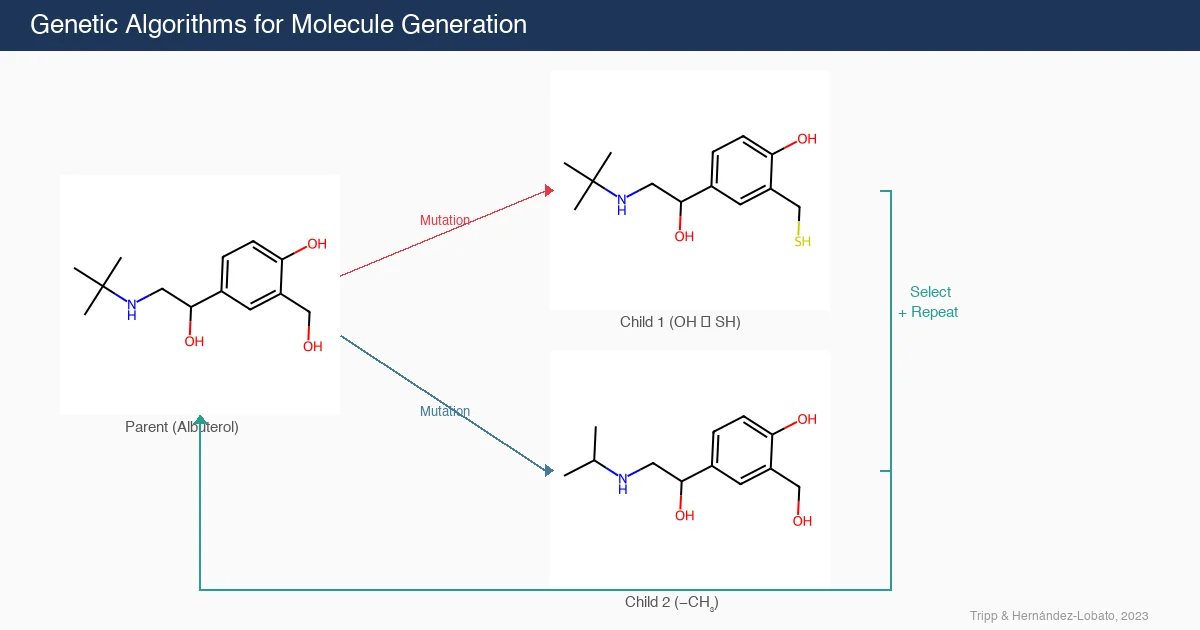
Graph-Based GA and MCTS Generative Model for Molecules
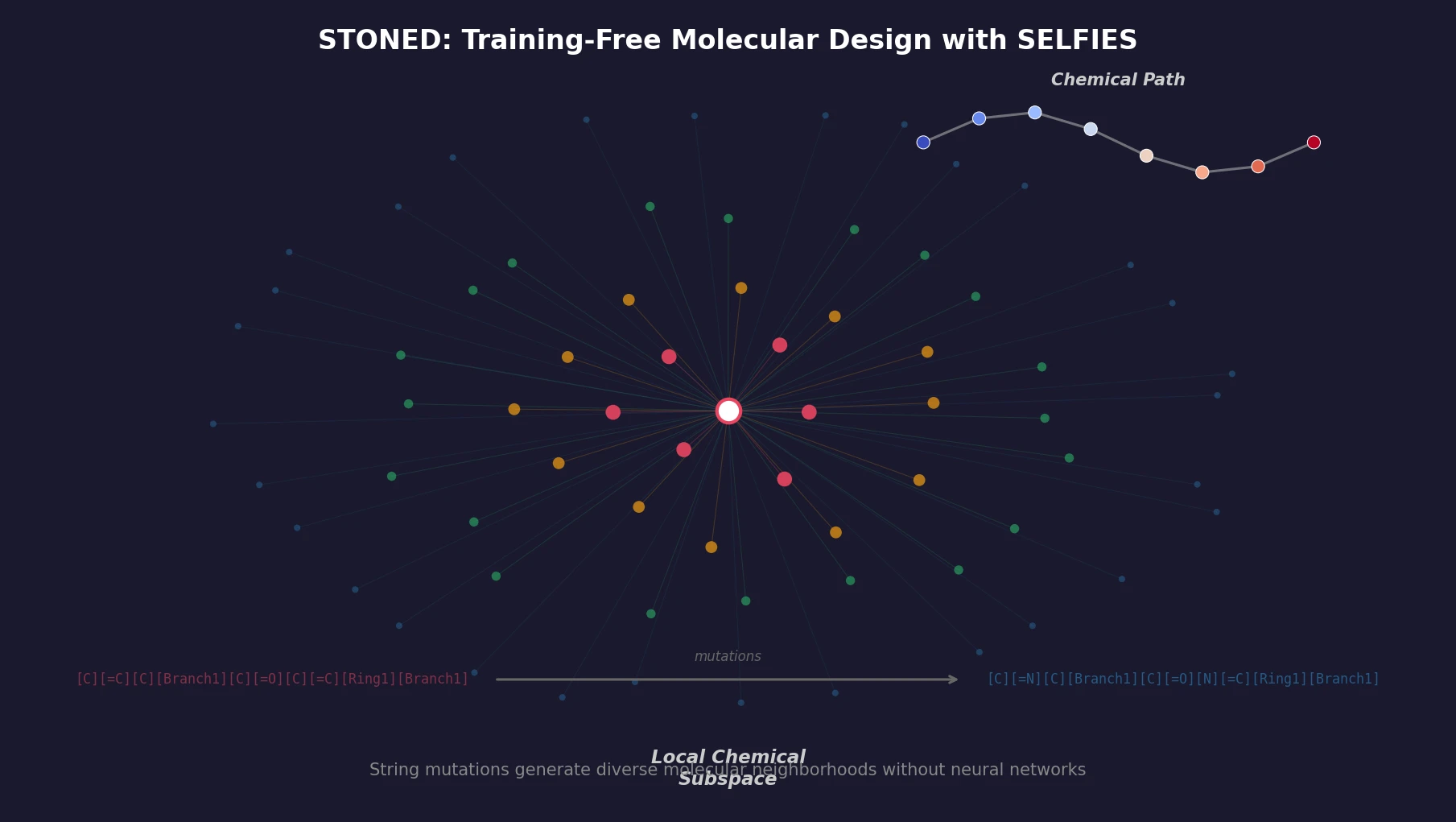
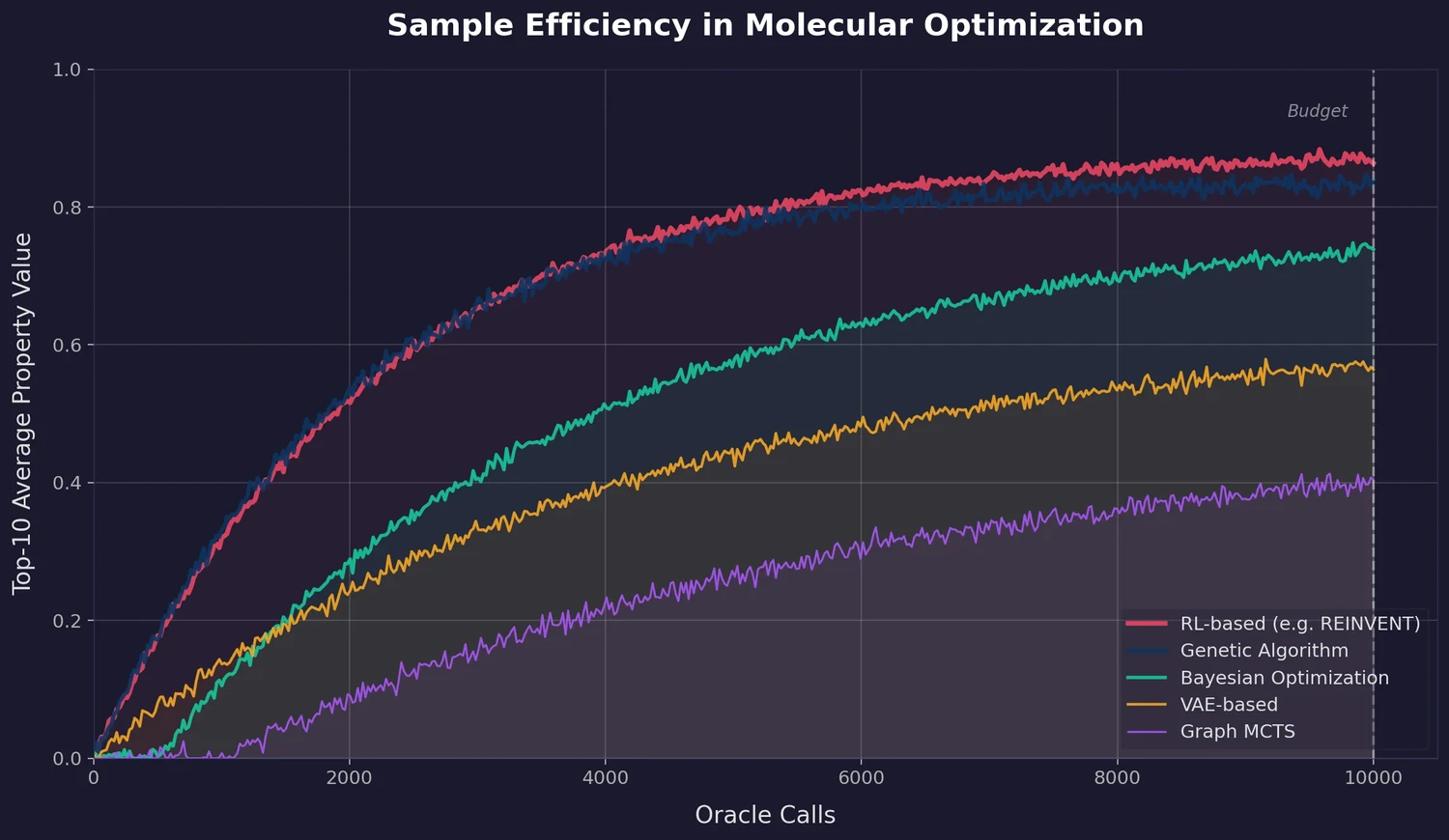
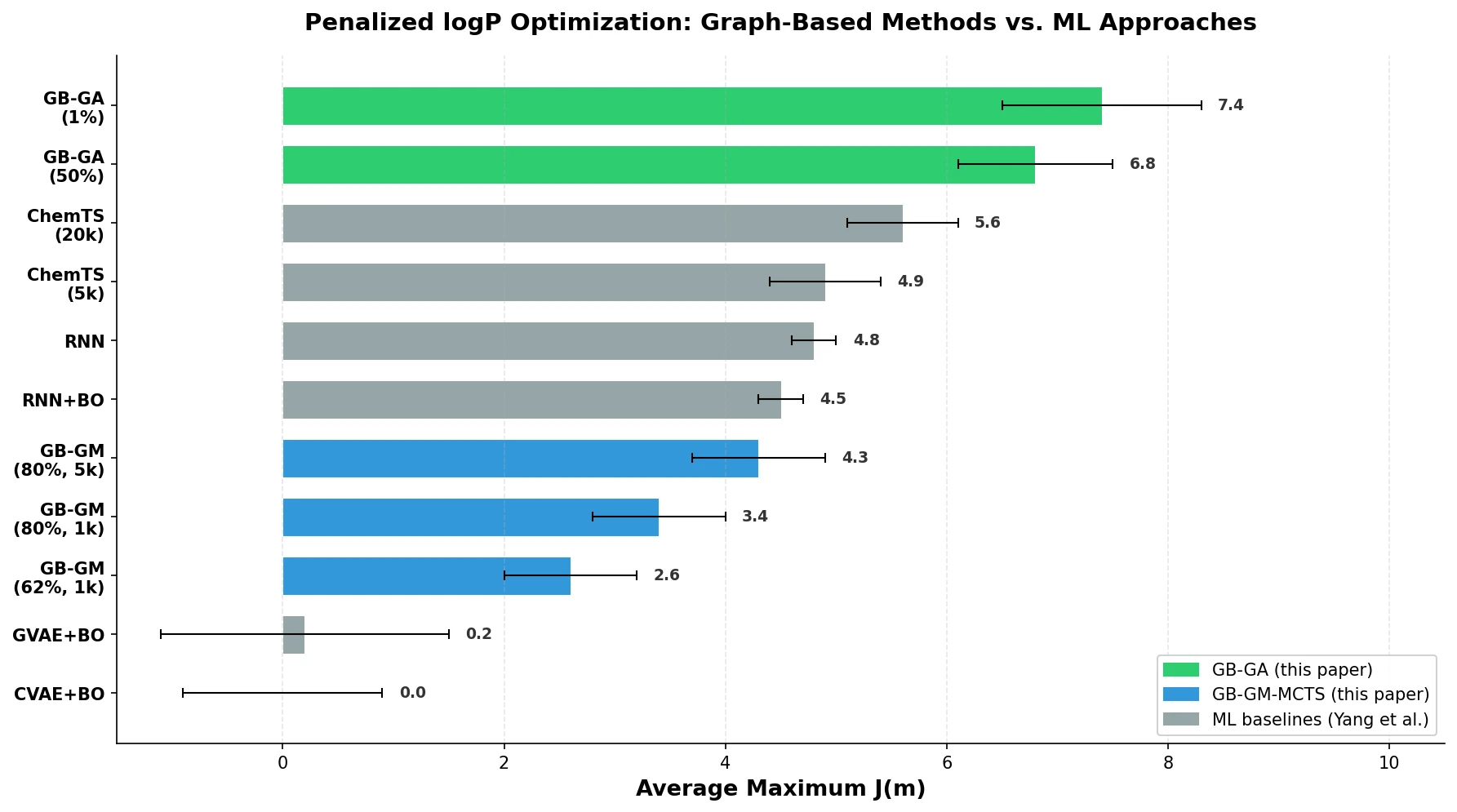
A graph-based genetic algorithm (GB-GA) and a graph-based generative model with Monte Carlo tree search (GB-GM-MCTS) for molecular optimization that match or outperform ML-based generative approaches while being orders of magnitude faster.