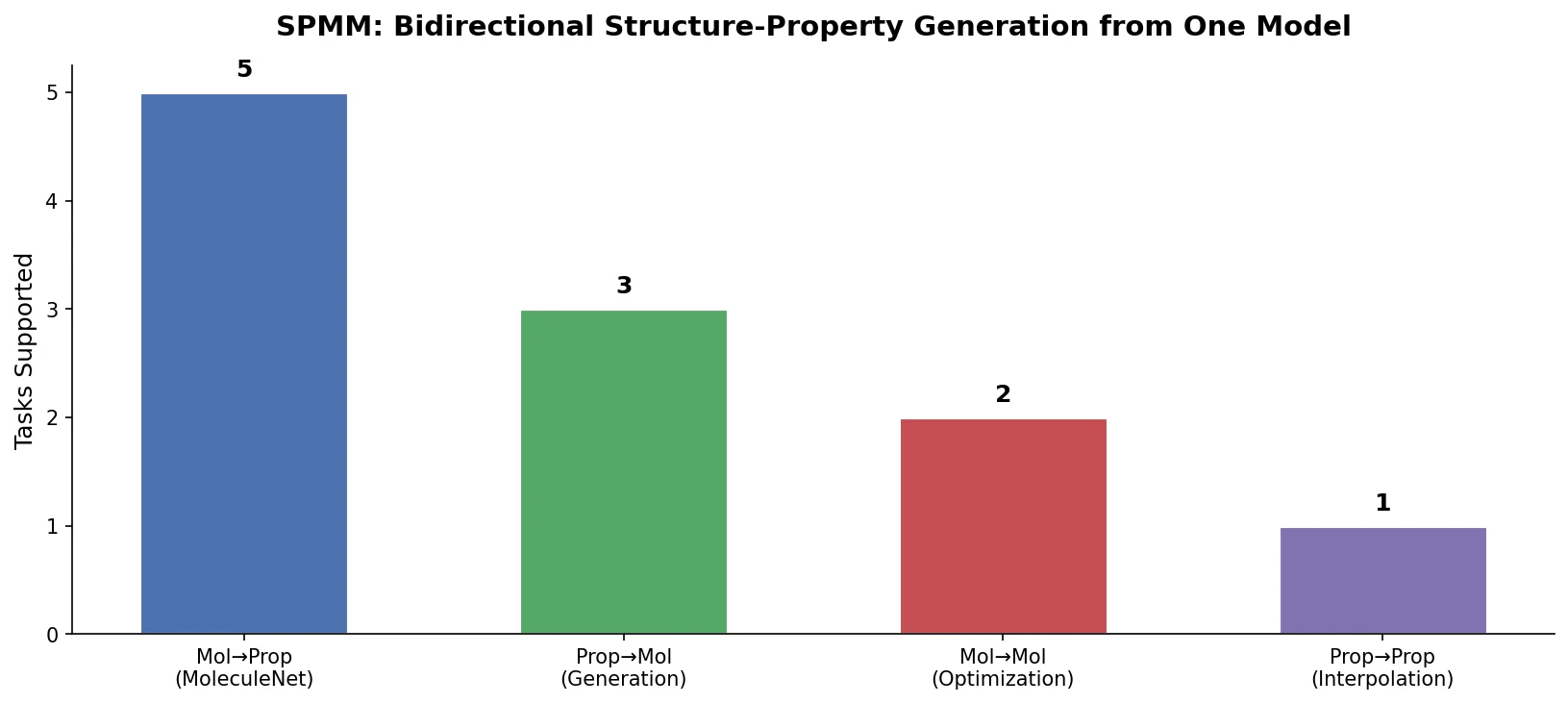
SPMM: A Bidirectional Molecular Foundation Model
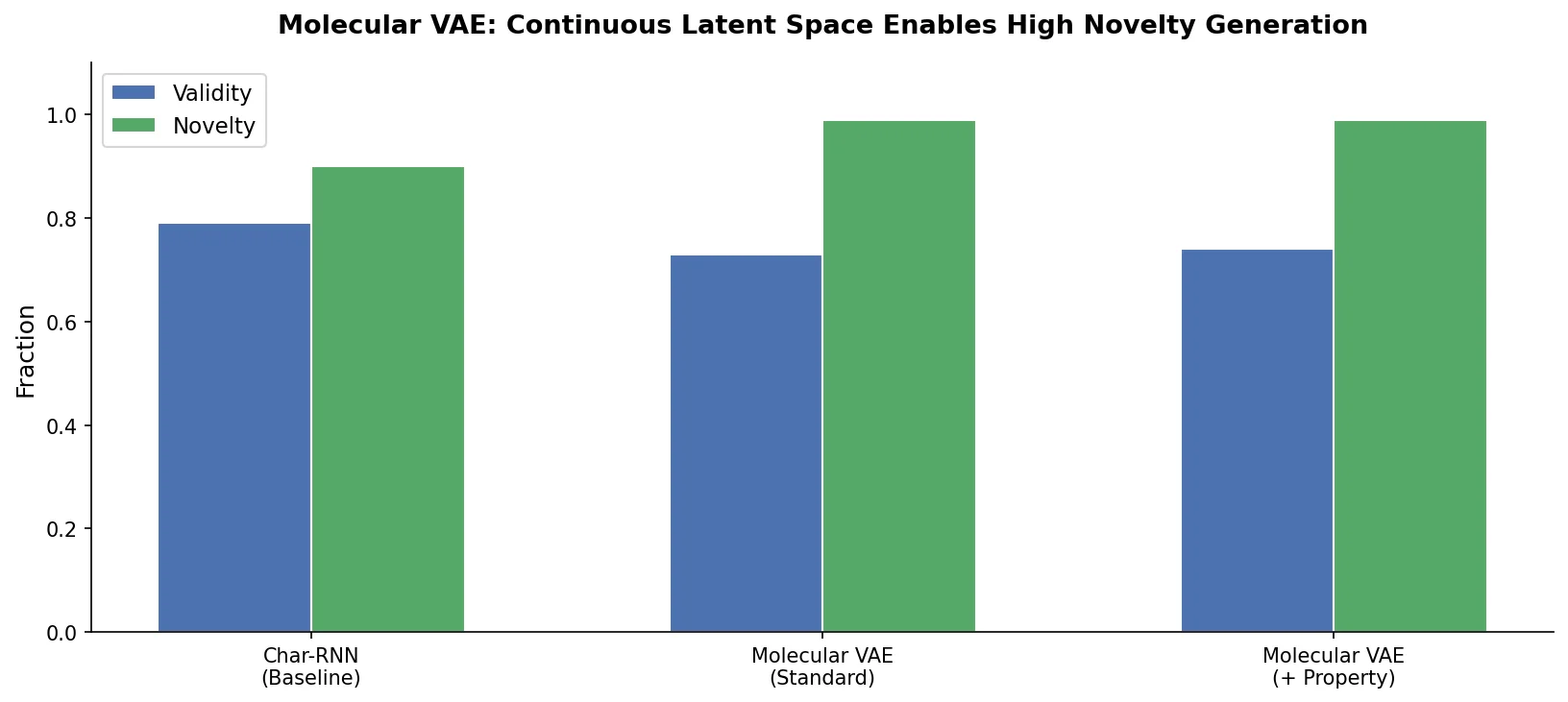
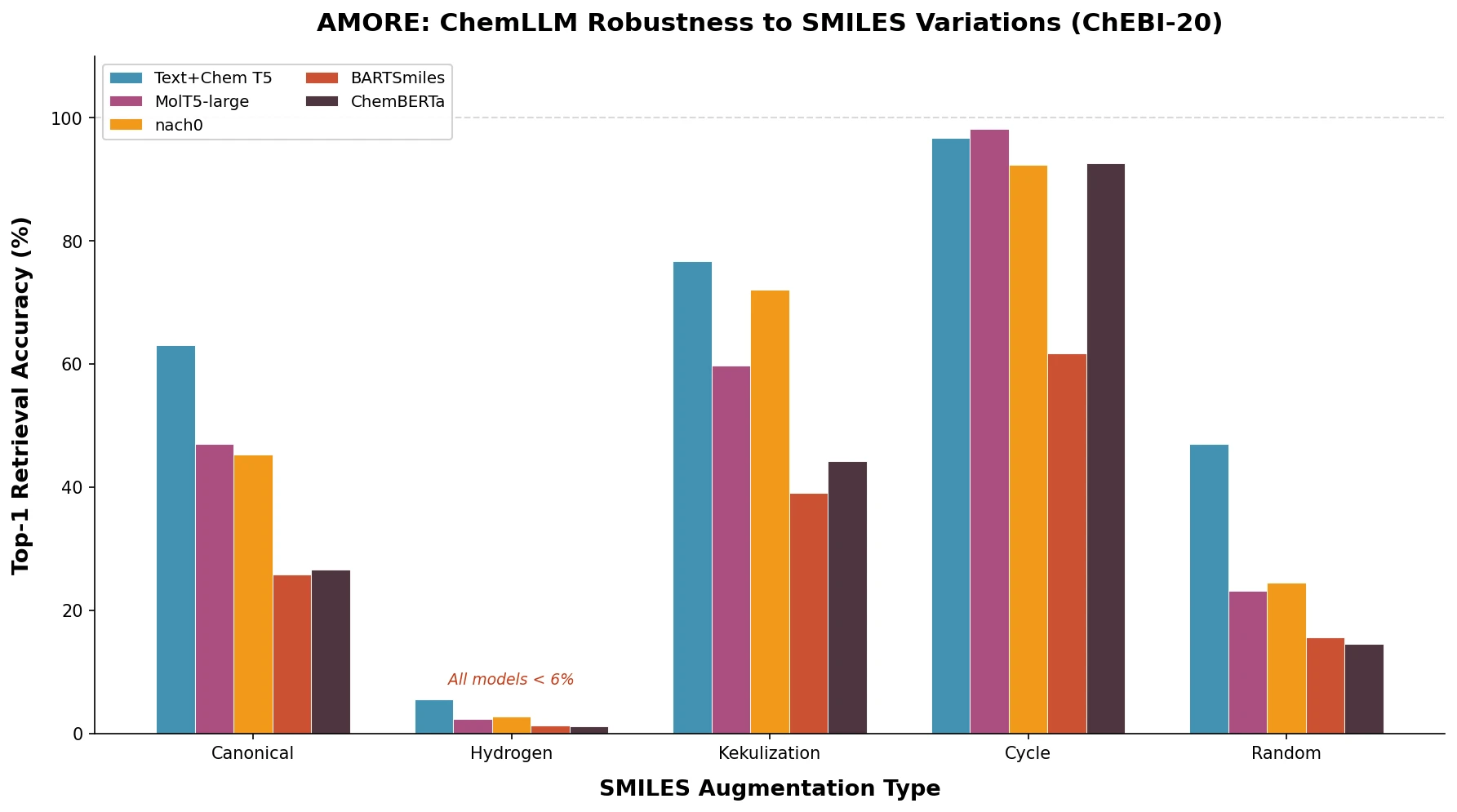
SPMM pre-trains a dual-stream transformer on SMILES and 53 molecular property vectors using contrastive learning and cross-attention, enabling bidirectional structure-property generation, property prediction, and reaction prediction through a single model.