Key Contribution
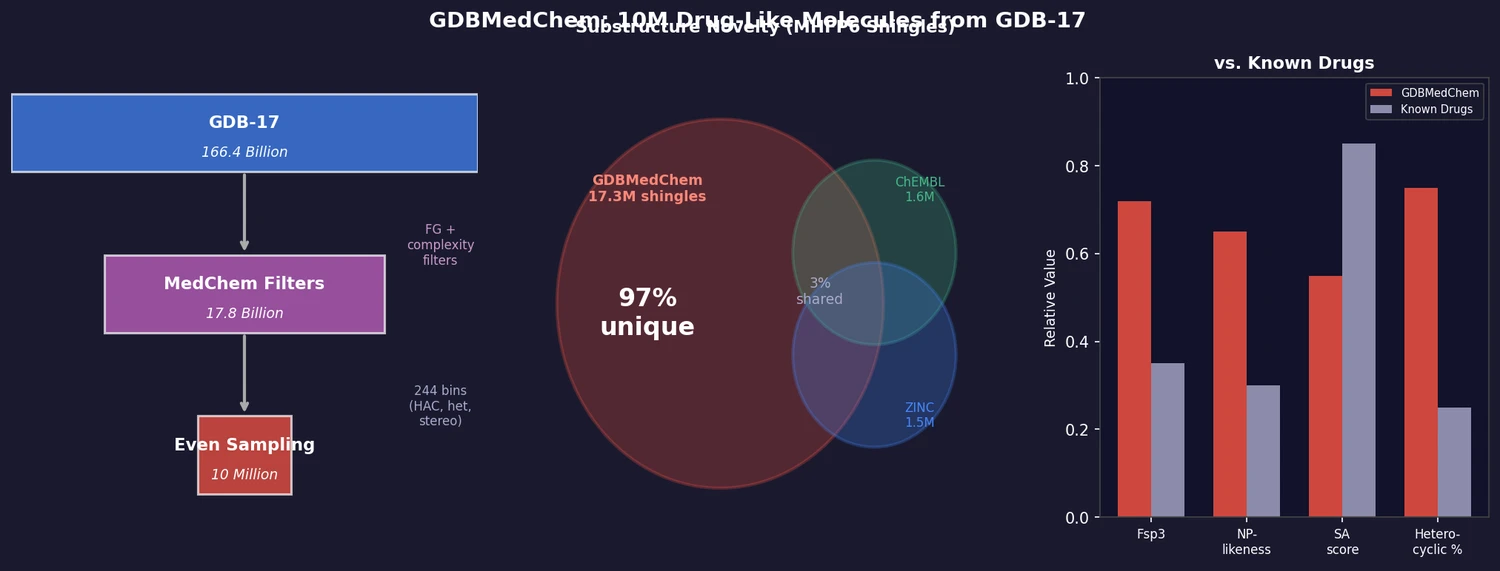
GDBMedChem is a 10 million molecule subset of GDB-17 selected using medicinal chemistry criteria rather than the fragment-likeness rules used for FDB-17. The resulting database has reduced complexity and better synthetic accessibility than the full GDB-17, while retaining higher Fsp3 carbon fraction and natural product likeness compared to known drugs. Critically, 97% of its MHFP6 substructure shingles are absent from DrugBank, ChEMBL, and ZINC, making it an unprecedented source of structural diversity for drug design.
Overview
GDB-17 enumerates 166.4 billion molecules following chemical stability and synthetic feasibility rules, but does not consider medicinal chemistry criteria such as acceptable functional group types, overall structural complexity, or drug-likeness. GDBMedChem addresses this gap with a different filtering philosophy than FDB-17: instead of enforcing fragment-likeness (rotatable bond limits, small size), it applies medicinal chemistry-inspired rules that allow larger, more flexible molecules while excluding problematic functional groups and overly complex scaffolds.
Assembly Pipeline
Stage 1: Medicinal chemistry filters (166.4B to 17.8B, ~9.4x reduction)
Three categories of filters, each benchmarked against ChEMBL, DrugBank, and UNPD (natural products) to ensure low elimination of known bioactives:
| Category | Key Filters | GDB-17 Eliminated |
|---|---|---|
| Functional groups | No amidines, imidates, aldehydes, aziridines, epoxides; no Br/I; no Cl/F on heterocycles; max 1 nitrile/alkyne/sulfone; max 2 ethers/amides/esters | 53% |
| Structural complexity | Max 18 avalon fingerprint density; max 1 cyclic tetravalent node; max 4 stereocenters; max 3 bonds in fused ring systems; max 3 rings | 62% |
| Polarity | Heteroatom-to-carbon ratio max 0.7 | 6% |
| Combined | All filters together | 86% |
These filters eliminate 86% of GDB-17 but only 36% of ChEMBL molecules and 50% of DrugBank drugs (the higher DrugBank rate is driven mainly by the heteroatom-to-carbon ratio filter removing highly polar drugs with negative clogP values).
Of the 21 filters, 16 are implemented as SMARTS queries and 5 (stereocenters, ring count, avalon density, heteroatom-to-carbon ratio, largest aromatic ring size) use other RDKit functions. Filters were applied progressively (simplest first), not in the order listed above. The benchmarking percentages for ChEMBL and DrugBank refer to ChEMBL 22 and DrugBank 5.011 molecules with HAC ≤ 17.
Stage 2: Even sampling (17.8B to 10M)
The 17,804,900,000 molecules in the filtered set are binned into 425 possible triplet combinations of HAC (1-17), heteroatoms (≤1, 2, 3, 4, ≥5), and stereocenters (0, 1, 2, 3, 4). Of these, 181 bins are unoccupied, leaving 244 bins. PySpark’s sampleBy function performs stratified sampling without replacement, using a round-robin allocation that increments each bin’s quota by one until the total reaches 10M. The resulting distribution is uniform except in low-HAC bins (HAC ≤ 10) where all available molecules are taken.
Comparison with FDB-17
GDBMedChem and FDB-17 are both 10M-molecule subsets of GDB-17 but take fundamentally different approaches:
| Property | GDBMedChem | FDB-17 |
|---|---|---|
| Parent set | 17.8B (medchem filters) | 4.6B (fragment filters) |
| Overlap | 480M molecules shared between parent sets | |
| Rotatable bonds | Similar to known drugs | Restricted to max 3 (fragment-like) |
| Key difference | Drug-like flexibility, medchem FG rules | Fragment-like rigidity, strict FG removal |
Both databases retain GDB-17’s characteristic high Fsp3 fraction and 3D molecular shape diversity compared to predominantly planar known molecules.
Substructure Novelty
MHFP6 (MinHash fingerprint with diameter 6) shingle analysis reveals striking structural novelty:
| Database | Molecules | Unique Shingles | Unique to Database |
|---|---|---|---|
| GDBMedChem | 10M | 17.3M | 97% |
| ChEMBL | 1.4M | 1.6M | 57% |
| ZINC | 15M | 1.5M | 53% |
| DrugBank | 8.3k | 82k | 12% |
GDBMedChem contains 17.3 million unique shingles, roughly 10x more than the 15 million-molecule ZINC database, with 97% appearing in no other database. The cumulative unique shingle count grows faster and more steadily with database size for GDBMedChem than for known molecule databases, reflecting greater internal diversity. Among the most frequent shingles, oxygen-containing saturated or singly unsaturated substructures dominate GDBMedChem, in contrast to aromatic and nitrogen heterocycles in ZINC.
Property Profiles
Compared to known drugs (DrugBank17, ChEMBL17):
- Synthetic accessibility: Slightly better than GDB-17 due to complexity filters, but still lower than known molecules
- Natural product likeness: Significantly higher than drugs, approaching natural products (UNPD17)
- Fsp3 fraction: Higher than drugs, reflecting more 3D-shaped molecules
- Compound categories: Much higher fraction of heterocyclic molecules, much lower fraction of aromatic molecules (a consequence of combinatorial enumeration favoring heteroatom-in-ring combinations)
Strengths & Limitations
Strengths:
- 97% structurally novel substructures provide unprecedented diversity for drug design
- Medicinal chemistry filters retain drug-relevant functional group patterns
- Even sampling corrects GDB-17’s combinatorial bias toward large, complex molecules
- Higher Fsp3 and natural product likeness compared to known drugs
- Available with interactive 3D visualization, MQN/MHFP6 similarity search, and download
Limitations:
- Synthetic accessibility scores remain lower than for known molecules
- Excludes Br, I, and Cl/F on heterocycles, which are common in medicinal chemistry
- Random sampling means specific molecules of interest from the 17.8B parent set may be absent
- Overlap with FDB-17 is limited (different filtering philosophies), so both databases complement rather than replace each other
Technical Notes
Molecule Preprocessing
Before filtering, each molecule undergoes: counter-ion removal, largest-fragment retention, conversion to non-chiral SMILES, valence-error checking, and protonation at pH 7.4 (using ChemAxon JChem). Duplicates are removed by canonical SMILES comparison within each database.
Reference Databases
The comparison databases used specific versions: ChEMBL 22 (1.4M compounds with HAC ≤ 50; 105,423 with HAC ≤ 17), DrugBank 5.011 (8,299 approved/experimental drugs with HAC ≤ 50; 2,284 with HAC ≤ 17), UNPD (20,302 natural products with HAC ≤ 17), and ZINC 12 (15M commercially available compounds).
MHFP6 Shingle Computation
Shingles were computed using the mhfp Python package (also on PyPI), specifically the shingling_from_smiles function from the MHFPEncoder class. Each shingle represents an extended-connectivity substructure around an atom with a diameter of up to 6 bonds, plus all ring structures, encoded as rooted SMILES strings.
Avalon Fingerprint Density
The avalon fingerprint density, used as the overall structural complexity filter (max 18), is defined as the number of on-bits in the avalon fingerprint scaled to the heavy atom count.
Reproducibility Details
| Artifact | Type | License | Notes |
|---|---|---|---|
| GDBMedChem download | Dataset | Non-commercial (no patents, no redistribution) | 10M molecules in SMILES format |
| GDB web tools | Other | Unknown | 3D visualization, MQN/MHFP6 similarity search |
mhfp Python package | Code | MIT | MHFP6 fingerprint and shingle computation |
| PCA visualization tools | Code | Unknown | MQN-to-3D PCA projection preprocessing |
Status: Partially Reproducible. The dataset itself is publicly available for download, and the paper describes the filtering and sampling pipeline in detail (RDKit 2017_09_03, PySpark 2.3.2, 98-node cluster with 252 GB RAM). The mhfp package for shingle analysis is open-source. However, no standalone filtering/sampling code is released: reproducing the pipeline from scratch requires reimplementing the 16 SMARTS filters and 5 RDKit-based filters, plus the PySpark stratified sampling procedure. The molecule preprocessing step also depends on ChemAxon JChem (commercial) for pH 7.4 protonation and MQN calculation.
The paper is published in the closed-access journal Molecular Informatics. An open-access preprint is available on ChemRxiv.
Citation
@article{awale2019medicinal,
title={Medicinal Chemistry Aware Database GDBMedChem},
author={Awale, Mahendra and Sirockin, Finton and Stiefl, Nikolaus and Reymond, Jean-Louis},
journal={Molecular Informatics},
volume={38},
number={8-9},
pages={e1900031},
year={2019},
publisher={Wiley},
doi={10.1002/minf.201900031}
}