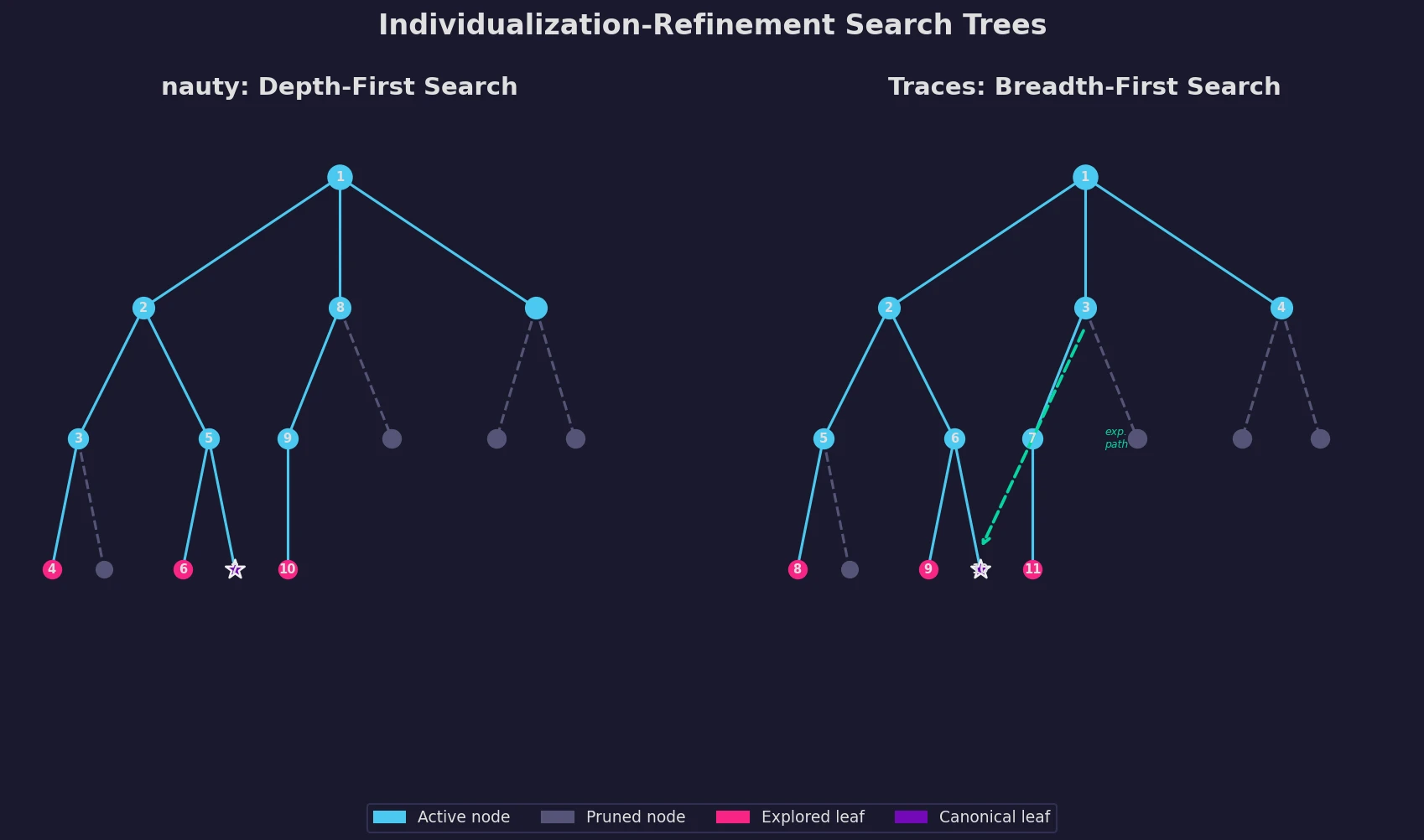

nauty and Traces: Graph Isomorphism Algorithms
An updated description of nauty and introduction of Traces, two programs for graph isomorphism testing and canonical labeling using the individualization-refinement paradigm.

An updated description of nauty and introduction of Traces, two programs for graph isomorphism testing and canonical labeling using the individualization-refinement paradigm.
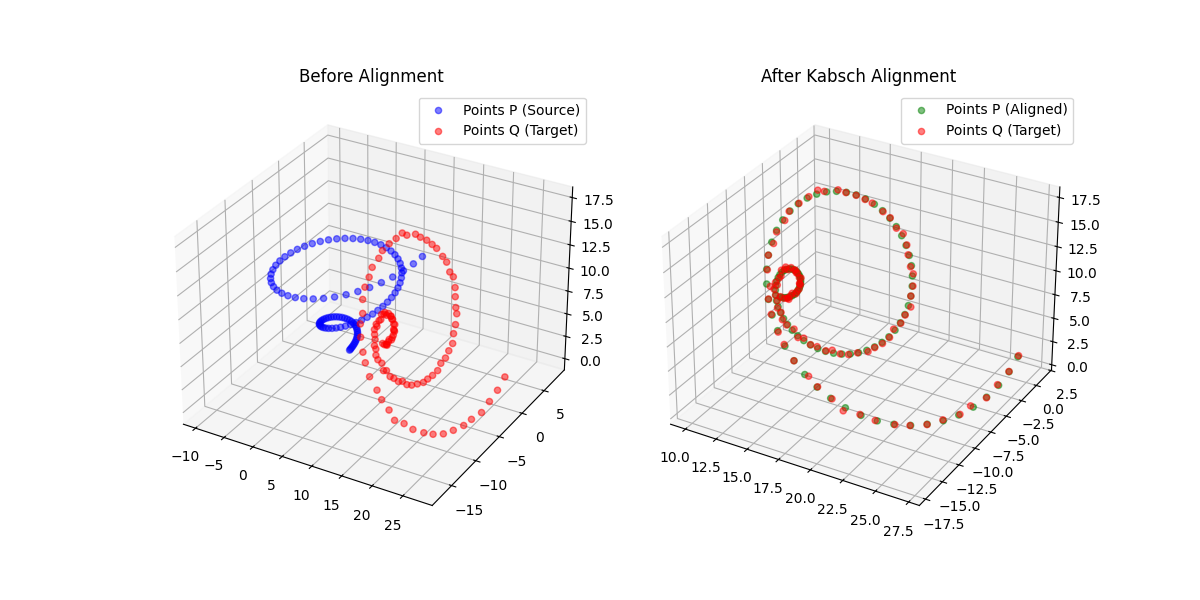
A differentiable point-set alignment library implementing N-dimensional Kabsch, Horn quaternion, and Umeyama scaling algorithms with per-point weights, batch dimensions, and custom autograd across NumPy, PyTorch, JAX, TensorFlow, and MLX.
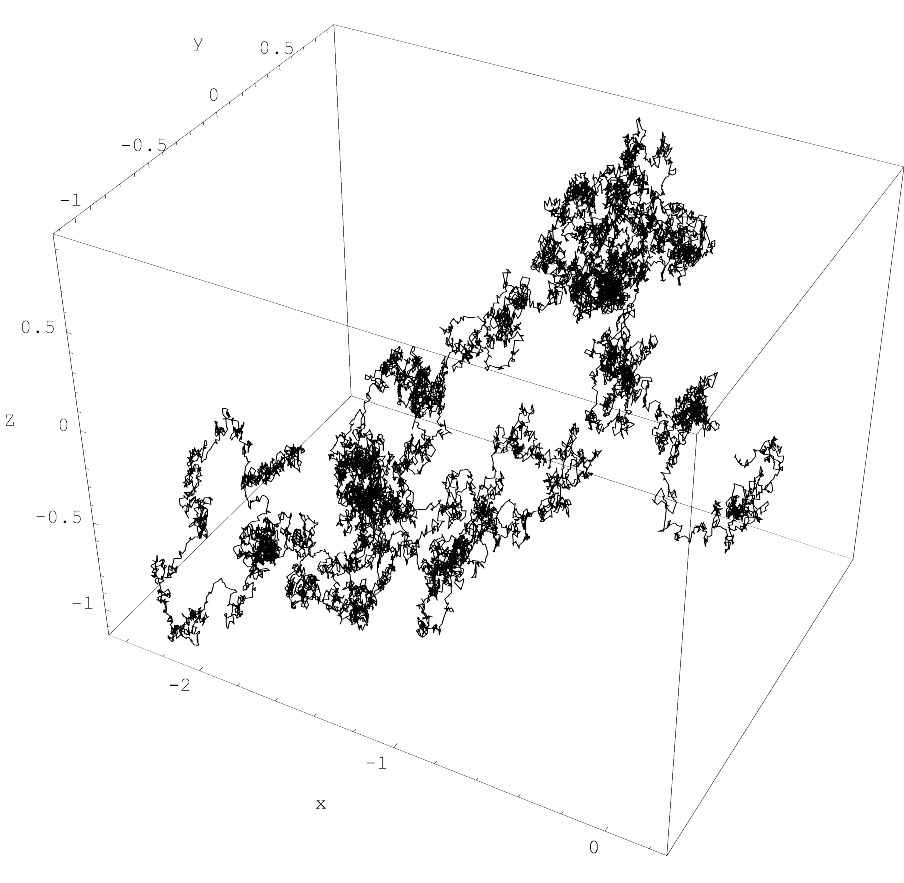
Proposes the Hyperbolic Algorithm for Euclidean field theory simulations. By adding a second-order fictitious time derivative to the Langevin equation, the method reduces systematic errors from O(ε) down to O(ε²).
Personal working taxonomy for categorizing papers as Method, Theory, Resource, Systematization, Position, Discovery, or Application contributions using a superposition model.
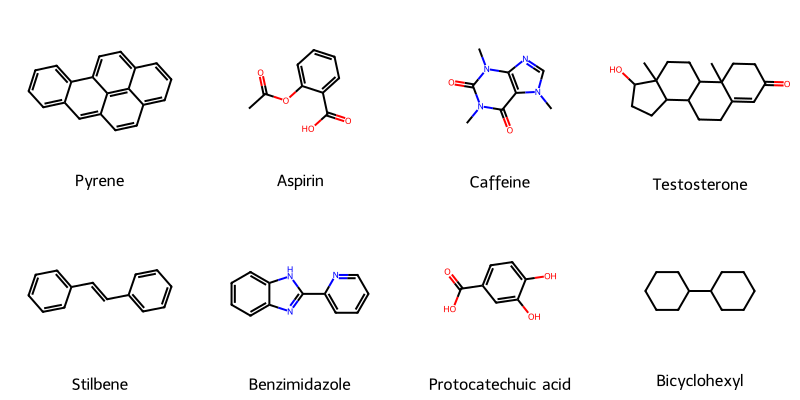
An RDKit wrapper treating molecular visualization as a software engineering problem, implementing strategy pattern for SVG generation with automatic raster fallback, native SELFIES support for generative AI workflows, and strict type safety for batch processing in molecular ML training pipelines.

Explore two fundamental approaches to generating exponentially distributed random numbers: the modern inverse transform method using logarithms and von Neumann’s ingenious 1951 comparison-based algorithm that avoids transcendental functions entirely.
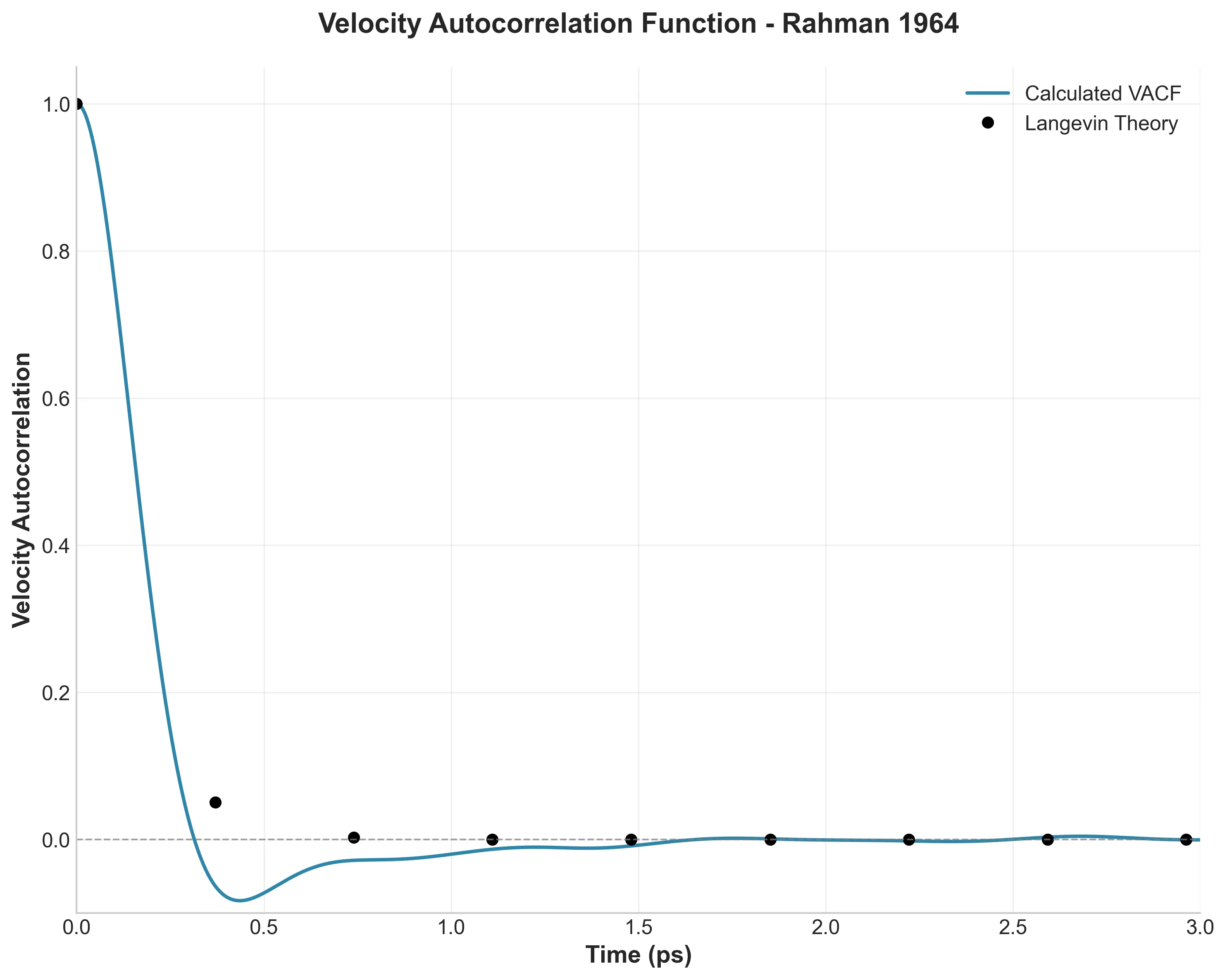
A digital restoration of Rahman’s seminal 1964 molecular dynamics paper using LAMMPS and a Python analysis pipeline featuring decorator-based caching, fully vectorized NumPy computations for O(N^2) operations, and modern tooling (uv, type hints, Makefile automation) transforming academic scripts into a reproducible research toolkit.

I replicated Rahman’s landmark 1964 liquid argon molecular dynamics simulation using modern tools, building a Python analysis pipeline with caching, vectorization, and type hints to bridge vintage science with modern software engineering.
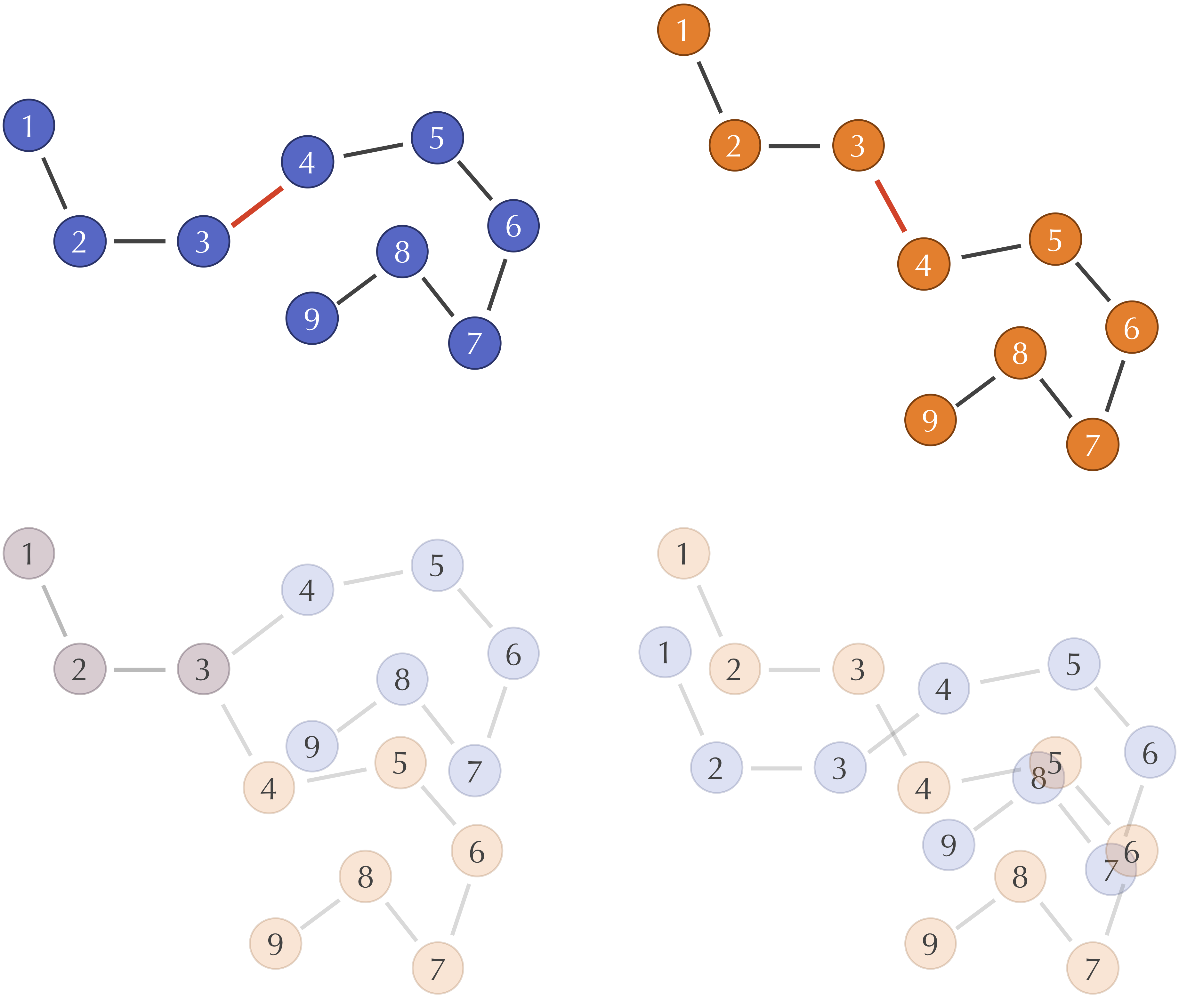
Learn to align molecular structures and point clouds using the Kabsch algorithm, with differentiable implementations for modern ML frameworks.
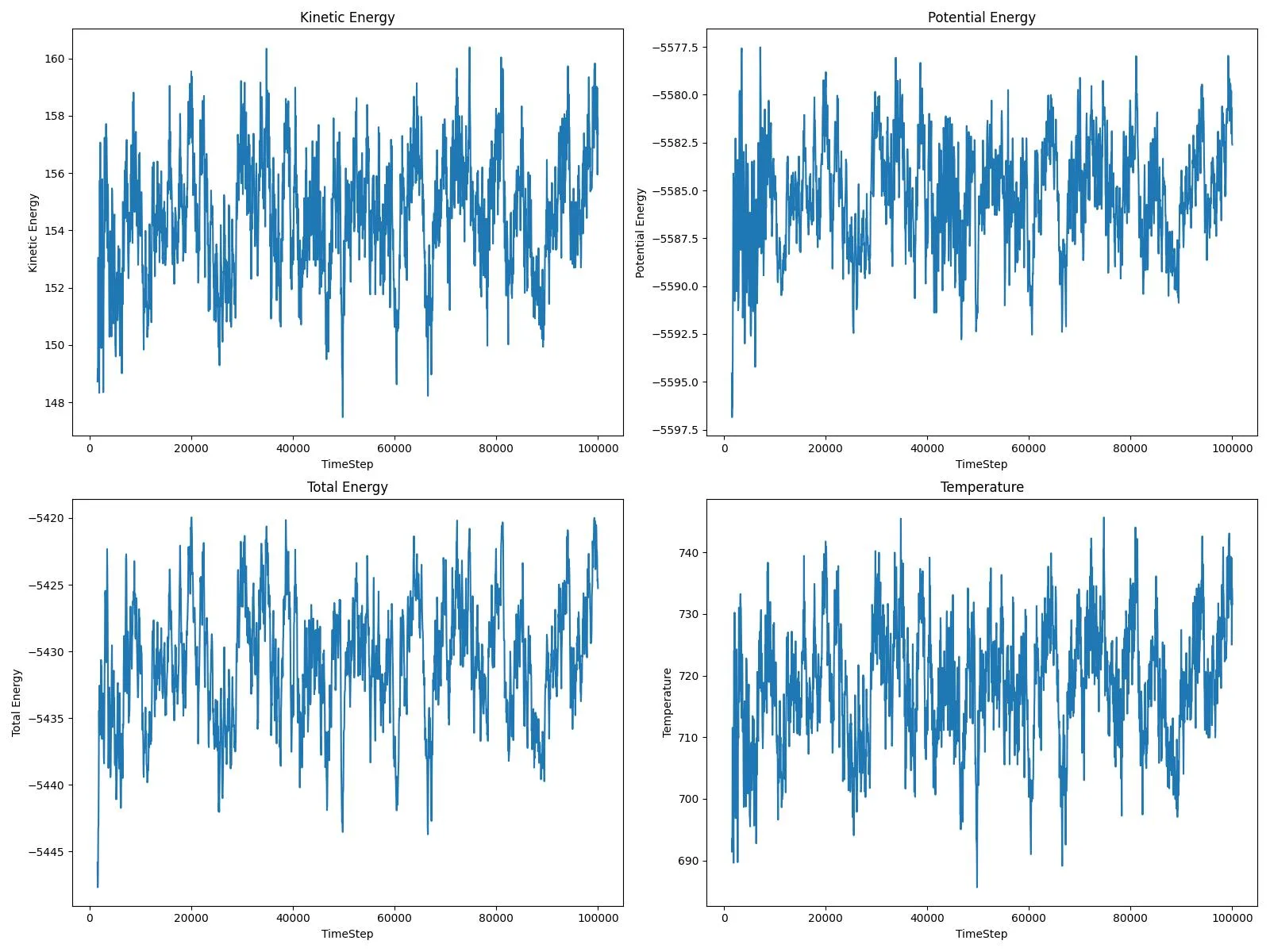
An input-to-analysis workflow for simulating adatom diffusion on FCC metal surfaces using LAMMPS and EAM potentials, covering copper and platinum to compare how atomic mass and bonding strength affect surface dynamics, with a Python analysis layer that generates energy and trajectory diagnostic plots. The LAMMPS setup is adapted from Eric N. Hahn’s adatom tutorial.
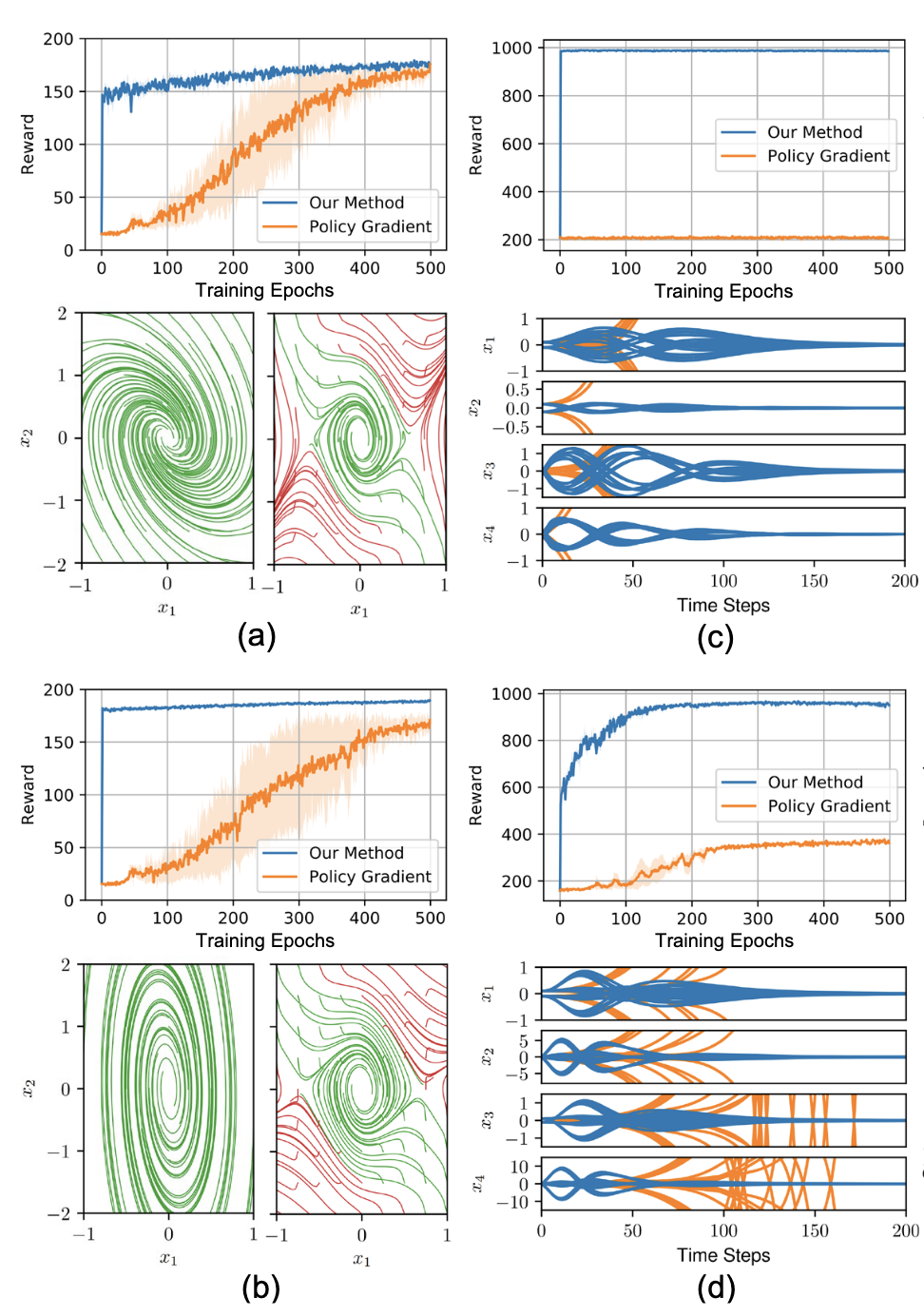
A PyTorch implementation enforcing strict Lyapunov stability guarantees on recurrent neural network controllers through Integral Quadratic Constraints, bridging 1990s robust control theory with modern deep reinforcement learning by solving semidefinite programs inside the gradient descent loop to provide mathematical certificates of safety.
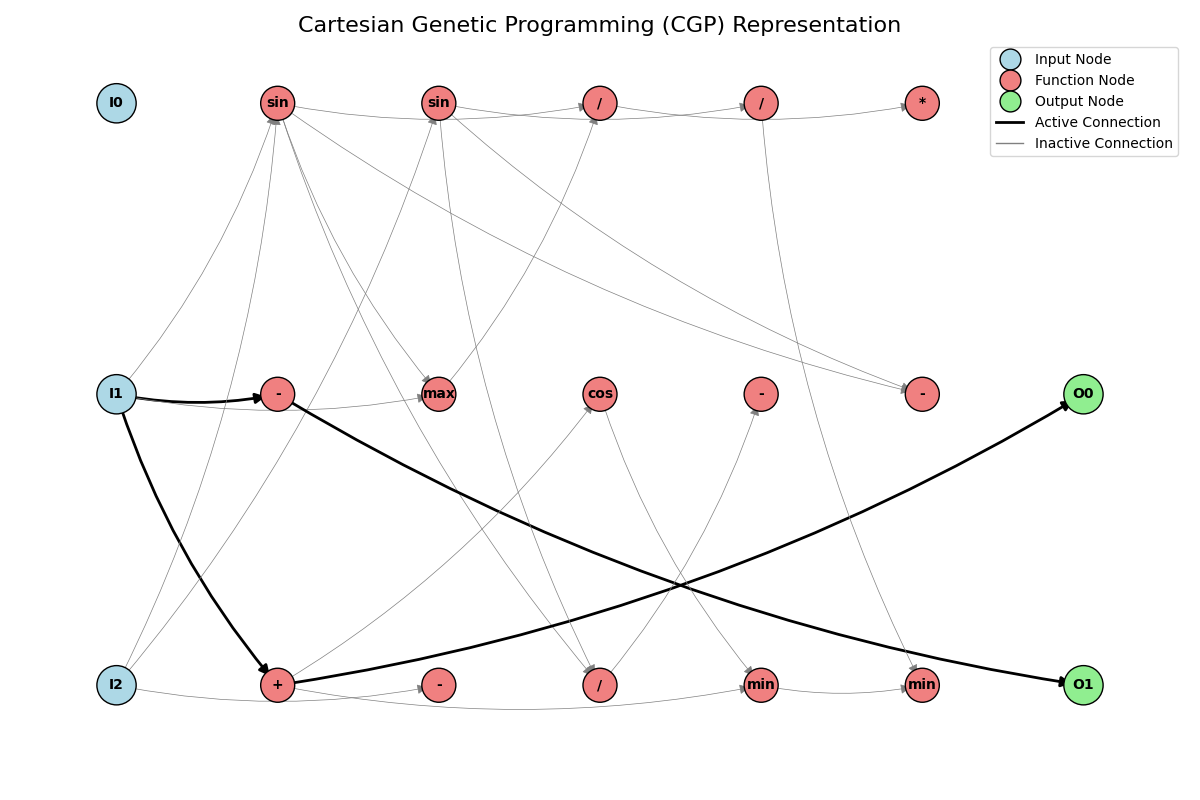
A 2018 exploration of evolutionary algorithms applied to Atari games, forking Dennis Wilson’s d9w/CGP.jl, where my contribution was the Atari reinforcement-learning experiment layer (experiment variants, fitness functions, hyperparameter tuning) on top of the upstream Cartesian Genetic Programming framework and its NEAT-inspired speciation.