Key Contribution
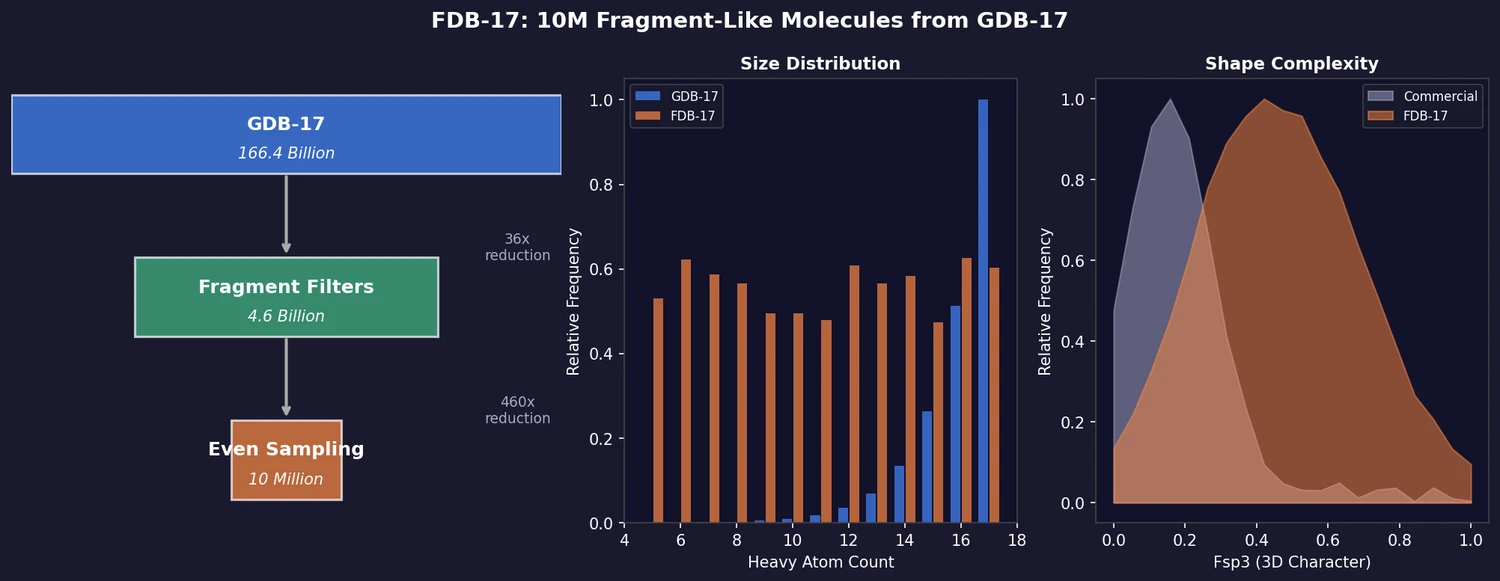
FDB-17 is a curated subset of 10 million fragment-like molecules extracted from the 166.4 billion molecules in GDB-17. It corrects the combinatorial bias of exhaustive enumeration (which overwhelmingly produces large, complex molecules) by evenly sampling across molecular size, polarity, and stereochemical complexity. The result is a database sized for practical virtual screening tools while retaining GDB-17’s distinctive 3D molecular shape diversity.
Overview
GDB-17 exhaustively enumerates molecules up to 17 heavy atoms, but the combinatorial explosion means the database is dominated by the largest, most functionalized, and stereochemically most complex entries. This makes it impractical for most virtual screening workflows and poorly suited for identifying simple, synthetically accessible fragments. FDB-17 addresses both problems through a two-stage reduction.
Assembly Pipeline
Stage 1: Fragment-likeness filters (166.4B to 4.6B, 36x reduction)
Criteria limiting structural and functional group complexity:
| Category | Constraints |
|---|---|
| Scaffolds | Max 3 rings, max 2 small (3/4-membered) rings, max 2 quaternary centers, max 4 stereocenters, max 3 rotatable bonds |
| FG density | Max 5 N+O atoms, max 1 positive/negative charge at neutral pH, max 3 HBA, max 2 HBD |
| Excluded groups | Aldehydes, epoxides, aziridines, carbonates, imidates, nitro groups, aromatic rings >6 atoms, ≤ 1 cyano group |
| Removed elements | Non-aromatic C=C, C triple bonds, halogens (approximated by saturated C-C and methyl) |
Stage 2: Even sampling (4.6B to 10M, 460x reduction)
The 4.6B fragment subset is binned into 175 cells defined by value triplets of (HAC, heteroatoms, stereocenters):
| Dimension | Bin values |
|---|---|
| HAC | ≤11, 12, 13, 14, 15, 16, 17 (7 bins) |
| Heteroatoms (N+O+S) | ≤1, 2, 3, 4, ≥5 (5 bins) |
| Stereocenters | 0, 1, 2, 3, 4 (5 bins) |
Individual bins ranged from 3,359 to 446,322,188 molecules, reflecting the extreme combinatorial skew toward large, complex structures. Bins with ≤70,000 molecules are taken entirely; larger bins are randomly sampled to approximately 60,000 molecules each. The filtering was implemented in Java using ChemAxon’s JChem libraries and executed on a 500-node cluster in 10,000 CPU hours. The resulting even distribution across molecular size, polarity, and complexity replaces the exponentially skewed distribution of the parent database.
Property Profiles vs. Commercial Fragments
FDB-17 was compared against 40,986 commercial fragments collected from 8 vendors (AnalytiCon, ChemBridge, Enamine, FRAGMENTA, BIONET, LifeChemical, Maybridge, Vitas) and filtered by Congreve’s rule of three (mass ≤300, HBA ≤3, HBD ≤3, logP ≤3, RBC ≤3, PSA ≤60). Only 31% (12,847) of these commercial fragments appeared in the 4.6B fragment subset at all, due to functional groups absent from GDB-17 (halogens, thiols, azides, thioethers). Of those, only 6.7% (2,740) appeared in FDB-17 due to the random sampling step.
Key differences:
- Size and polarity: FDB-17’s even sampling produces distributions comparable to commercial fragments, unlike the parent GDB-17 which peaks sharply at HAC = 17
- Compound categories: Half are heteroaromatic in both sets, but FDB-17’s second half is predominantly heterocyclic vs. aromatic for commercial fragments
- 3D character: FDB-17 retains GDB-17’s coverage of the full PMI (principal moments of inertia) shape triangle, with a frequency peak at center-left (PMI computed from single low-energy CORINA conformers). Commercial fragments are predominantly planar. FDB-17 has significantly higher Fsp3 values
- Ring count: Fragment subsets of GDB-17 are enriched in 2- and 3-ring molecules (a consequence of the rotatable bond limit, which constrains monocyclic molecules more than polycyclic ones)
Virtual Screening Validation
Nearest-neighbor searches were performed using two fingerprint spaces: MQN (42-dimensional molecular quantum numbers counting atoms, bonds, polarity, and topology) and Xfp (55-dimensional extended pharmacophore fingerprint capturing shape and pharmacophore features). Four fragment-like drugs were used as queries: fencamfamine, gabapentin, rimantadine, and levetiracetam. For each drug, 10,000 nearest neighbors were retrieved and scored by 3D-shape similarity using ROCS (Rapid Overlay of Chemical Structures). 3D conformers were generated with OMEGA (all possible stereoisomers, keeping the highest-scoring one). Molecules with ROCS Tanimoto Combo > 1.4 were considered virtual hits.
FDB-17 delivered comparable numbers of virtual hits to the full 4.6B fragment subset and the entire GDB-17, despite being 460x and 16,640x smaller respectively. Both close analogs (high substructure similarity, Tsfp > 0.7) and scaffold-hopping compounds (low substructure similarity but high shape similarity) were identified. Random sampling from FDB-17 and searches in the 41k commercial fragment set returned far fewer hits.
Strengths & Limitations
Strengths:
- Manageable size (10M) compatible with docking and 3D-shape virtual screening tools
- Even coverage of molecular size, polarity, and complexity avoids combinatorial bias
- High 3D shape diversity compared to predominantly flat commercial fragment libraries
- Available with interactive visualization (MQN/SMIfp-mapplet) and web-based nearest neighbor search
Limitations:
- Only the 10M FDB-17 is released, not the 4.6B fragment-filtered intermediate. Practitioners who want a different sampling strategy or the full fragment subset cannot access it
- Random sampling means specific molecules of interest from the 4.6B subset may be absent
- Excludes halogens, non-aromatic unsaturations, and several functional group classes present in commercial fragments
- Only 6.7% overlap with commercial fragments limits direct comparison
- Still derived from GDB-17’s enumeration rules, so molecules outside those rules (e.g., containing metals or larger rings) are excluded
Reproducibility Details
FDB-17 is publicly available for download from the GDB project page as a single SMILES file (62.2 MB), hosted on Zenodo. Interactive visualization via the MQN/SMIfp-mapplet and web-based nearest neighbor search tools are also accessible through the same site. The multi-fingerprint browser supports nearest-neighbor search across six fingerprints: MQN (42D), SMIfp (34D), APfp (21D), Xfp (55D), Sfp (1024-bit Daylight-type), and ECfp4 (1024-bit circular). The filtering code was written in Java using JChem libraries (ChemAxon) and executed on a 500-node cluster in 10,000 CPU hours. The filtering code itself is not publicly released. Virtual screening additionally requires OMEGA (conformer generation) and ROCS (3D-shape scoring), both commercial tools from OpenEye.
| Artifact | Type | License | Notes |
|---|---|---|---|
| FDB-17 SMILES | Dataset | Custom (no patents, no redistribution) | 10M fragment-like molecules from GDB-17 |
| MQN/SMIfp-mapplet | Other | Web tool | Interactive PCA visualization on 1000x1000 grids |
| Multi-fingerprint browser | Other | Web tool | Nearest neighbor search across 6 fingerprints (MQN, SMIfp, APfp, Xfp, Sfp, ECfp4) |
Reproducibility status: Partially Reproducible. The 10M FDB-17 is freely downloadable, but the 4.6B fragment-filtered intermediate is not released. The filtering criteria are fully documented, but the Java filtering code is not released and depends on proprietary ChemAxon libraries. Reproducing the virtual screening experiments requires commercial tools (OMEGA, ROCS from OpenEye; CORINA for PMI analysis).
Citation
@article{visini2017fragment,
title={Fragment Database FDB-17},
author={Visini, Ricardo and Awale, Mahendra and Reymond, Jean-Louis},
journal={Journal of Chemical Information and Modeling},
volume={57},
number={4},
pages={700--709},
year={2017},
publisher={American Chemical Society},
doi={10.1021/acs.jcim.7b00020}
}