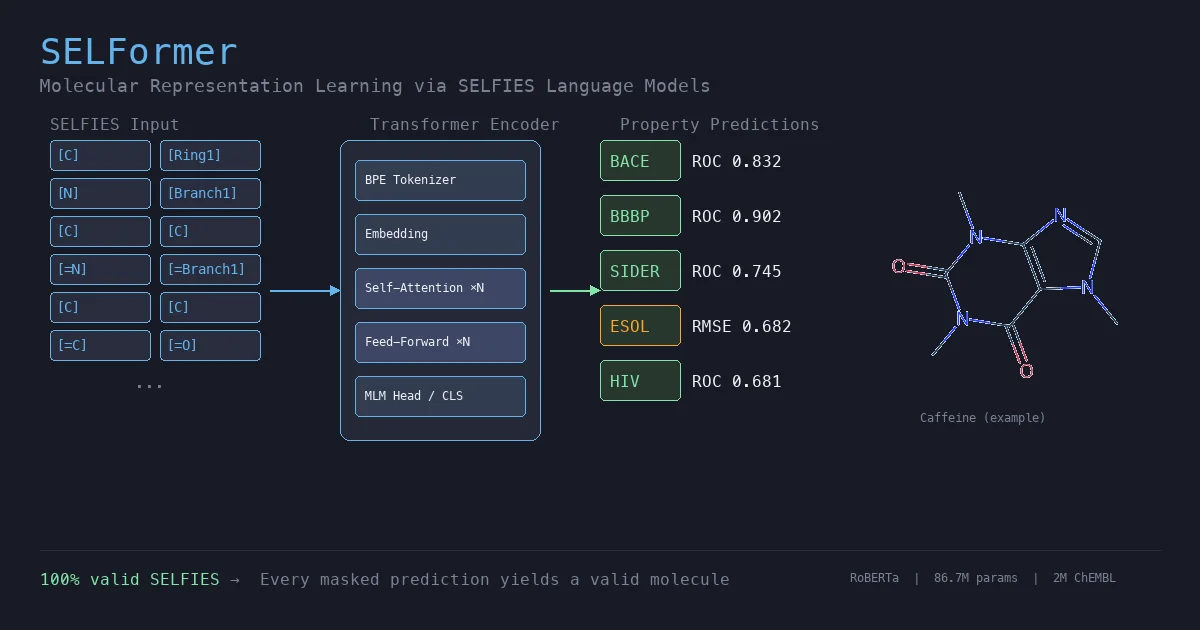
SELFormer: A SELFIES-Based Molecular Language Model
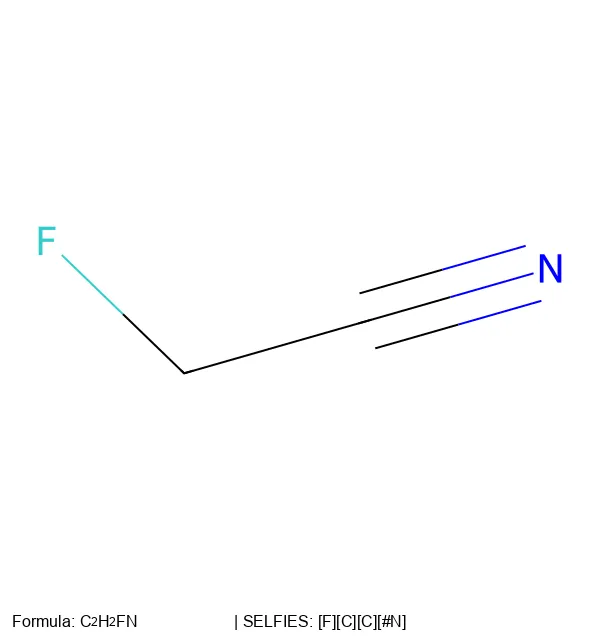
SELFormer is a transformer-based chemical language model that uses SELFIES instead of SMILES as input. Pretrained on 2M ChEMBL compounds via masked language modeling, it achieves strong classification performance on MoleculeNet tasks, outperforming ChemBERTa-2 by ~12% on average across BACE, BBBP, and HIV.