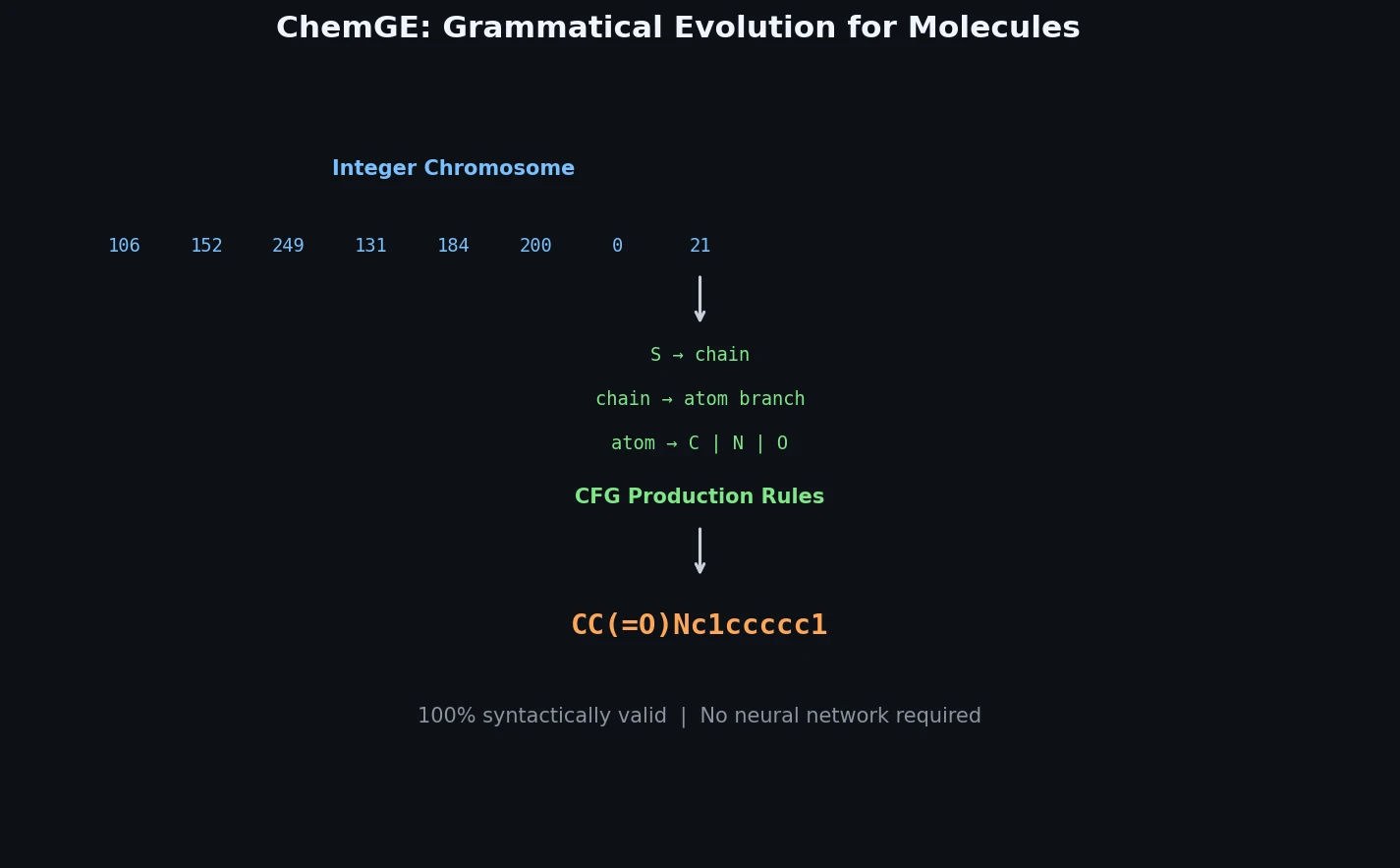
ChemGE: Molecule Generation via Grammatical Evolution
ChemGE uses grammatical evolution over SMILES context-free grammars to generate diverse drug-like molecules in parallel, outperforming deep learning baselines in throughput and molecular diversity.