Research notes spanning three related areas: protein folding (Levinthal’s paradox, energy landscape theory, inverse folding with Potts models, dynamics-informed drug design), the 3D point set alignment algorithms that underpin structural comparison (Kabsch, Horn, Arun, Umeyama), and computational metabolic pathway design (graph-grammar chemical space expansion with ILP flow queries).
Protein Folding
| Year | Paper | Key Idea |
|---|---|---|
| 1969 | How to Fold Graciously | Levinthal’s paradox: random search cannot explain protein folding |
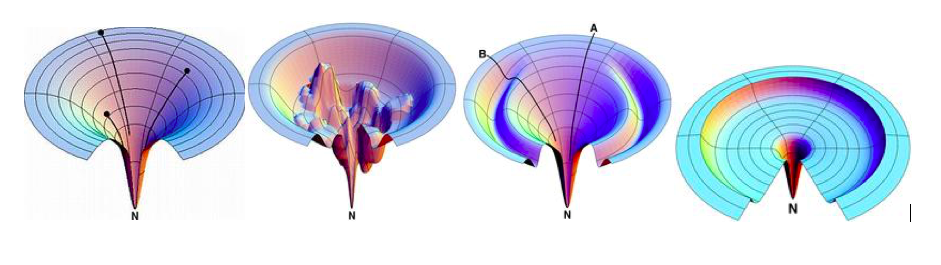
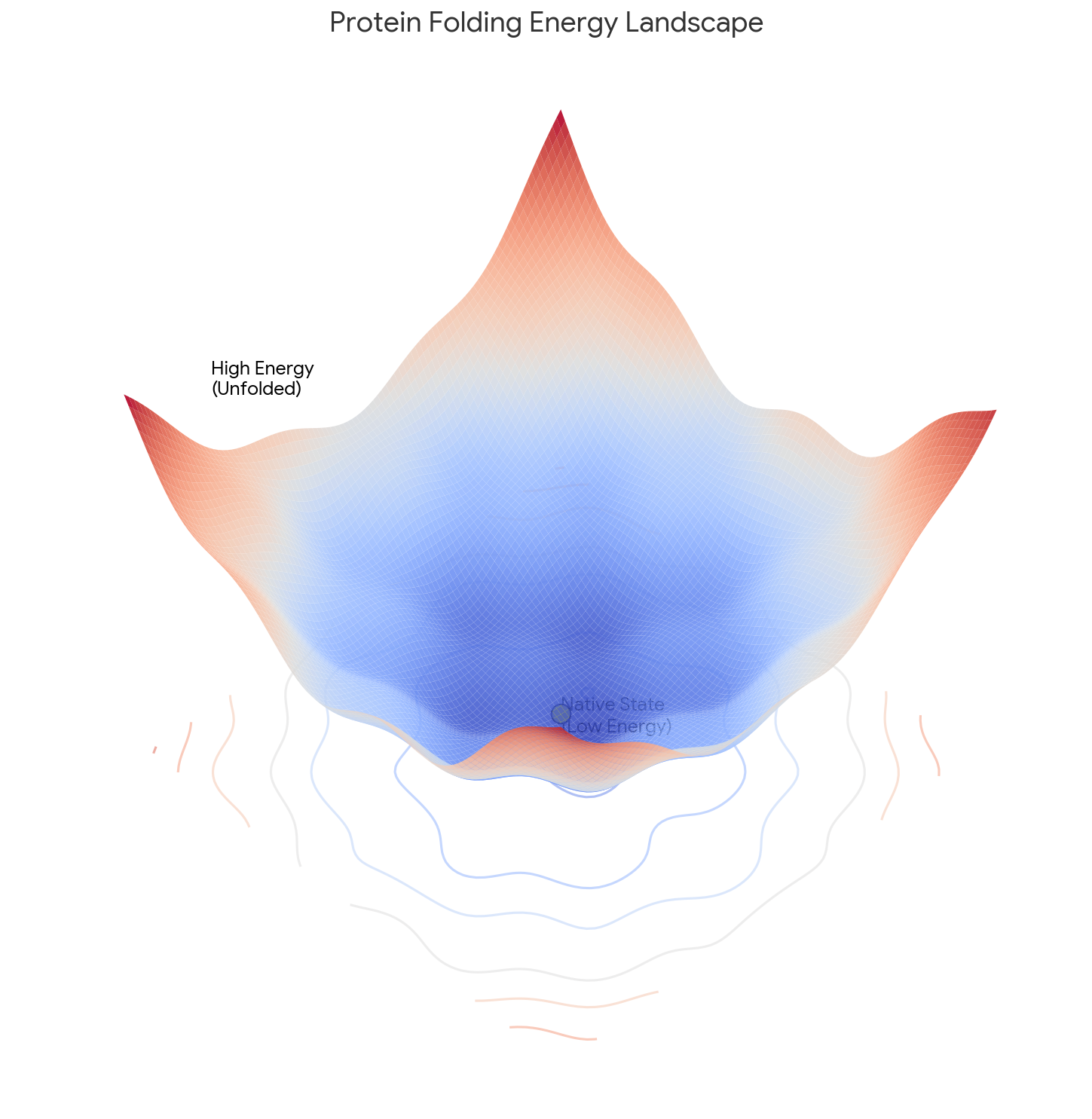
| 1995 | Funnels, Pathways, and Energy Landscapes | Energy landscape theory with folding funnels resolves Levinthal’s paradox |
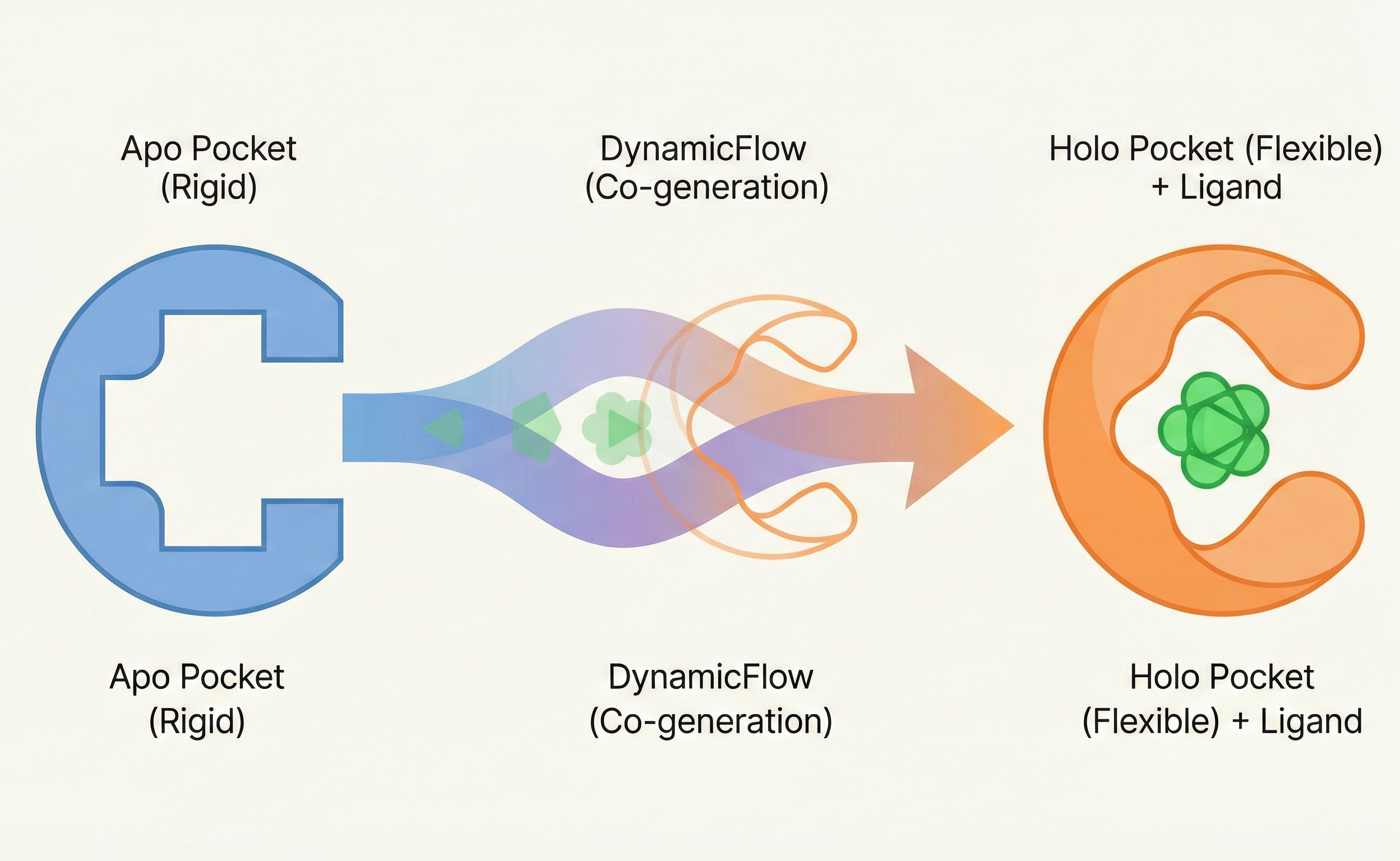
| 2025 | DynamicFlow | Flow matching co-generating ligands and flexible protein pockets |
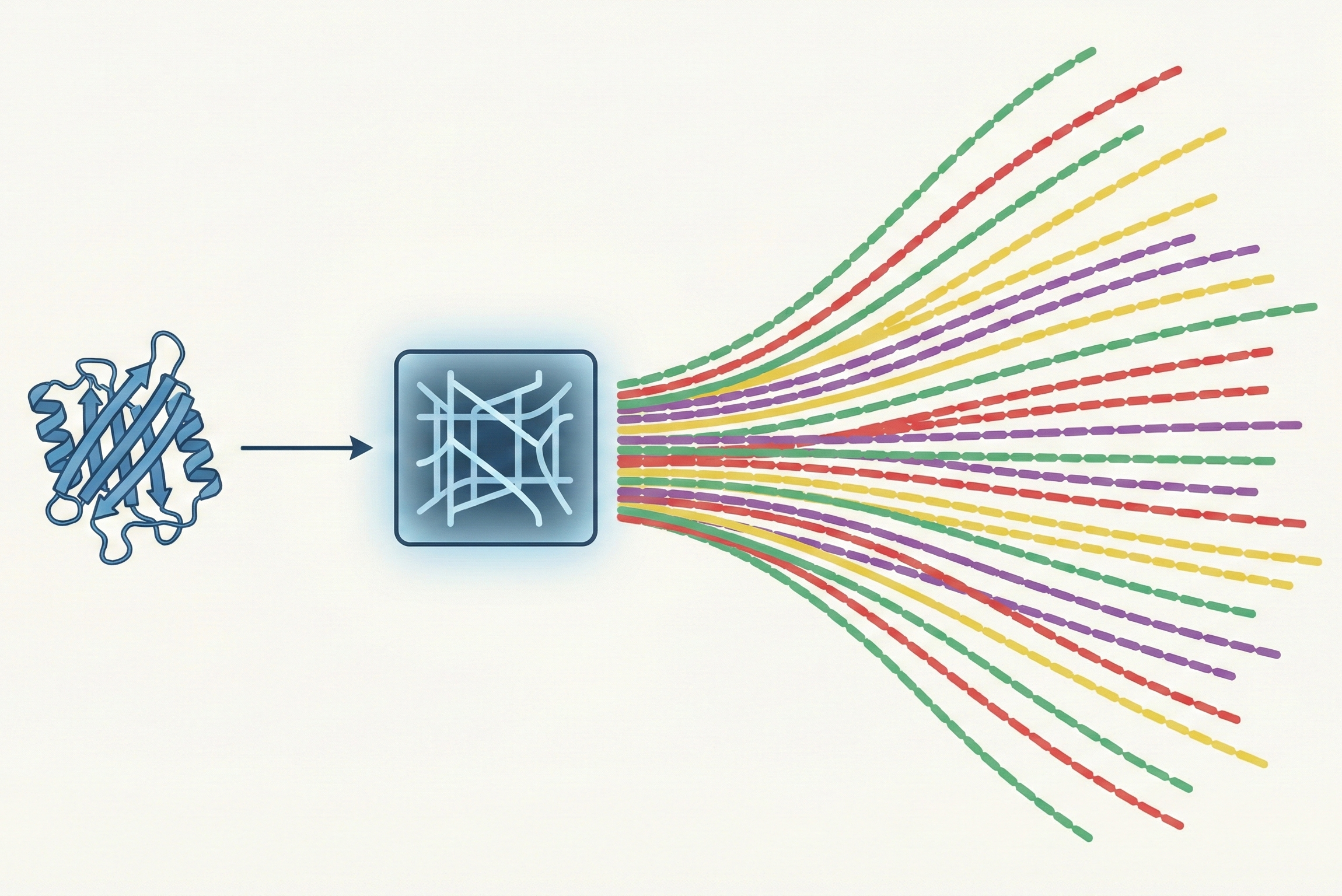
| 2025 | InvMSAFold | Potts model parameter prediction for fast diverse inverse folding |
Point Set Alignment
| Year | Paper | Key Idea |
|---|---|---|
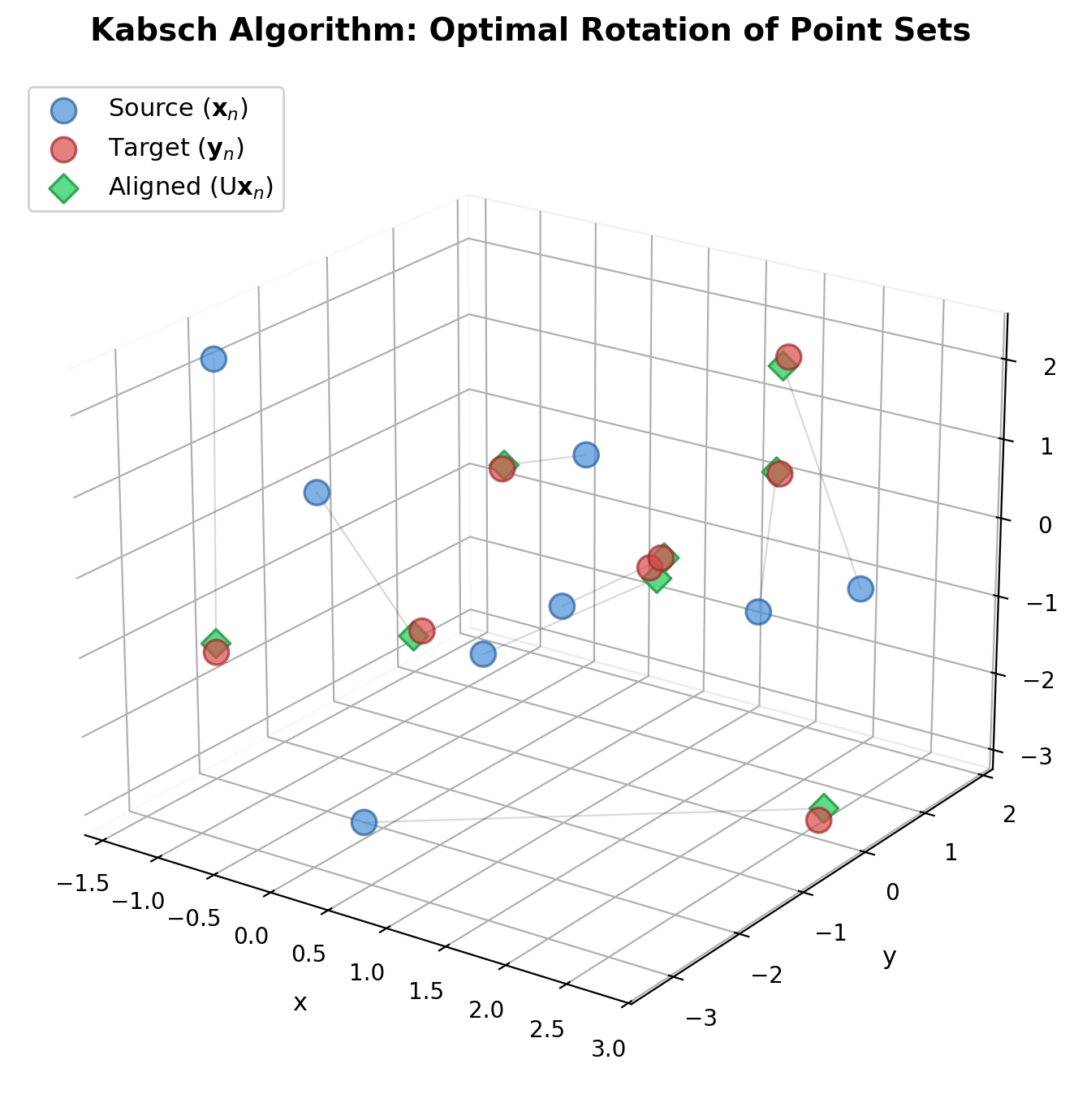
| 1976 | Kabsch Algorithm | Closed-form optimal rotation via cross-covariance eigendecomposition |
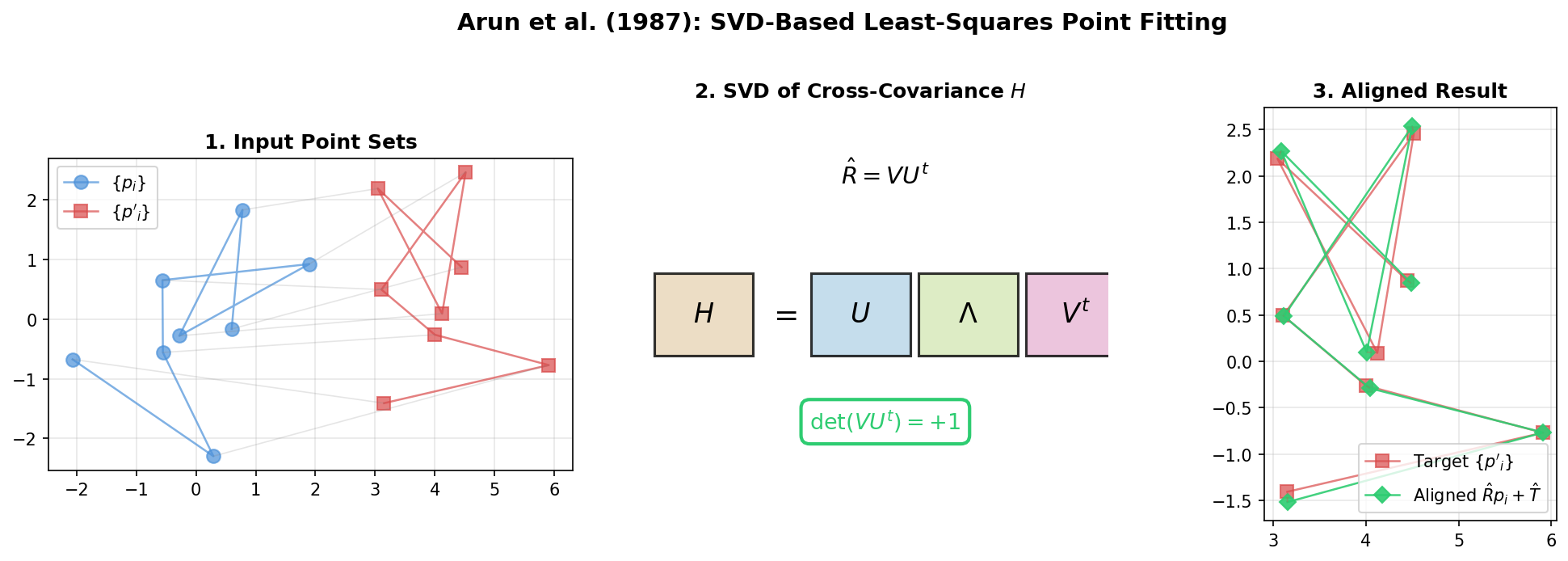
| 1987 | Arun et al.: SVD-Based Point Fitting | SVD-based rotation and translation for 3D point sets |
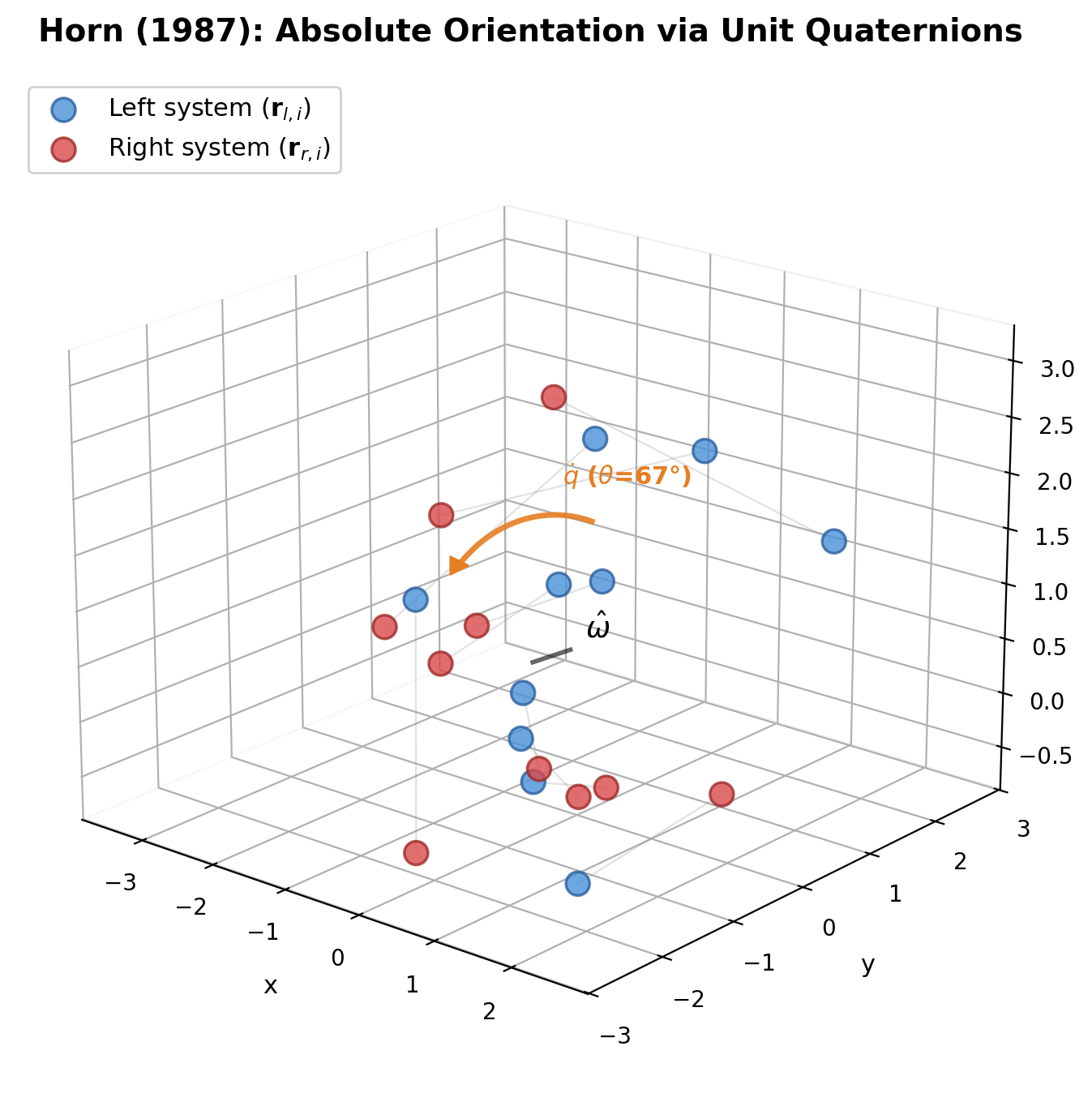
| 1987 | Horn’s Quaternion Method | Absolute orientation via unit quaternion eigenvector solution |
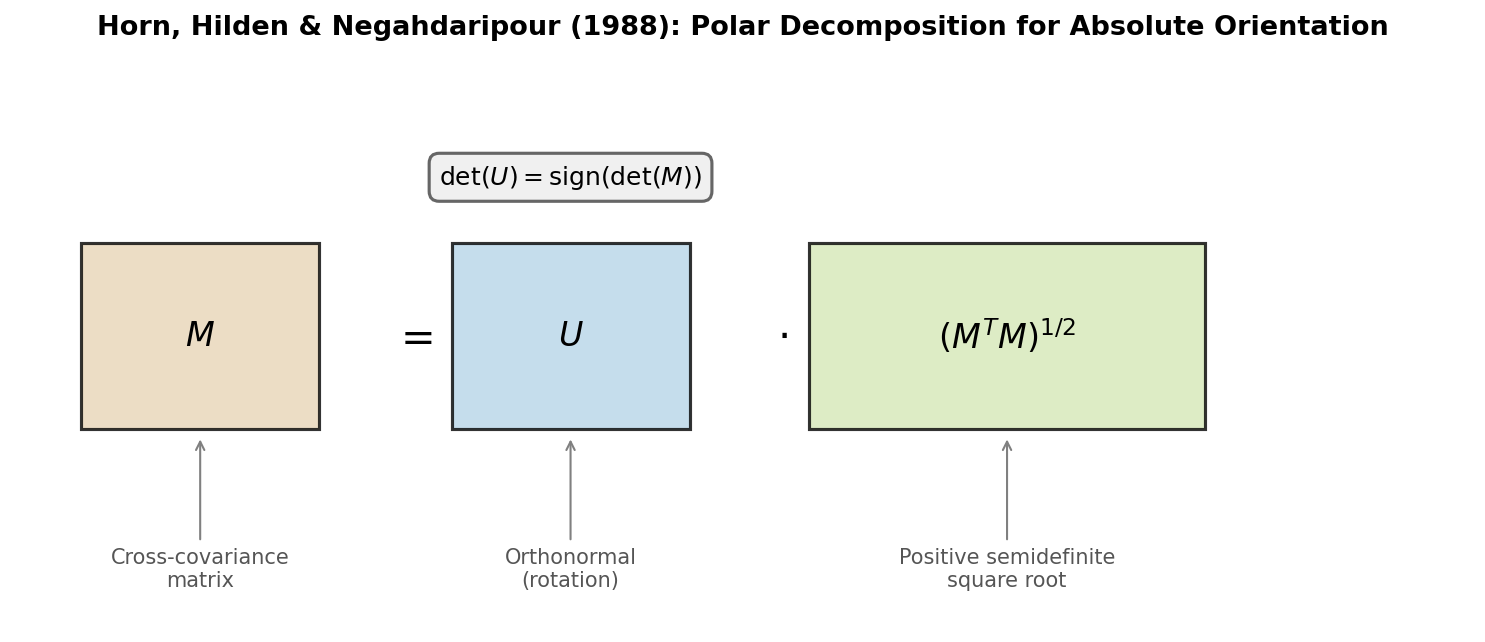
| 1988 | Horn et al.: Orthonormal Matrices | Polar decomposition alternative to quaternion-based alignment |
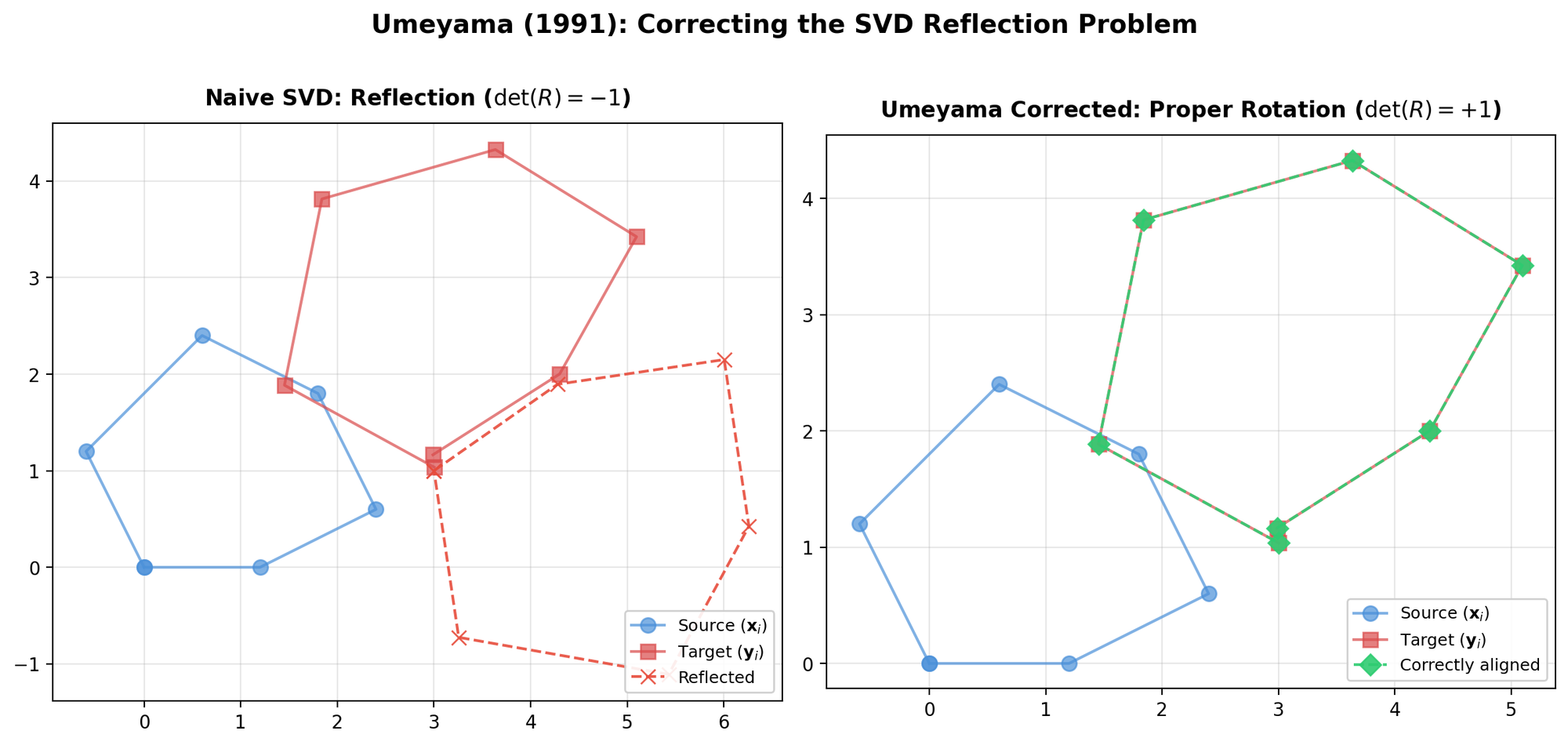
| 1991 | Umeyama’s Method | Corrected SVD alignment that always produces proper rotations |
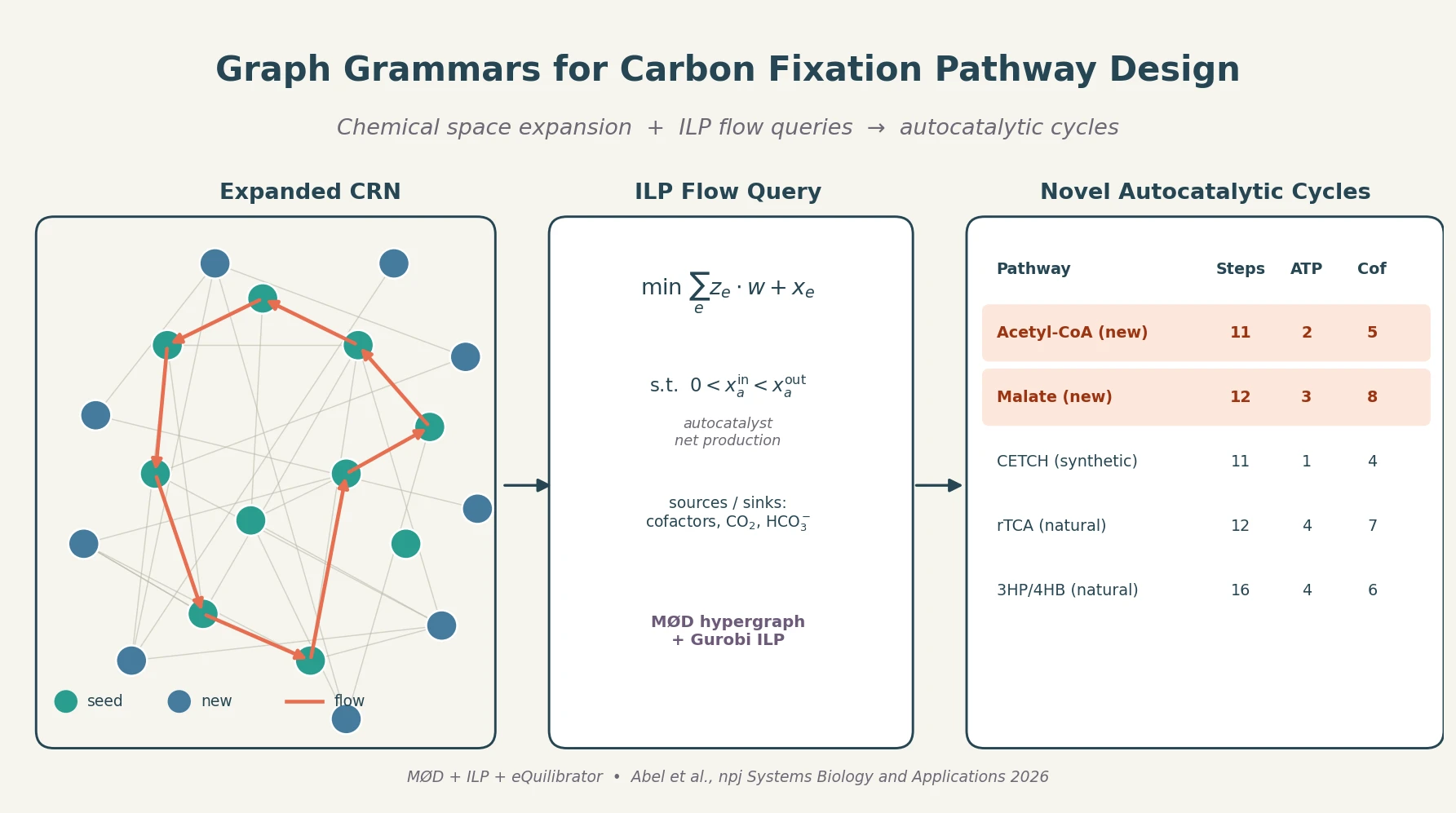
Metabolic Pathway Design
| Year | Paper | Key Idea |
|---|---|---|
| 2026 | Graph Grammar and ILP for Carbon Fixation Pathways | MØD chemical space expansion plus ILP flow queries discover novel autocatalytic carbon fixation cycles |