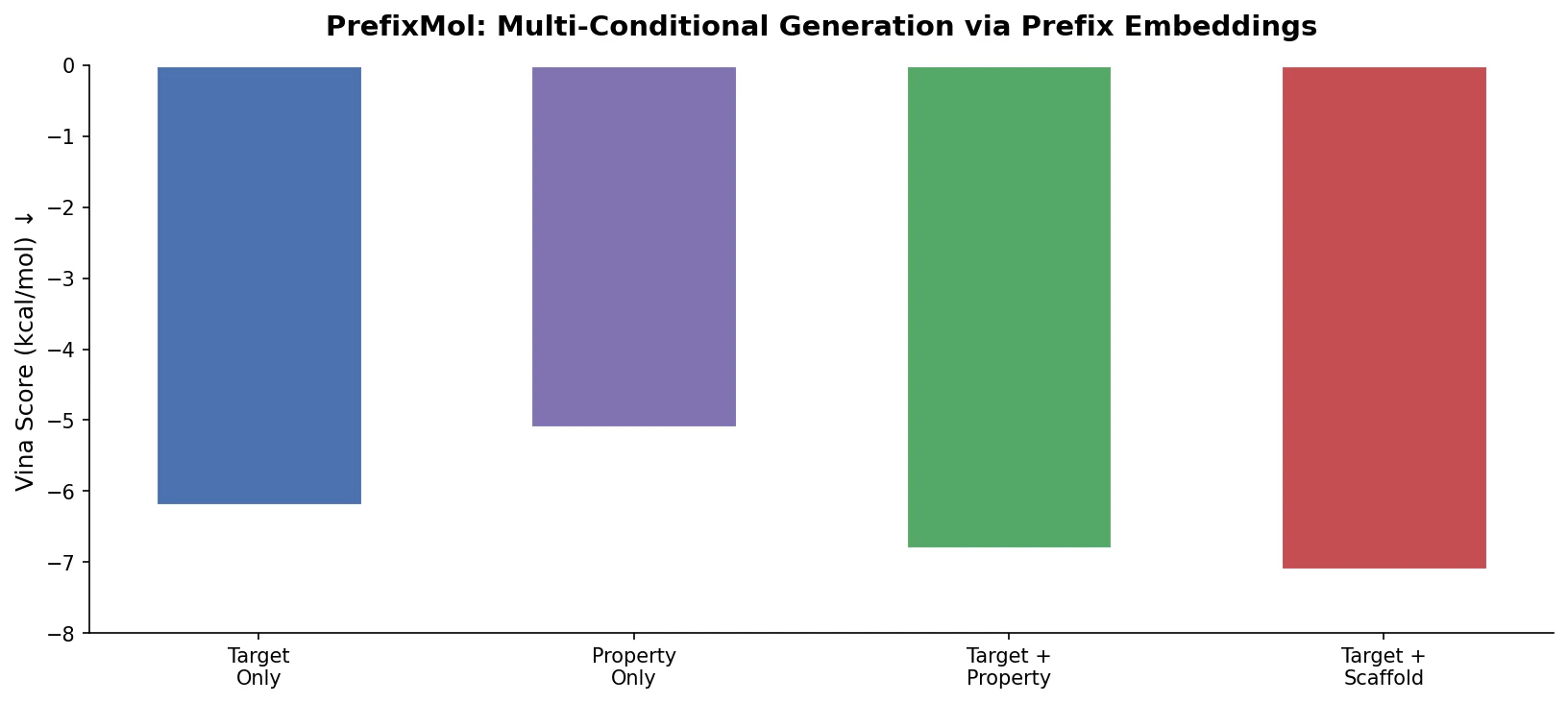
PrefixMol: Prefix Embeddings for Drug Molecule Design
PrefixMol prepends learnable condition vectors to a GPT transformer for SMILES generation, enabling joint control over binding pocket targeting and chemical properties like QED, SA, and LogP.

PrefixMol prepends learnable condition vectors to a GPT transformer for SMILES generation, enabling joint control over binding pocket targeting and chemical properties like QED, SA, and LogP.
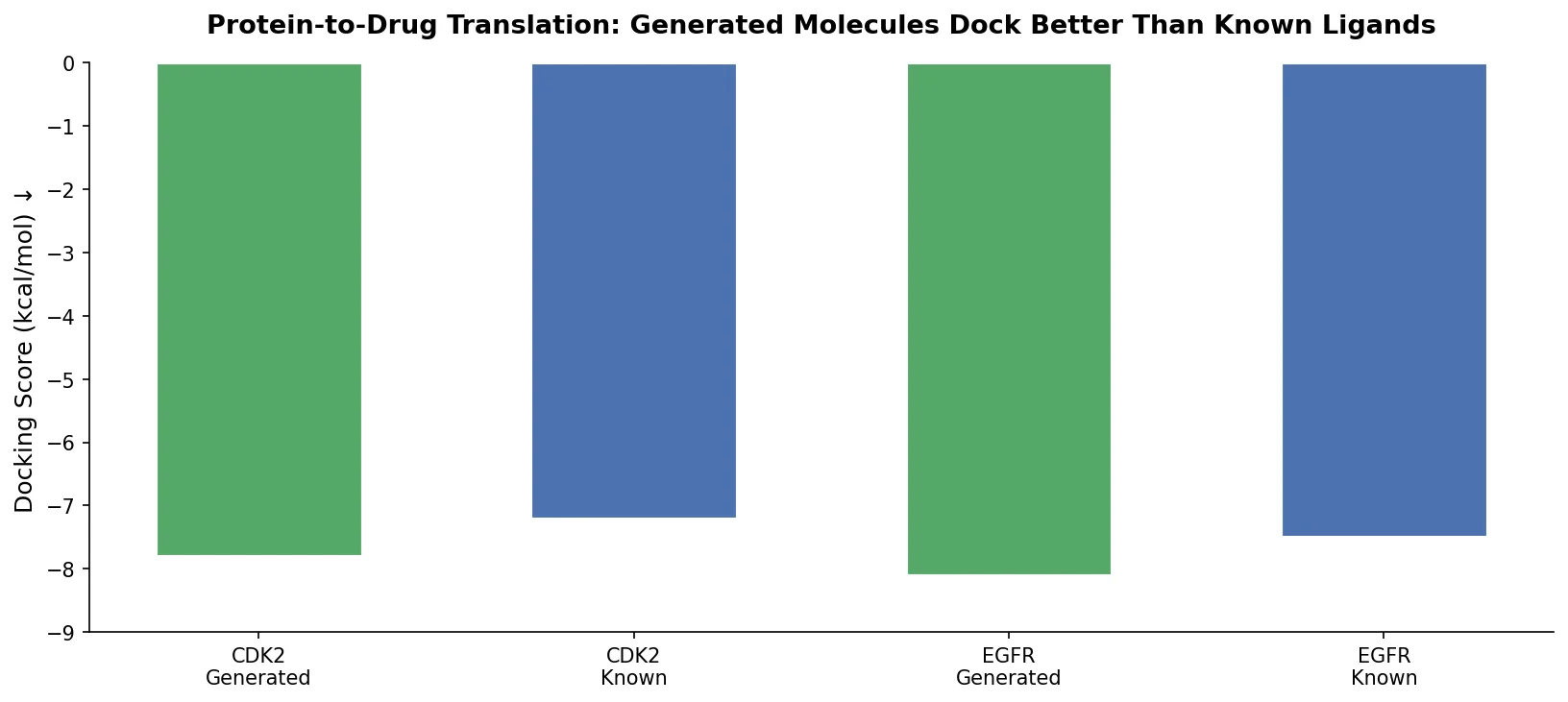
Applies the Transformer architecture to generate drug-like molecules conditioned on protein amino acid sequences, treating target-specific de novo drug design as a sequence-to-sequence translation problem.
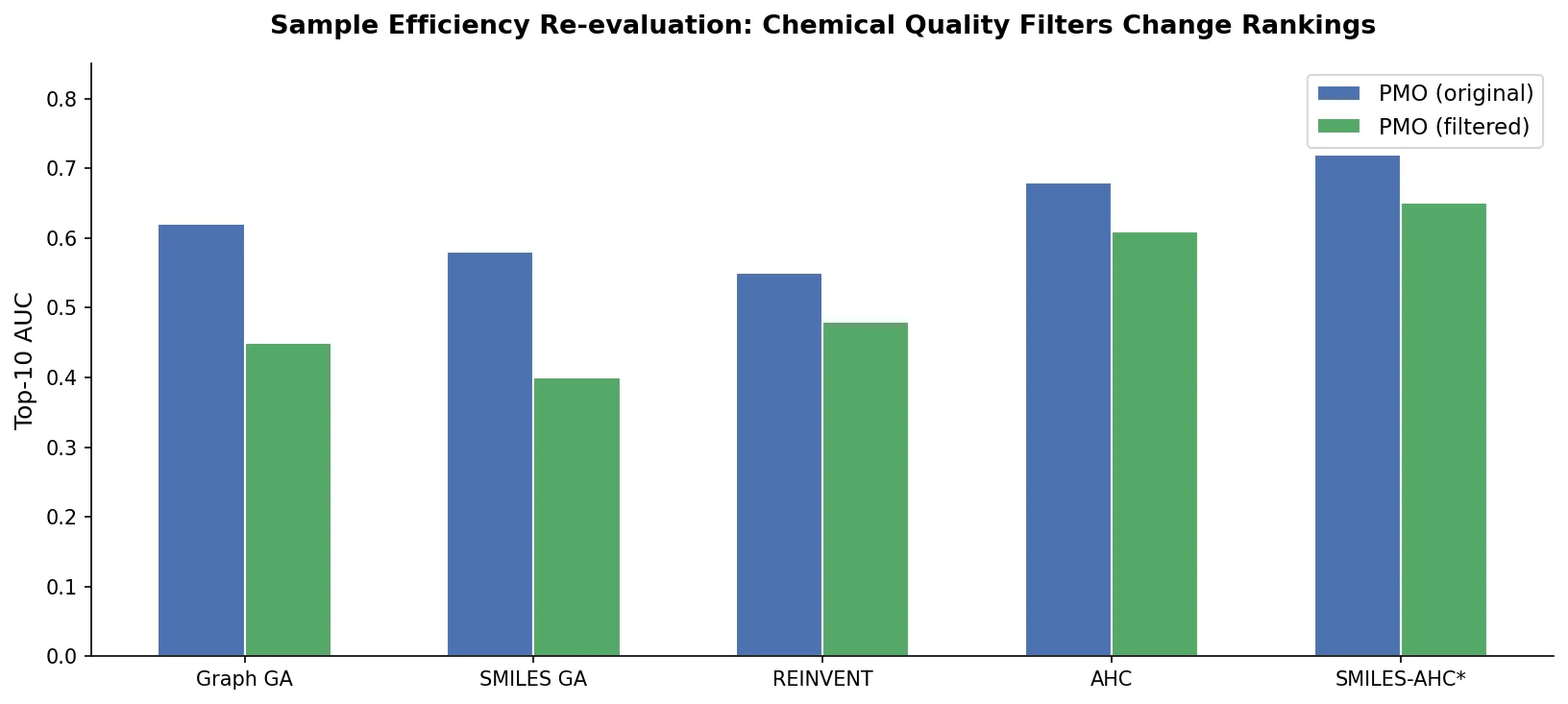
A critical reassessment of the PMO benchmark for de novo molecule generation, showing that adding molecular weight, LogP, and diversity filters substantially re-ranks generative models, with Augmented Hill-Climb emerging as the top method.
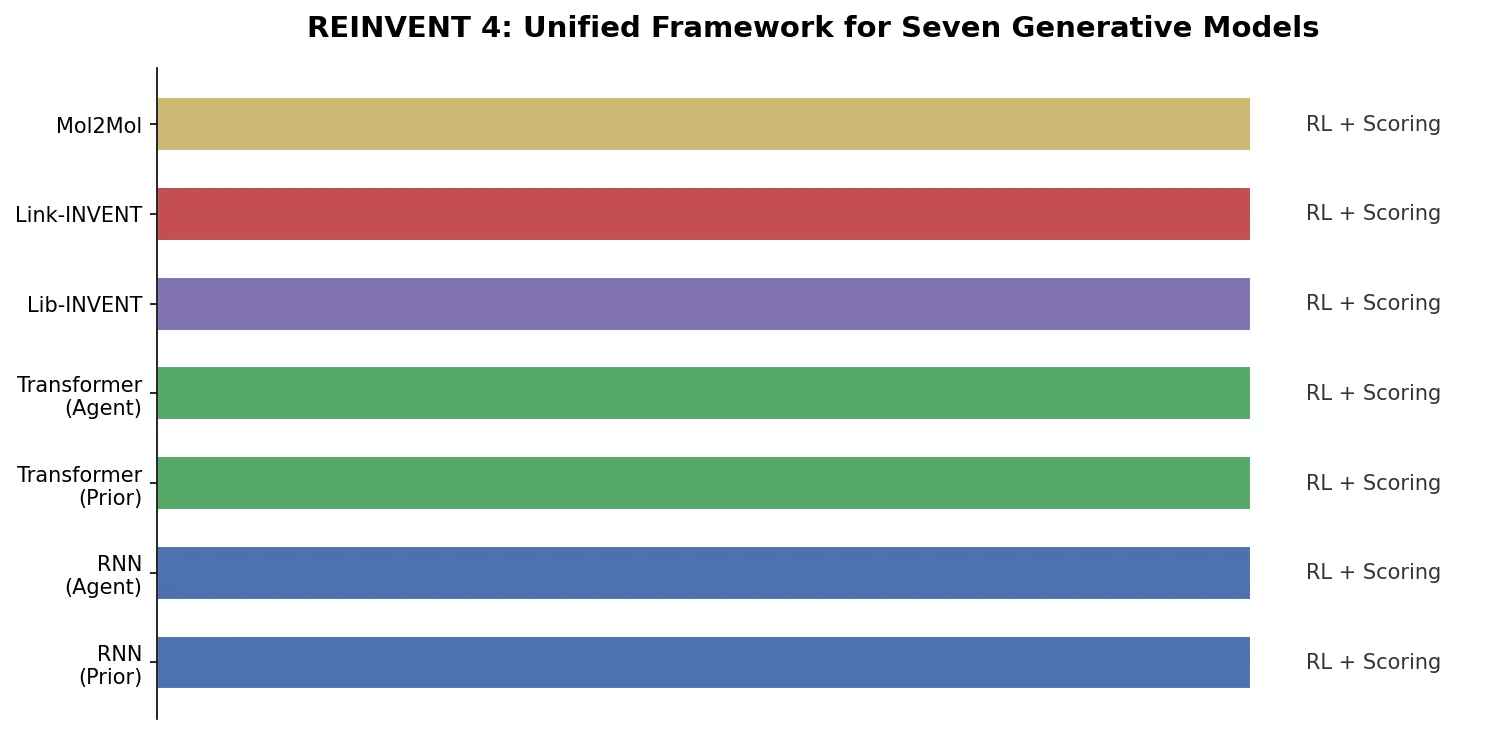
Overview of REINVENT 4, an open-source generative molecular design framework from AstraZeneca that unifies RNN and transformer generators within reinforcement learning, transfer learning, and curriculum learning optimization algorithms.
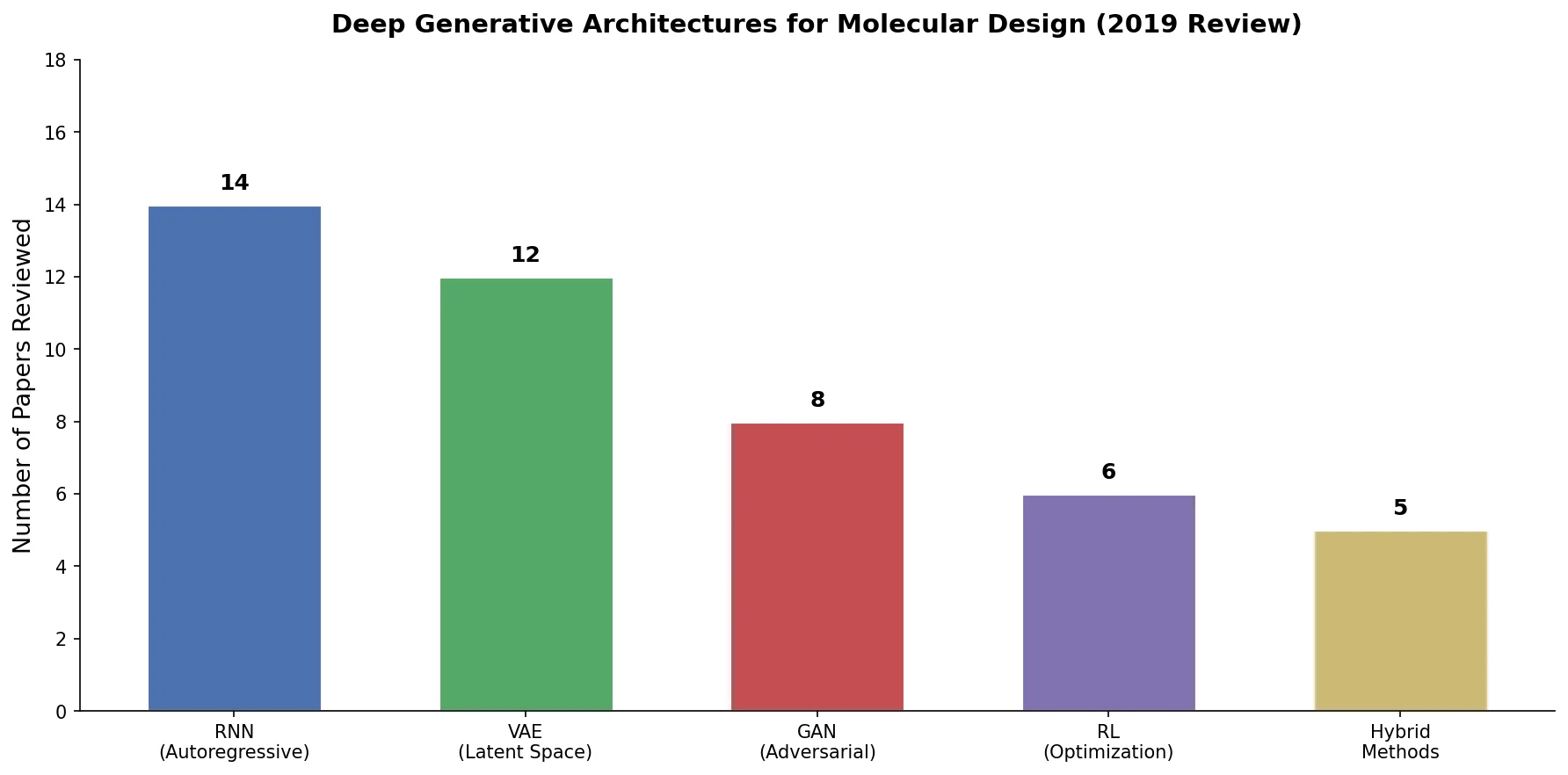
An early and influential review cataloging 45 papers on deep generative modeling for molecules, comparing RNN, VAE, GAN, and reinforcement learning architectures across SMILES and graph-based representations.
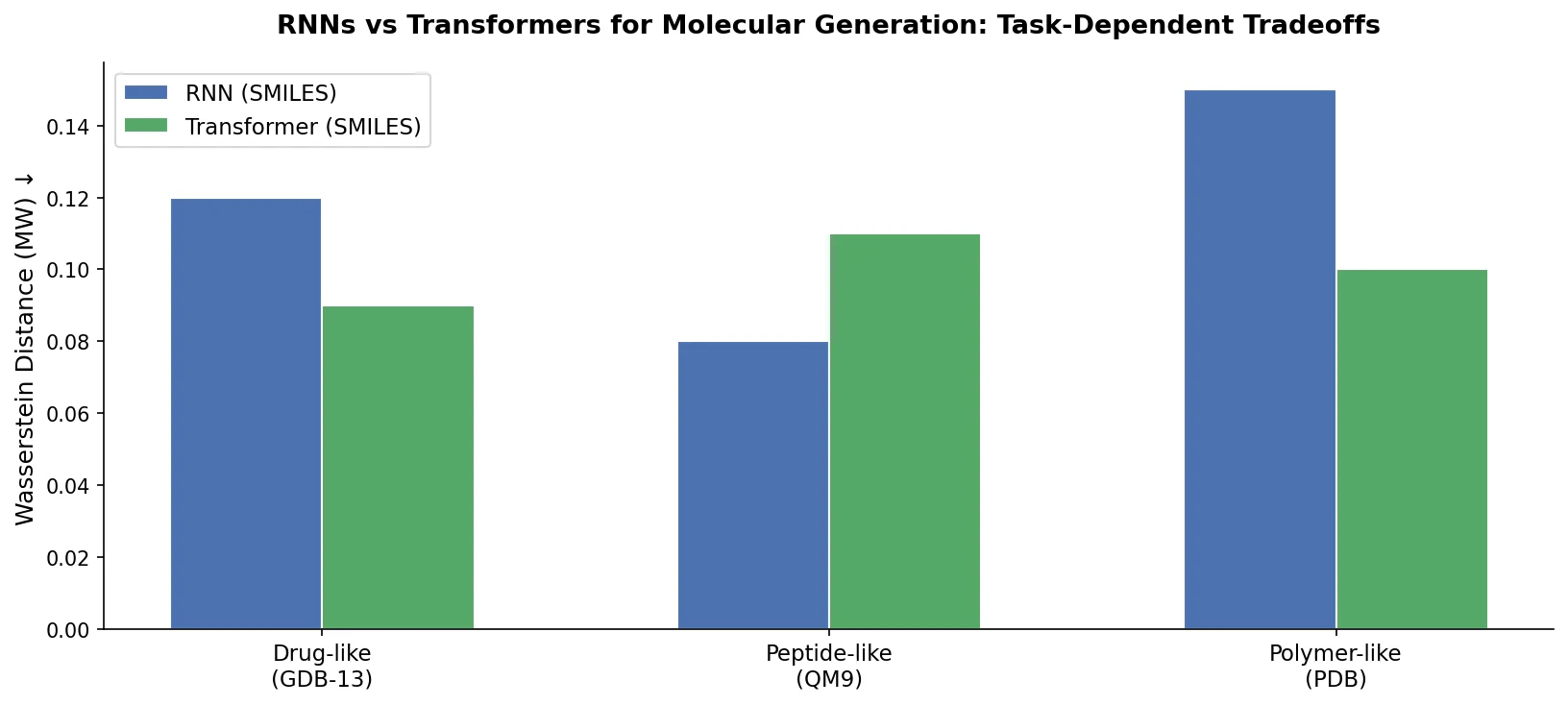
Compares RNN-based and Transformer-based chemical language models across three molecular generation tasks of increasing complexity, finding that RNNs excel at local features while Transformers handle large molecules better.
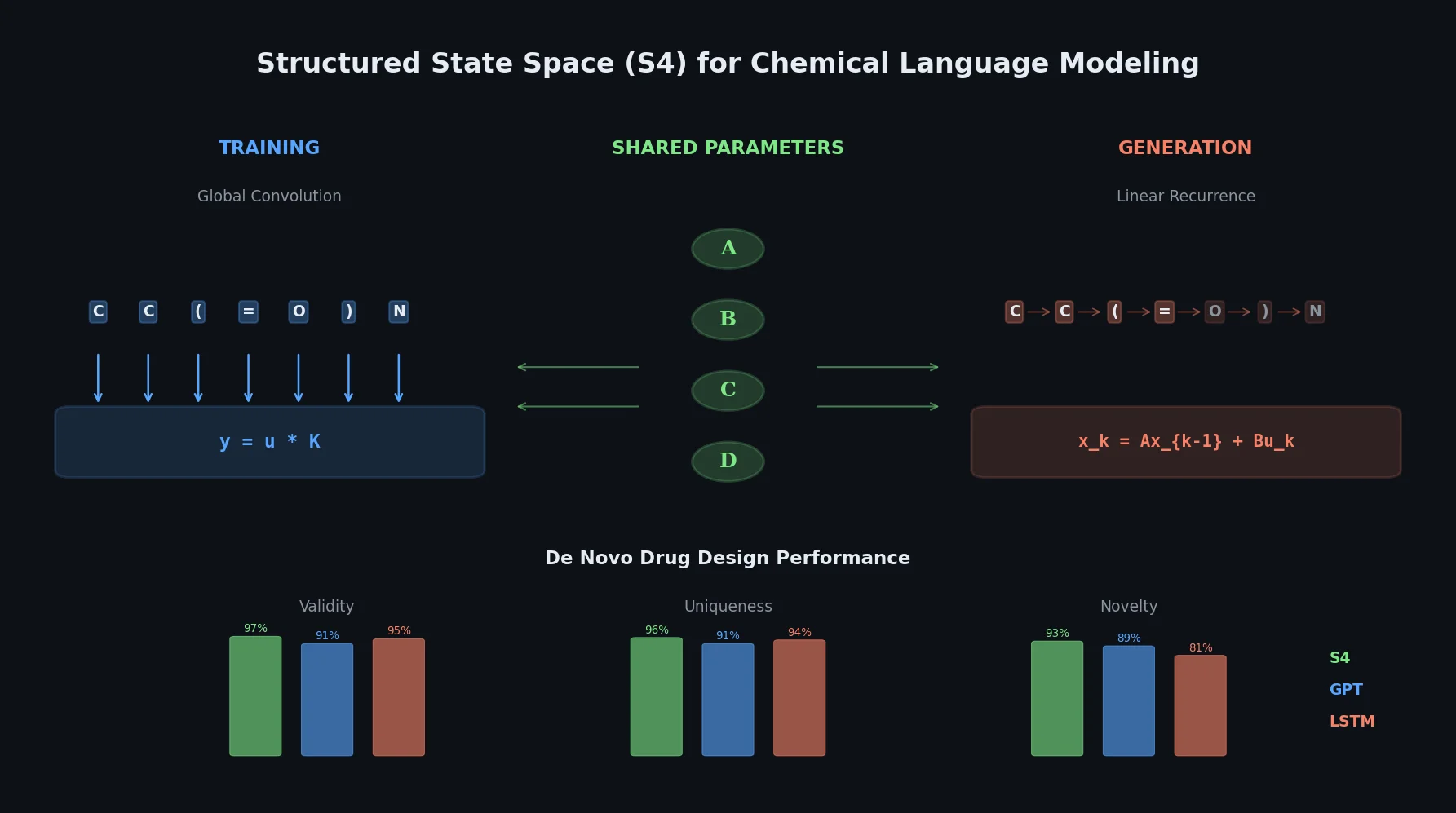
This paper introduces structured state space sequence (S4) models to chemical language modeling, showing they combine the strengths of LSTMs (efficient recurrent generation) and GPTs (holistic sequence learning) for de novo molecular design.
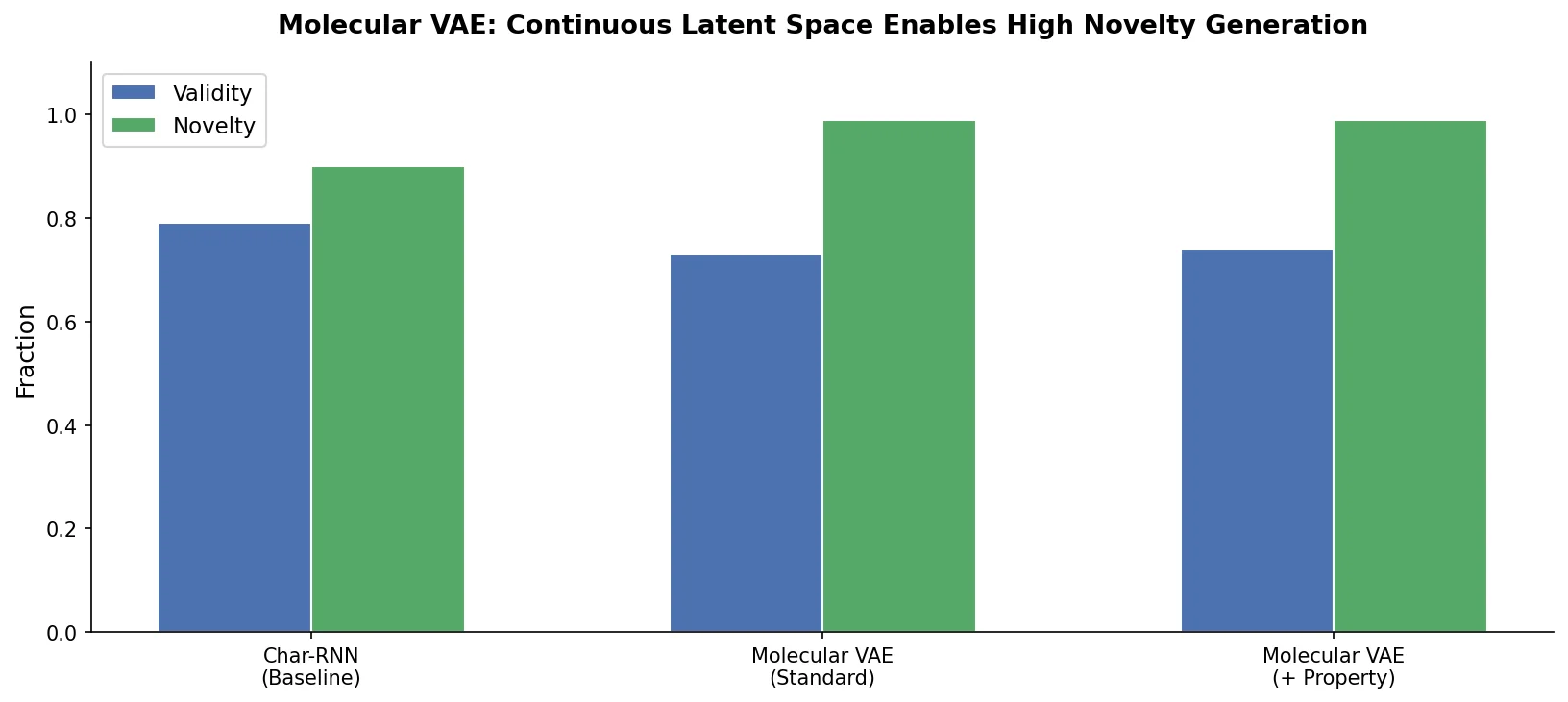
This foundational paper introduces a variational autoencoder (VAE) that encodes SMILES strings into a continuous latent space, allowing gradient-based optimization of molecular properties. Joint training with a property predictor organizes the latent space by chemical properties, and Bayesian optimization over the latent surface discovers drug-like molecules with improved QED and synthetic accessibility.
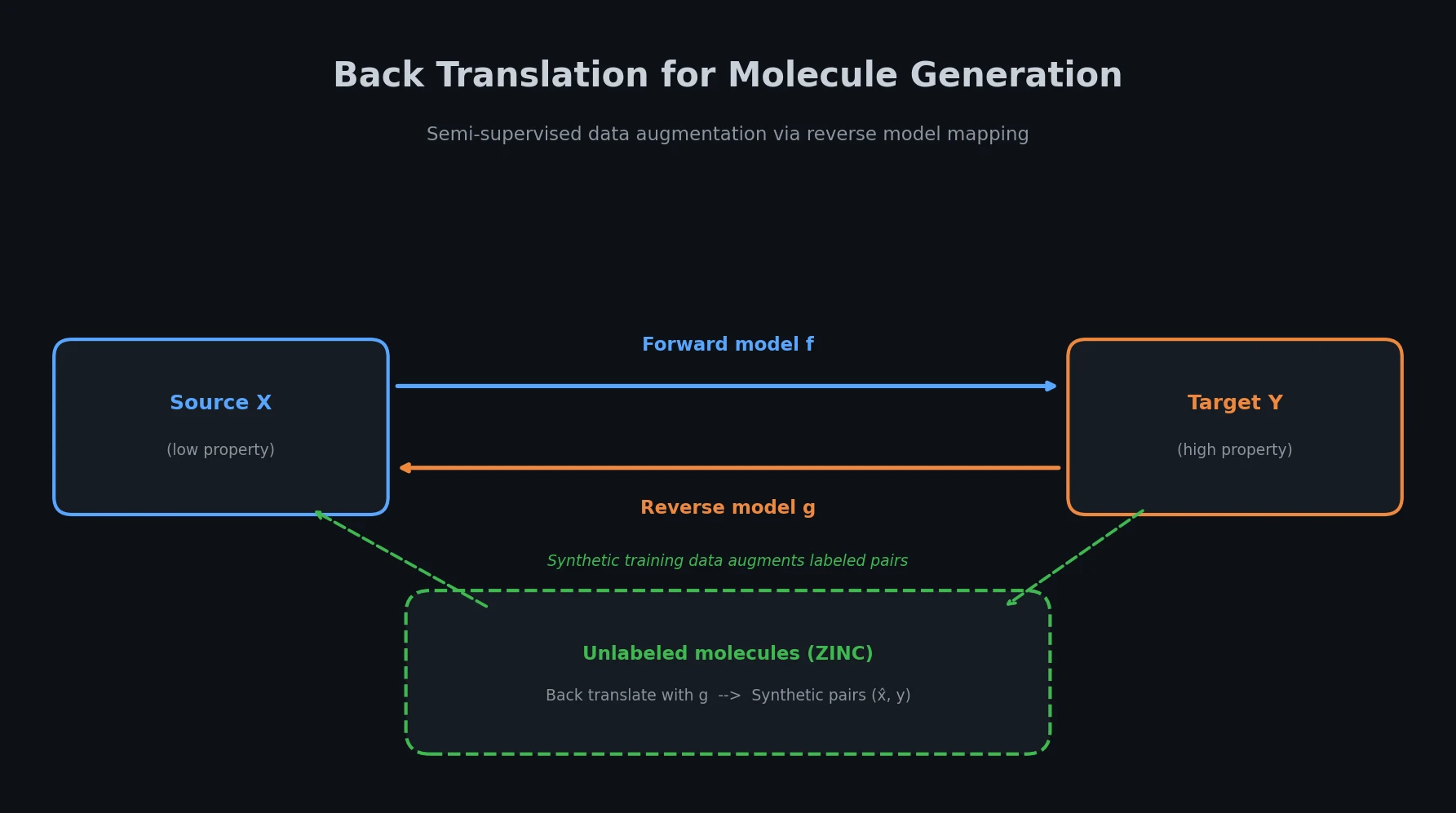
Adapts back translation from NLP to molecular generation, using unlabeled molecules from ZINC to create synthetic training pairs that improve property optimization and retrosynthesis prediction across Transformer and graph-based architectures.
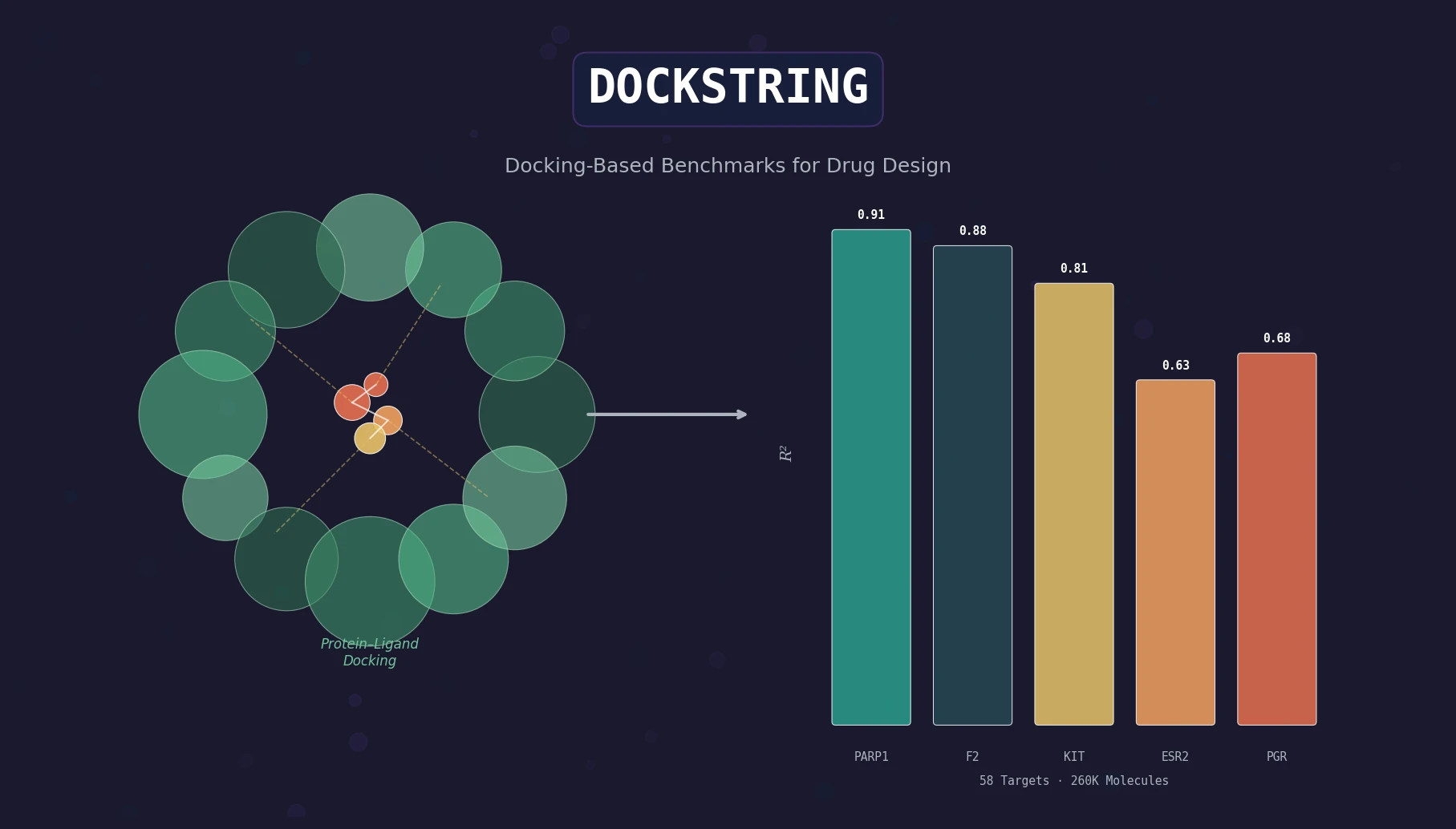
DOCKSTRING bundles an AutoDock Vina wrapper, a 260K-molecule docking dataset across 58 protein targets, and pharmaceutically relevant benchmarks for regression, virtual screening, and de novo design.
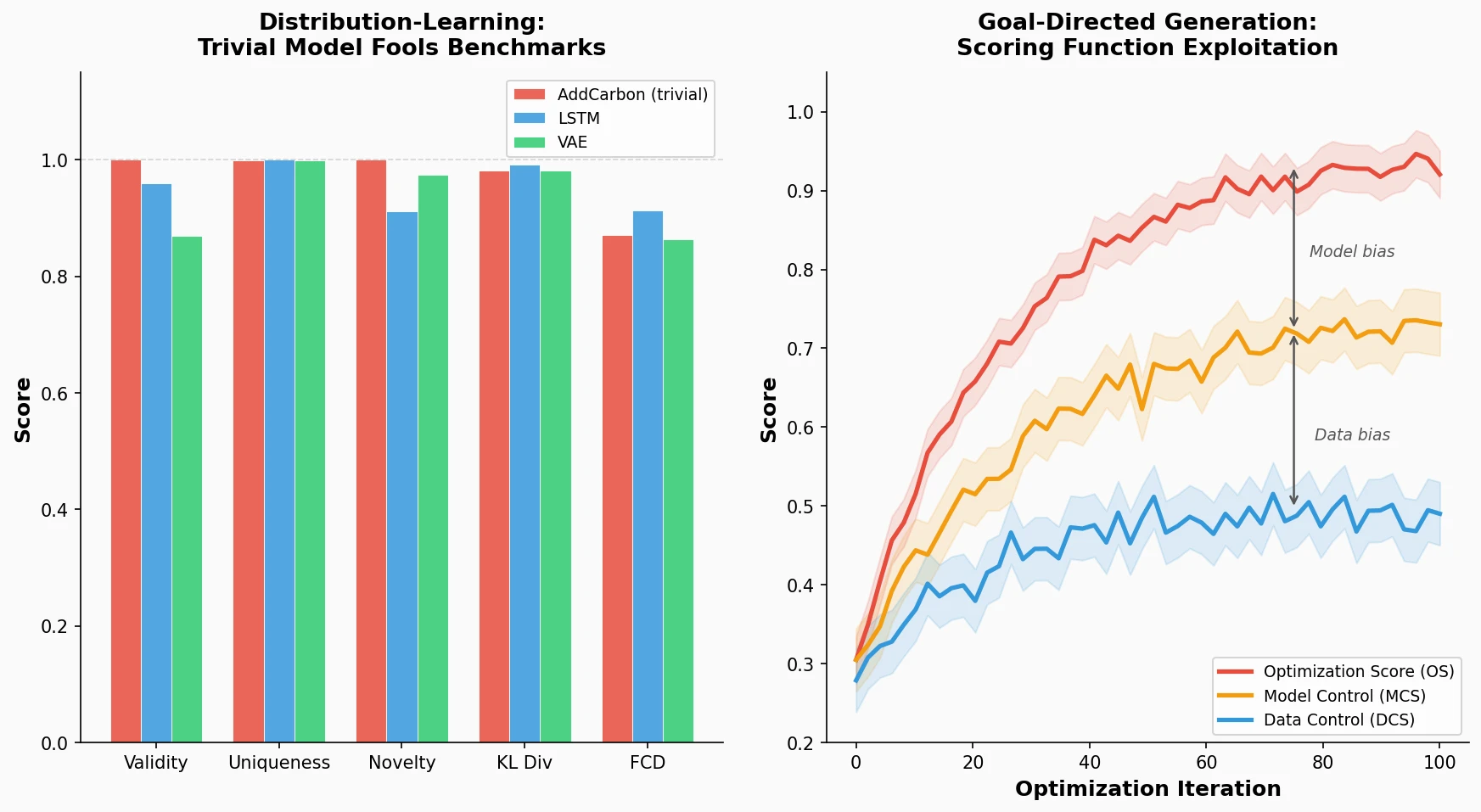
Identifies failure modes in molecular generative models, showing that trivial edits fool distribution-learning benchmarks and that ML-based scoring functions introduce exploitable model-specific and data-specific biases during goal-directed optimization.
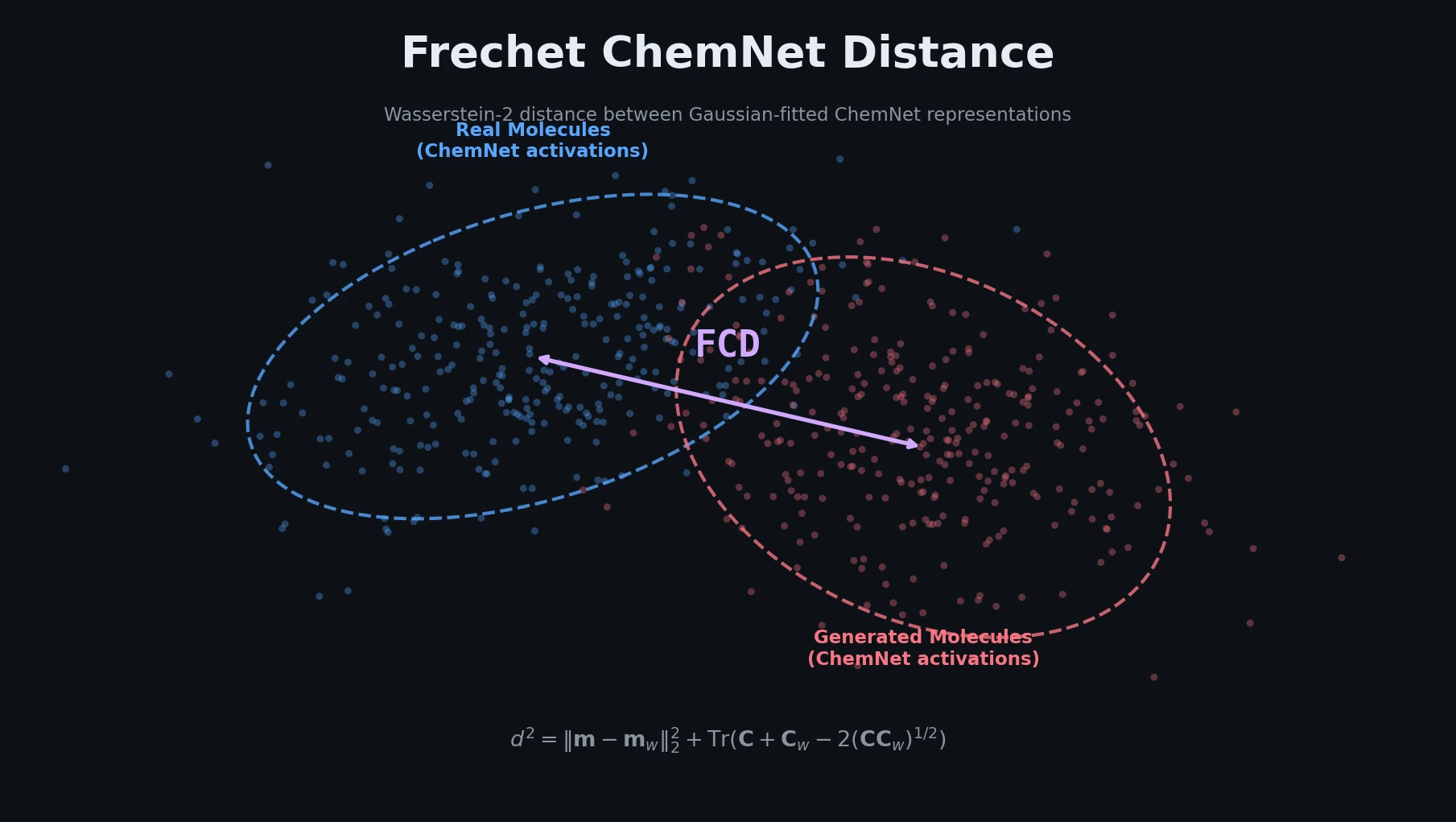
Introduces the Frechet ChemNet Distance (FCD), a single metric that captures chemical validity, biological relevance, and diversity of generated molecules by comparing distributions of learned ChemNet representations.