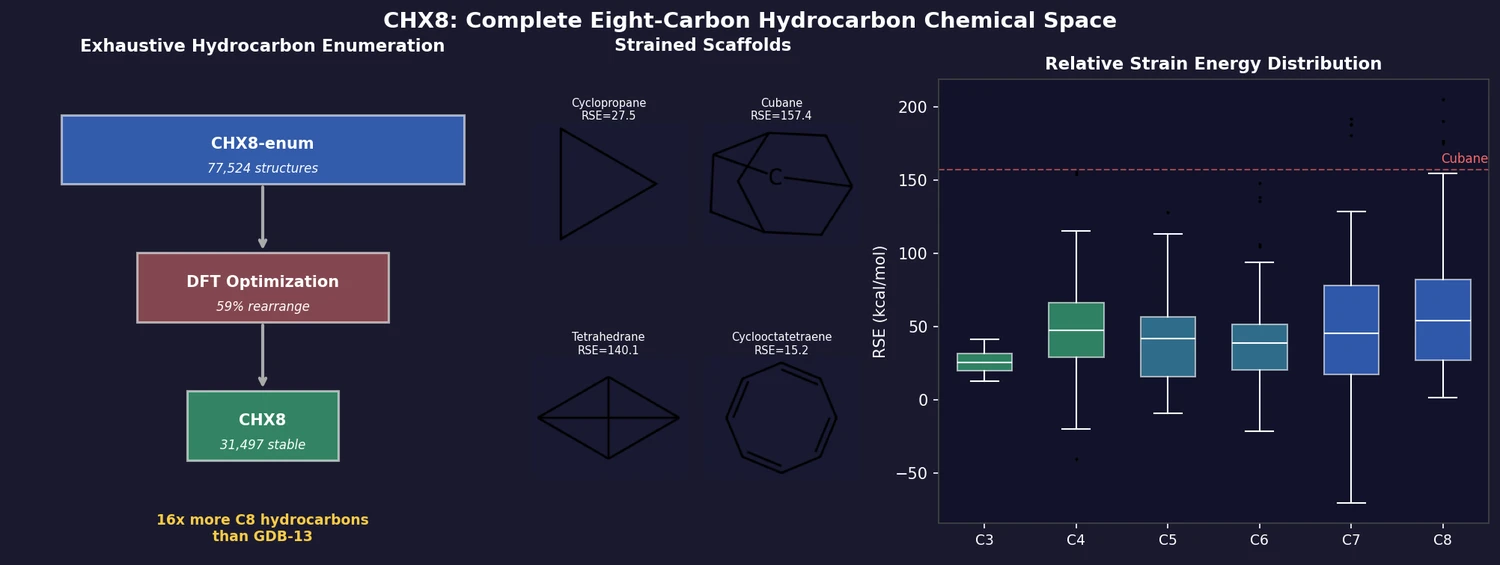
CHX8 is the first dataset to fully enumerate all closed-shell hydrocarbons with up to eight carbon atoms, deliberately including strained, anti-Bredt, and unconventional architectures that prior enumerations (e.g., GDB-13, GDB-17) excluded. Of 77,524 enumerated structures, 31,497 are stable under DFT optimization, covering 16x more C8 hydrocarbons than GDB-13. A universal relative strain energy (RSE) metric provides a quantitative synthesizability proxy for every molecule.
Motivation: Strained Scaffolds Are No Longer Inaccessible
GDB-series databases applied strict filters during enumeration, excluding highly strained polycyclic systems, cyclic allenes, anti-Bredt frameworks, and other “unconventional” motifs. Recent synthetic advances have shown that many of these structures can be accessed and exploited: 3D strained bioisosteres improve pharmacokinetic properties, cyclic allenes enable rapid construction of complex skeletons, and anti-Bredt olefins can be generated and trapped stereospecifically. CHX8 deliberately retains all of these motifs to provide a future-proofed database that remains relevant as synthetic capabilities expand.
Enumeration and Optimization
CHX8-enum (77,524 structures): All mathematically feasible hydrocarbons generated by exhaustively enumerating saturated carbon frameworks using the GENG tool from the nauty graph-isomorphism package (all 1-to-8-node connected graphs with 1-4 edges per node), then converting graphs to 3D coordinates via OpenBabel’s --Gen3D with the MMFF94 force field. Unsaturations (double bonds, triple bonds, allenes) were introduced iteratively in all valid positions by identifying C-C bonds flanked by hydrogen atoms (SMARTS: [#1]~[#6]~[#6]~[#1]), removing H atoms, and incrementing bond order. Point diastereoisomers and E/Z isomers were generated by manipulating InChI chiral layers. Duplicate detection relied on canonical InChI strings; residual duplicates account for no more than 1.5% of CHX8.
| HAC | Graphs | Saturated | Unsaturated | CHX8-enum | CHX8 (stable) |
|---|---|---|---|---|---|
| 1 | 1 | 1 | 0 | 1 | 1 |
| 2 | 1 | 1 | 2 | 3 | 3 |
| 3 | 2 | 2 | 7 | 9 | 8 |
| 4 | 6 | 7 | 31 | 38 | 30 |
| 5 | 21 | 25 | 138 | 163 | 117 |
| 6 | 78 | 114 | 753 | 867 | 522 |
| 7 | 353 | 746 | 4,939 | 5,685 | 2,917 |
| 8 | 1,929 | 12,903 | 57,856 | 70,758 | 27,899 |
| Total | 2,391 | 13,799 | 63,726 | 77,524 | 31,497 |
DFT optimization: All structures were geometry-optimized at the PBE0-D4/def2-TZVP level of theory. 66.5% of structures converged after a single optimization; the remainder required one or two additional passes. 59% of CHX8-enum structures underwent $\sigma$-framework rearrangements during optimization and were classified as unstable. Rearranged structures were identified by comparing input and output InChI strings. Analysis confirmed that all rearrangement products (closed-shell, zwitterionic, or carbene species) were already present in the enumeration, so no new compounds were missed.
Relative Strain Energy as a Synthesizability Proxy
A universal RSE metric, referenced to cyclohexane (zero strain), was developed and assigned to every molecule. The RSE for a molecule of interest (subscript $n$) relative to a reference structure (subscript $r$) is:
$$ \text{RSE} = E_{n} - E_{r} - (c_{n} - c_{r}),E_{\text{CH}_2} + E_{\text{unsat}} $$
where $E_{n}$ and $E_{r}$ are Gibbs energies, $c_{n}$ and $c_{r}$ are carbon counts, $E_{\text{CH}_2}$ is the average energy cost of adding an unstrained CH$_2$ unit, computed from the Gibbs energy differences between consecutive linear alkanes (ethane through octane, six increments), and $E_{\text{unsat}}$ corrects for differences in unsaturation:
$$ E_{\text{unsat}} = (r_{n} - r_{r}),E_{\text{ring}} + (d_{n} - d_{r}),E_{\text{double}} + (t_{n} - t_{r}),E_{\text{triple}} $$
$E_{\text{double}}$ and $E_{\text{triple}}$ are each derived from internal transformations between the second and third carbon of linear chains, averaged over four chain lengths (n-butane through n-octane). Initial attempts using terminal unsaturations systematically underestimated RSE for structures containing double and triple bonds. $E_{\text{ring}}$ is derived separately using the Dudev-Lim homolytic bond dissociation approach:
$$ E_{\text{ring}} = 2E_{\text{C-H}} - E_{\text{C-C}} $$
where the individual bond energies are obtained from ethane:
$$ E_{\text{C-H}} = E_{\text{ethane}} - E_{\text{ethyl radical}}, \quad E_{\text{C-C}} = E_{\text{ethane}} - 2E_{\text{methyl radical}} $$
The highest-RSE molecule with synthetic precedent (a C6 structure detected by atomic force microscopy on a metal surface) has an RSE of 201.4 kcal/mol. Using this as a threshold, over 90% of the novel structures in CHX8 should be considered synthetically accessible in principle.
Notable reference points on the RSE scale:
- Cyclopropane: 27.5 kcal/mol
- Tetrahedrane: 140.1 kcal/mol (substituted variants synthesized, unsubstituted not yet)
- Cubane: 157.4 kcal/mol (synthesized)
- Highest synthesized: 201.4 kcal/mol (C6 structure on metal surface)
Key Findings on Strained Motifs
The exhaustive enumeration enables systematic analysis of structural classes previously excluded:
- Trans-cycloalkenes: All trans-cycloalkenes in 6-membered rings or larger should be synthetically feasible. The stability of multi-trans systems depends on the relative position of double bonds: parallel trans-double bonds in a ring can undergo thermally accessible 4$\pi$-electrocyclisation, while non-parallel arrangements may be conformationally locked and stable.
- Cyclic alkynes and allenes: 37% of the CHX8 dataset consists of cyclic alkynes or allenes. All cyclic alkynes except cyclopropyne, and all cyclic allenes, should be synthesizable (in singlet or triplet states), with RSE values below cubane.
- Trans-fused rings: All but [3,3]- and [3,4]-unsubstituted trans-fused rings should be accessible. The proposed lower limit for trans-ring junctions is either (i) a 3-membered ring trans-fused to a ring of five or more atoms, or (ii) a 4-membered ring trans-fused to another 4-membered ring.
- Anti-Bredt structures: CHX8 contains seven hydrocarbon skeletons with a bridging section, yielding fourteen possible anti-Bredt (bridgehead-unsaturated) derivatives. Of these, thirteen are stable under DFT optimization, and over 200 substituted anti-Bredt structures are present in the dataset. All stable anti-Bredt structures have RSE values below cubane. Stability is classified using Fawcett’s S parameter (the number of non-bridgehead ring atoms): CHX8 finds structures with S $\geq$ 4 are stable to optimization, consistent with recent experimental work that has accessed anti-Bredt intermediates at S values as low as 4.
Comparison to Existing Databases
- vs. GDB-13: CHX8 contains 31,497 C1-C8 hydrocarbons vs. 1,966 in GDB-13 (16x more). For C8 hydrocarbons specifically, GDB-13 has more coverage than GDB-17 (1,966 vs. 1,121). All GDB-13 hydrocarbons appear in CHX8-enum, though some were unstable to DFT optimization.
- vs. VQM24: For C1-C5 hydrocarbons, VQM24 contains 123 closed-shell isomers vs. 154 in CHX8 (14-25% more). Many missing structures in VQM24 are diastereoisomers not generated by the SURGE process.
- vs. PubChem: Less than 44% of CHX8 structures appear in PubChem
- vs. Reaxys: Only 25% of CHX7 (up to 7 carbons) structures are commercially available
Reproducibility Details
The enumeration pipeline uses open-source tools: GENG from the nauty package for graph generation, RDKit for molecular manipulation and InChI canonicalization, and OpenBabel for 3D coordinate generation (MMFF94). DFT calculations used the PBE0-D4/def2-TZVP level of theory via the ORCA quantum chemistry package. The paper does not report total compute time or hardware specifications.
| Artifact | Type | License | Notes |
|---|---|---|---|
| CHX8 Dataset (Nottingham Repository) | Dataset | Unknown | All optimized 3D structures, optimization/frequency output files, organized into CHX7, CHX8-sat, and CHX8-unsat subsets |
Missing components for full reproduction: No source code for the enumeration or unsaturation-introduction scripts is released. The RSE calculation scripts and DFT input templates are not provided. Hardware/compute requirements are not reported.
Reproducibility status: Partially Reproducible. The dataset itself is deposited, but the enumeration and analysis code is not released.
Paper Information
- Preprint: ChemRxiv, January 2, 2026
@article{harman2026complete,
title={Complete Computational Exploration of Eight-Carbon Hydrocarbon Chemical Space},
author={Harman, Stephen J. and Ermanis, Kristaps},
journal={ChemRxiv},
year={2026},
doi={10.26434/chemrxiv-2026-qjr5r}
}