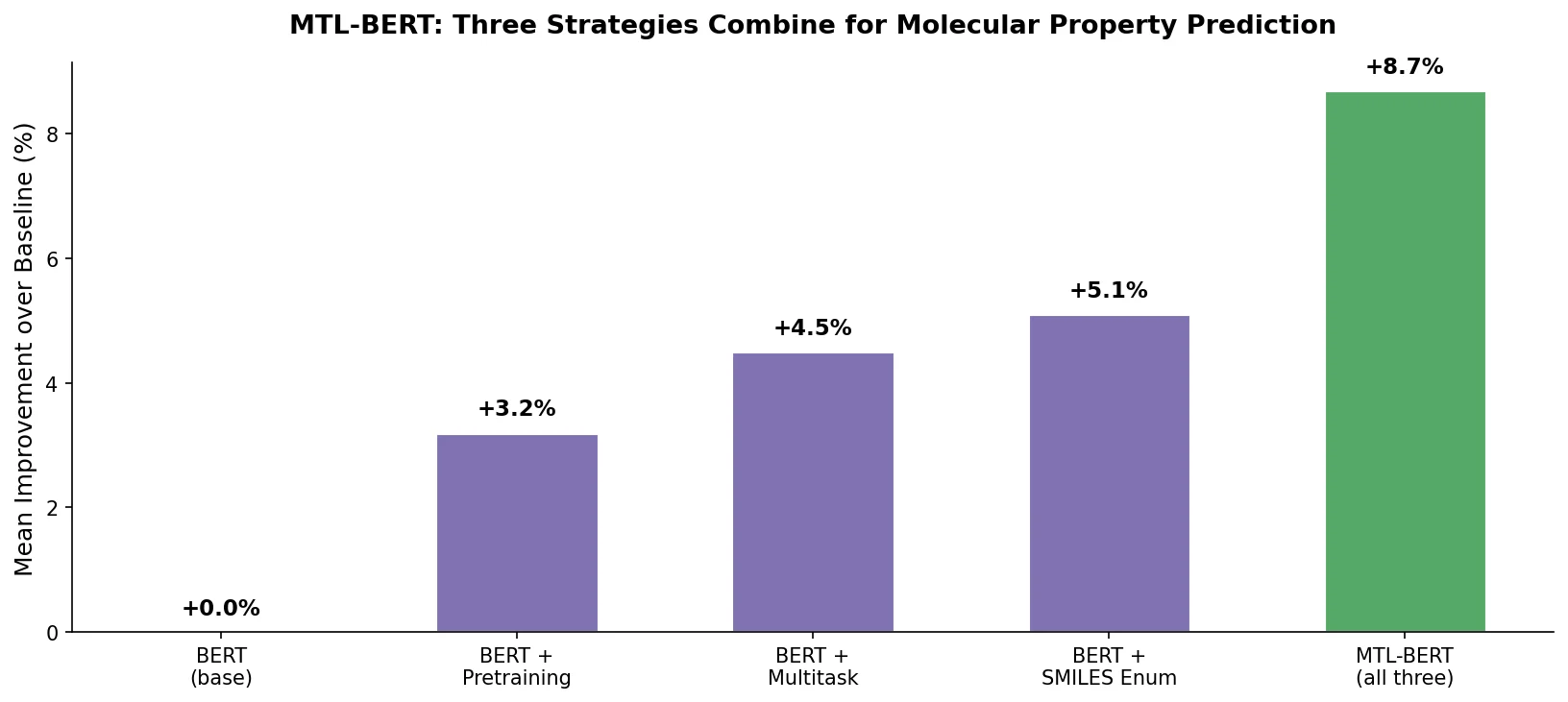
MTL-BERT: Multitask BERT for Property Prediction
MTL-BERT pretrains a BERT model on 1.7M unlabeled SMILES, then fine-tunes jointly on 60 ADMET and molecular property tasks using SMILES enumeration as data augmentation in all phases.

MTL-BERT pretrains a BERT model on 1.7M unlabeled SMILES, then fine-tunes jointly on 60 ADMET and molecular property tasks using SMILES enumeration as data augmentation in all phases.
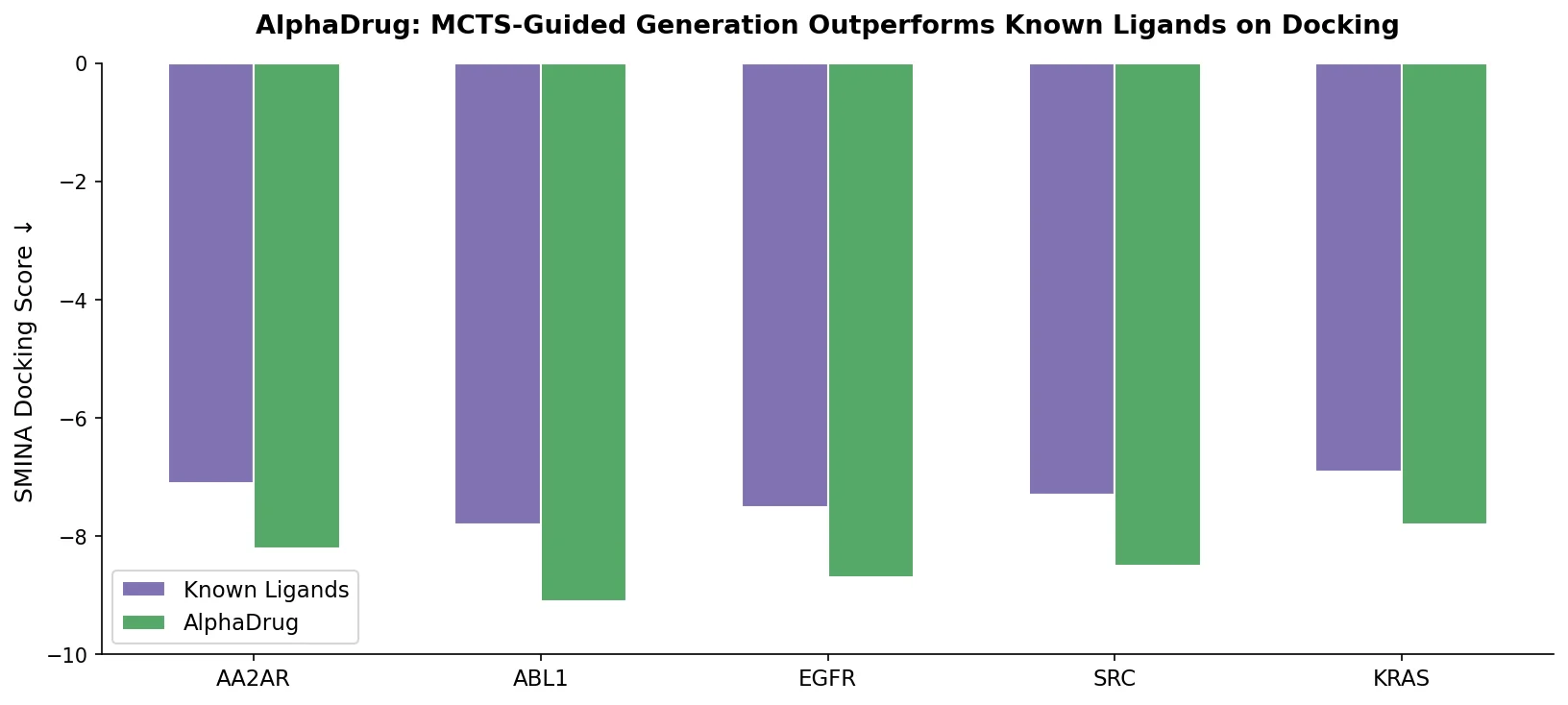
AlphaDrug generates drug candidates for specific protein targets by combining an Lmser Transformer (with hierarchical encoder-decoder skip connections) and Monte Carlo tree search guided by docking scores, achieving higher binding affinities than known ligands on 86% of test proteins.
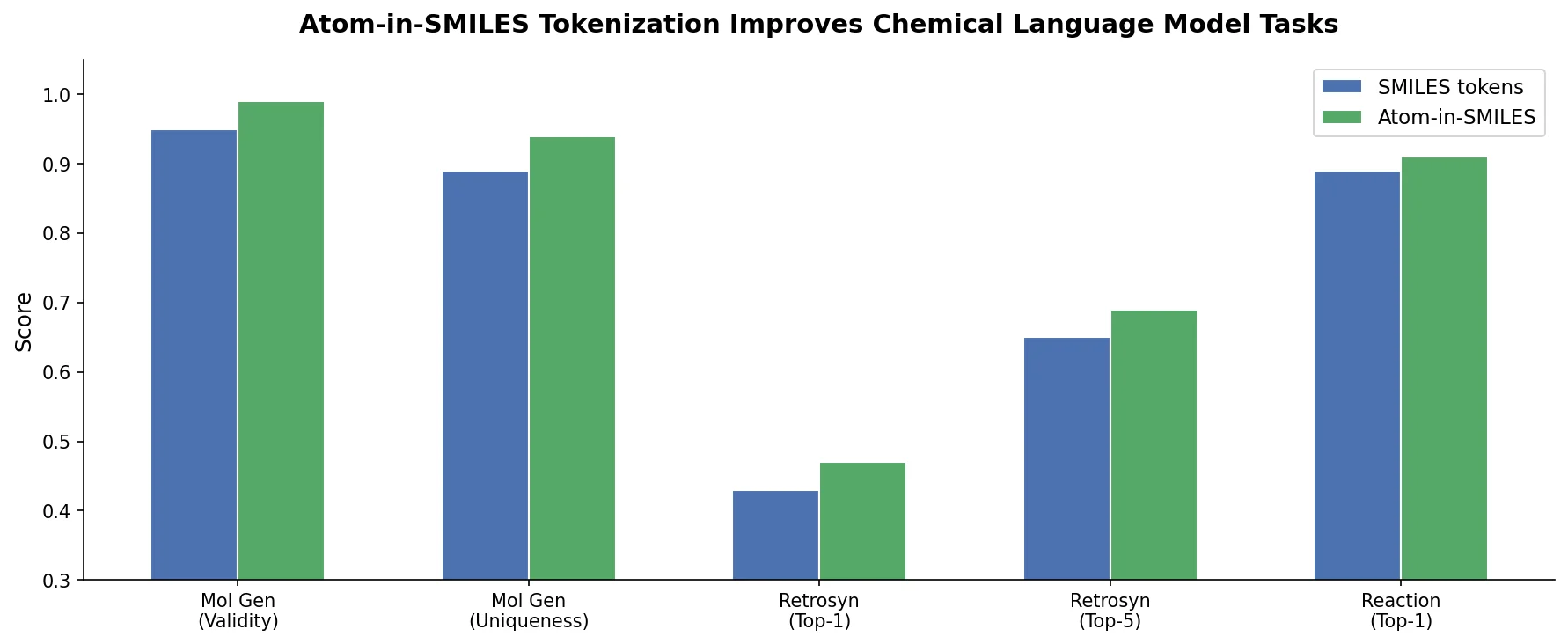
Introduces Atom-in-SMILES (AIS), a tokenization scheme that encodes local chemical environments into SMILES tokens, improving prediction quality across canonicalization, retrosynthesis, and property prediction tasks.
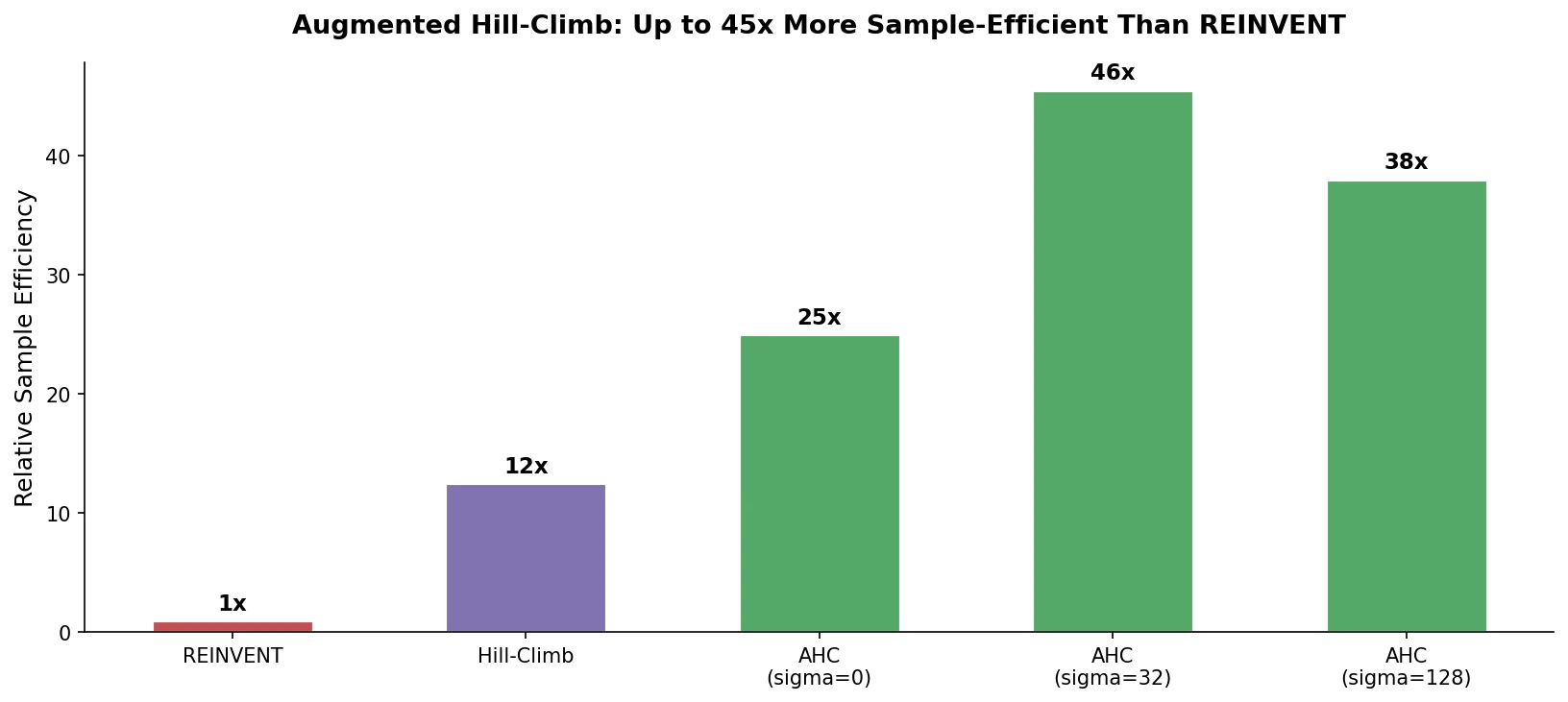
Proposes Augmented Hill-Climb, a hybrid RL strategy for SMILES-based generative models that improves sample efficiency ~45-fold over REINVENT by filtering low-scoring molecules from the loss computation, with diversity filters to prevent mode collapse.
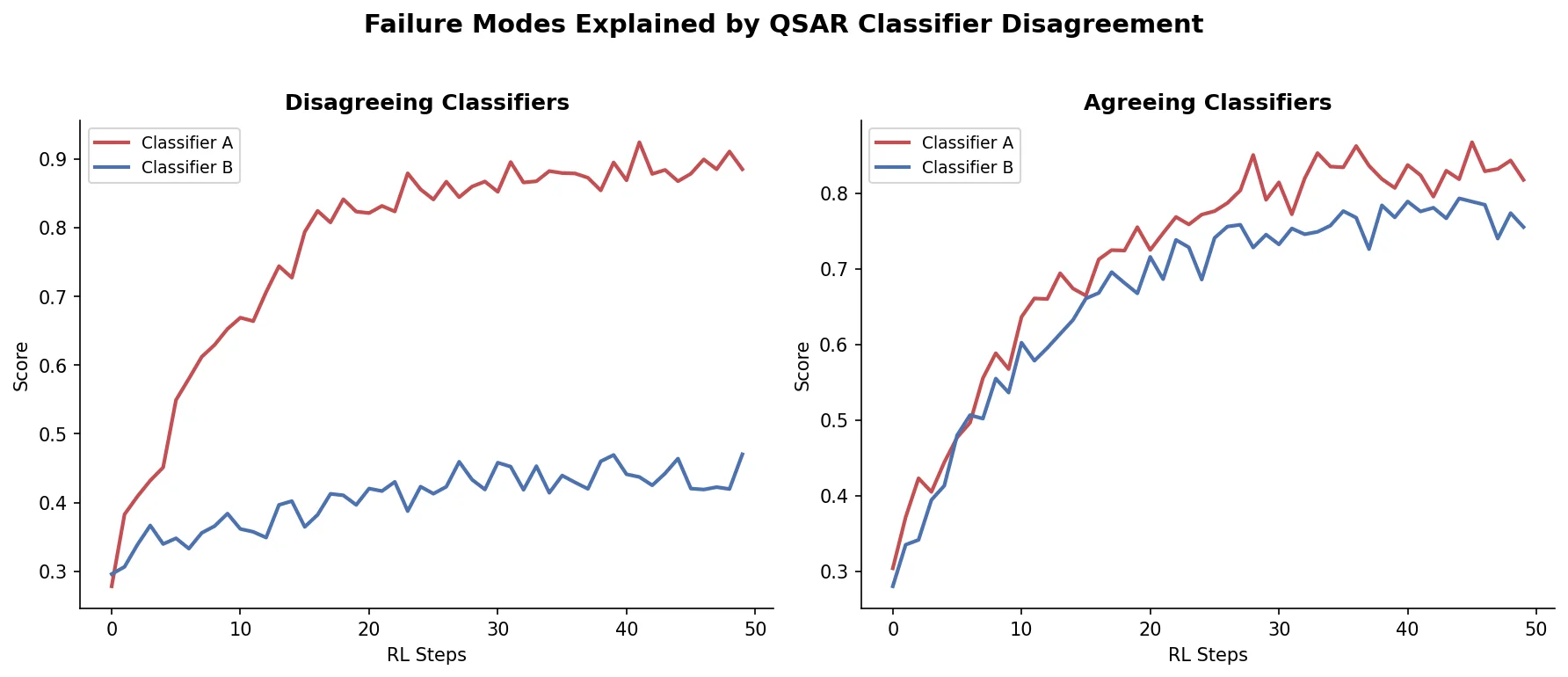
Shows that divergence between optimization and control scores during goal-directed molecular generation is explained by pre-existing disagreement among QSAR models on the training distribution, not by algorithmic exploitation of model-specific biases.
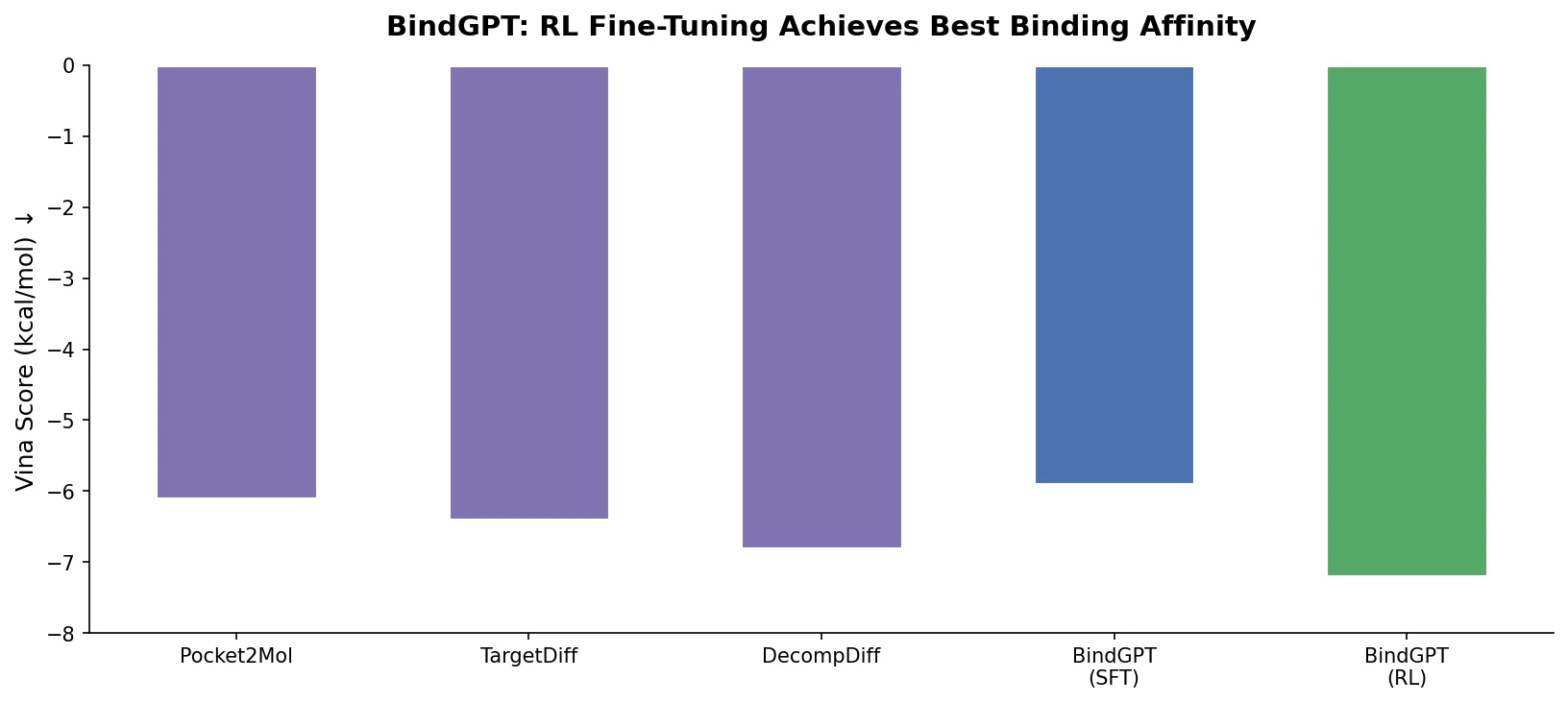
BindGPT formulates 3D molecular design as autoregressive text generation over combined SMILES and XYZ tokens, using large-scale pre-training and reinforcement learning to achieve competitive pocket-conditioned molecule generation.
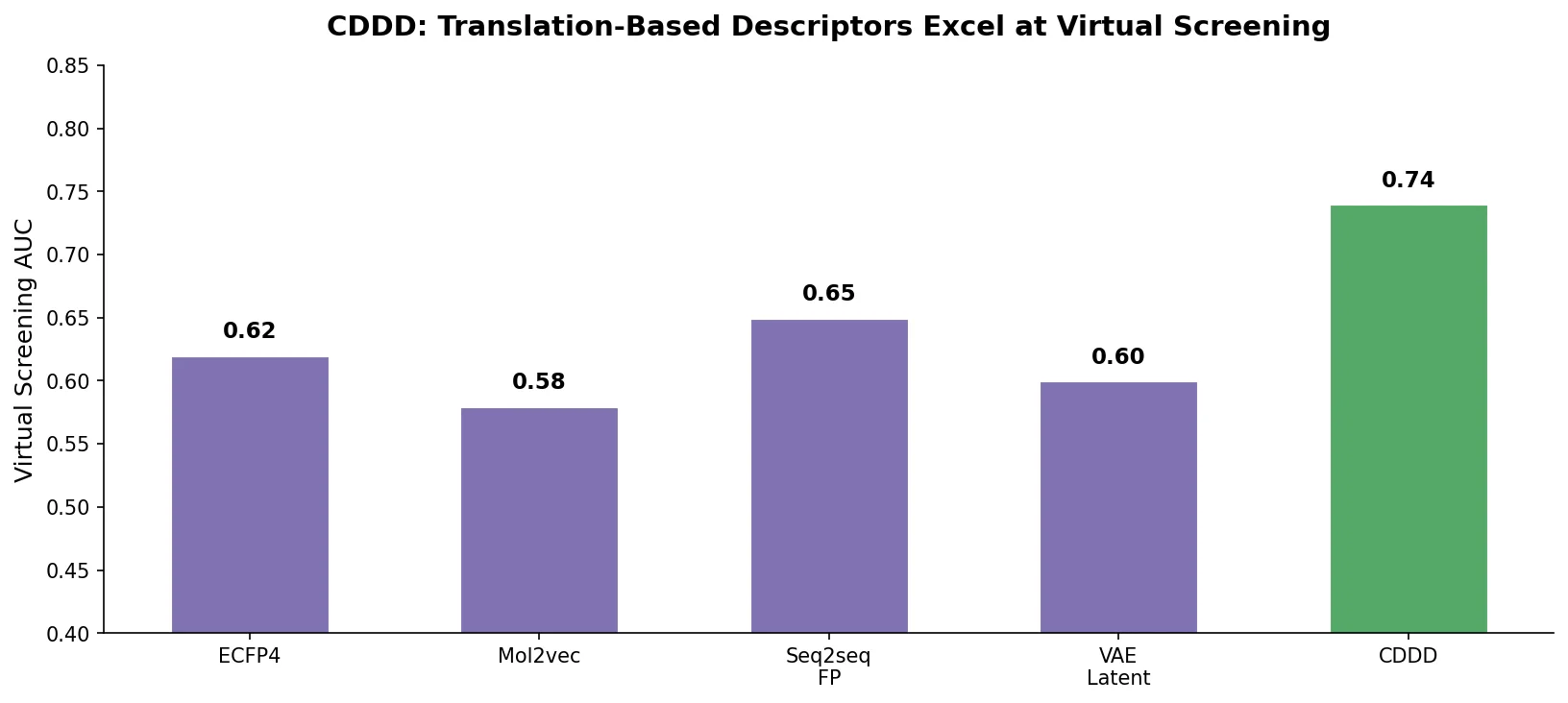
Winter et al. propose CDDD, a translation-based encoder-decoder that learns continuous molecular descriptors by translating between equivalent chemical representations like SMILES and InChI, pretrained on 72 million compounds.
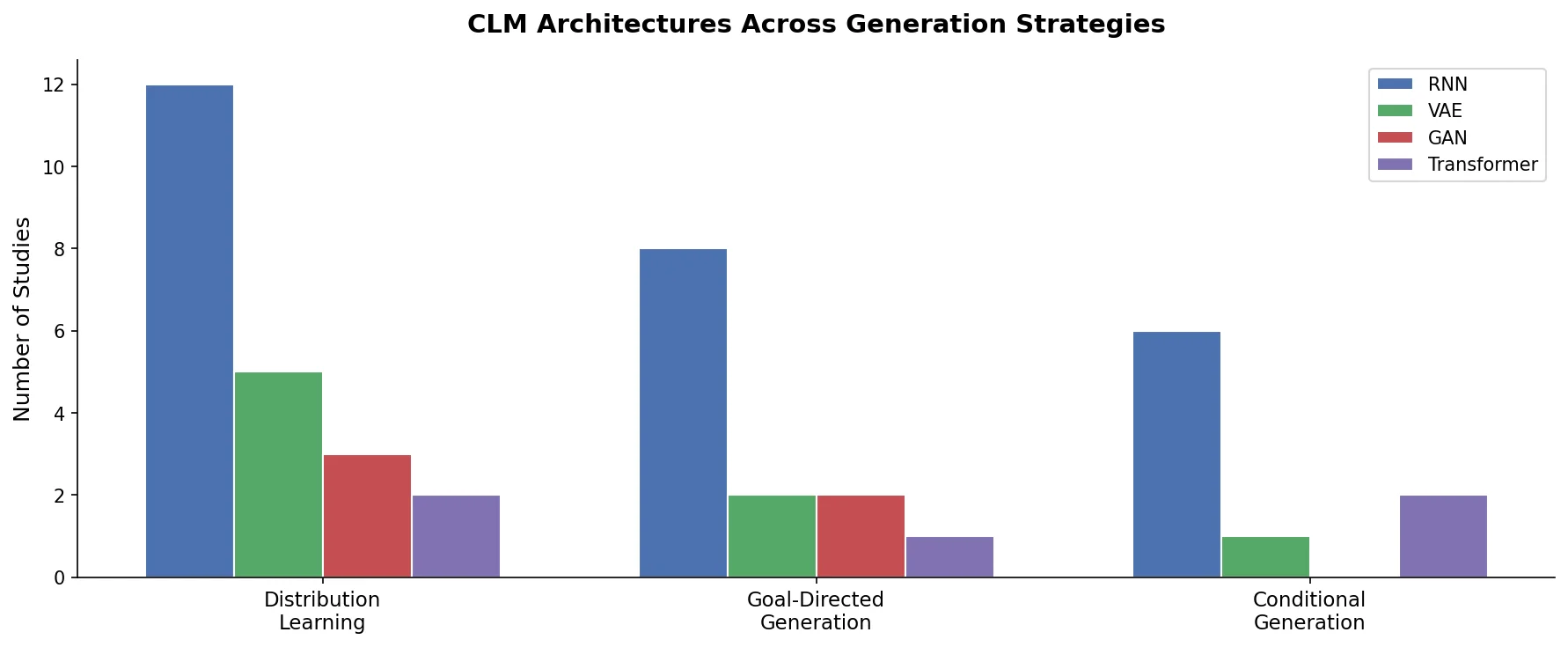
A minireview of chemical language models for de novo molecule design, covering SMILES and SELFIES representations, RNN and Transformer architectures, distribution learning, goal-directed and conditional generation, and prospective experimental validation.
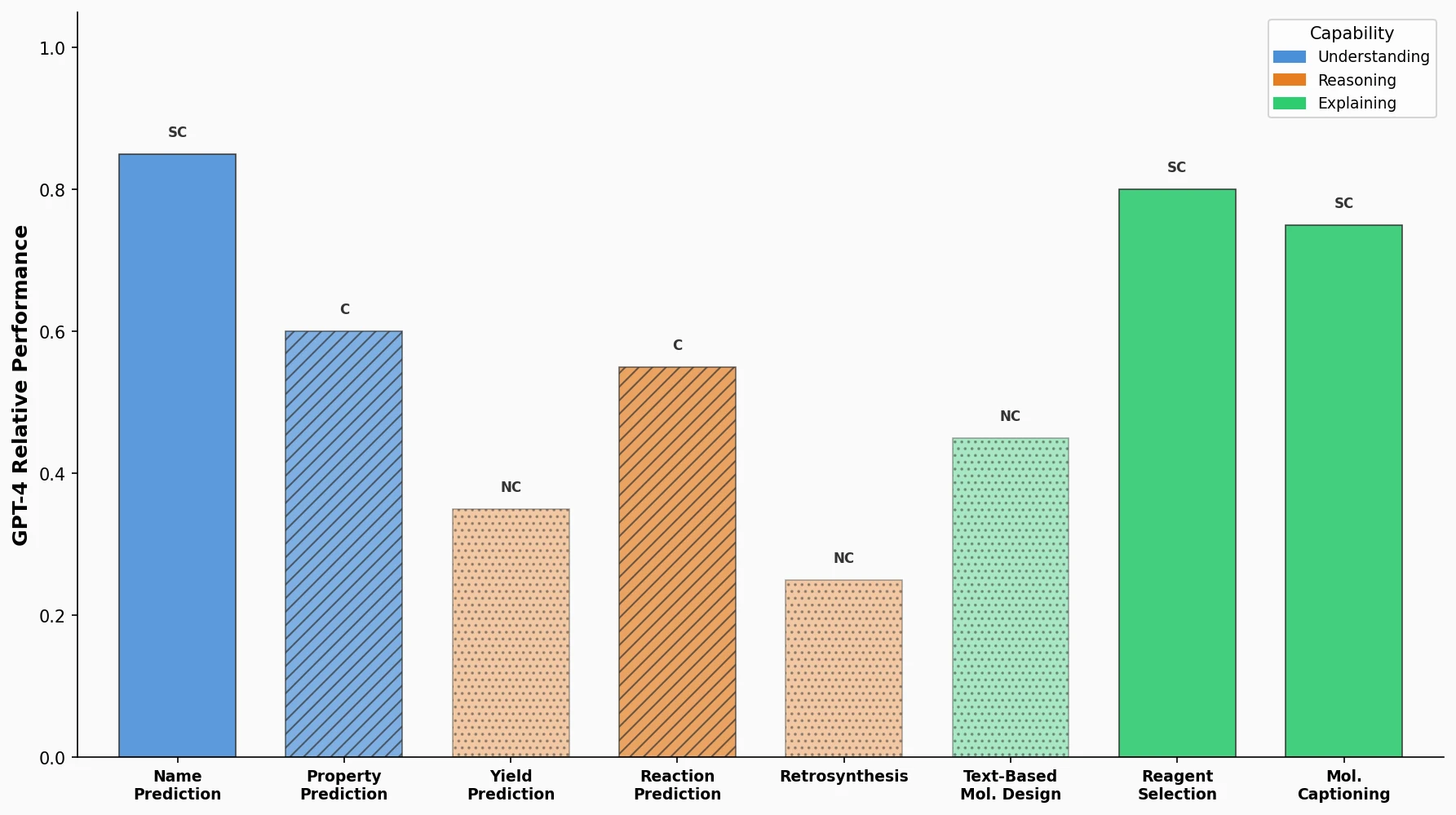
A comprehensive benchmark evaluating GPT-4, GPT-3.5, Davinci-003, Llama, and Galactica on eight practical chemistry tasks, revealing that LLMs are competitive on classification and text tasks but struggle with SMILES-dependent generation.
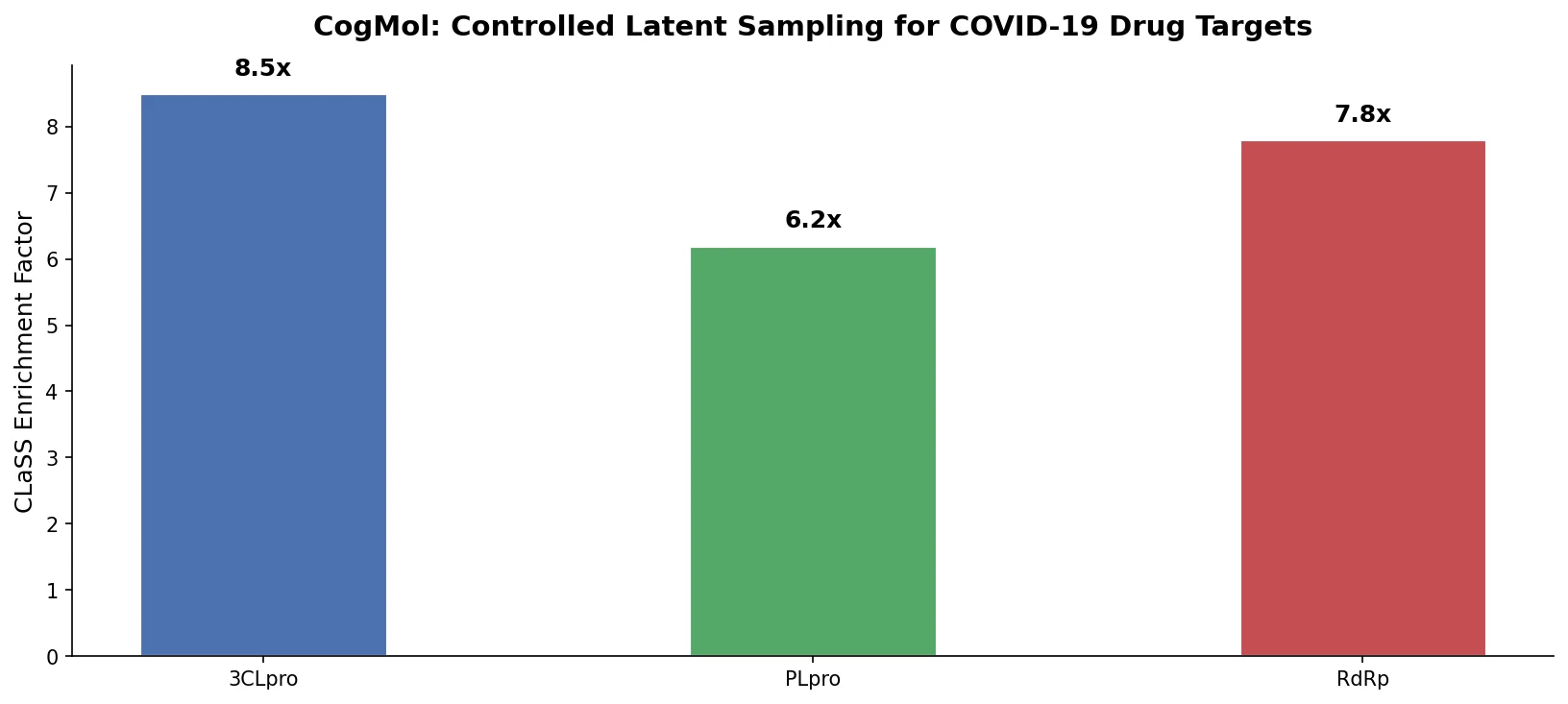
CogMol uses a SMILES VAE and multi-attribute controlled sampling (CLaSS) to generate novel, target-specific drug molecules for unseen SARS-CoV-2 proteins without model retraining.
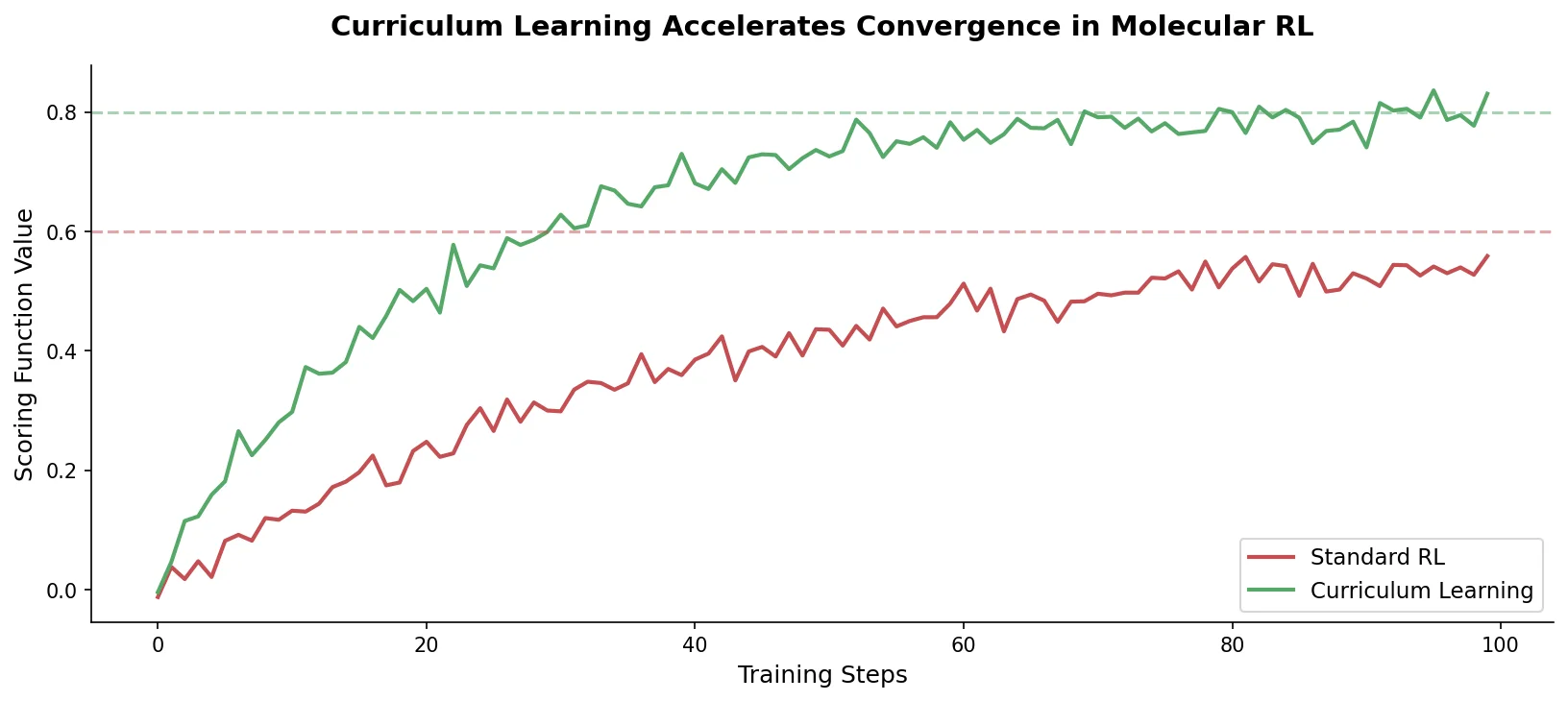
Introduces curriculum learning to the REINVENT de novo design platform, decomposing complex drug design objectives into simpler sequential tasks that accelerate agent convergence and improve output quality over standard reinforcement learning.
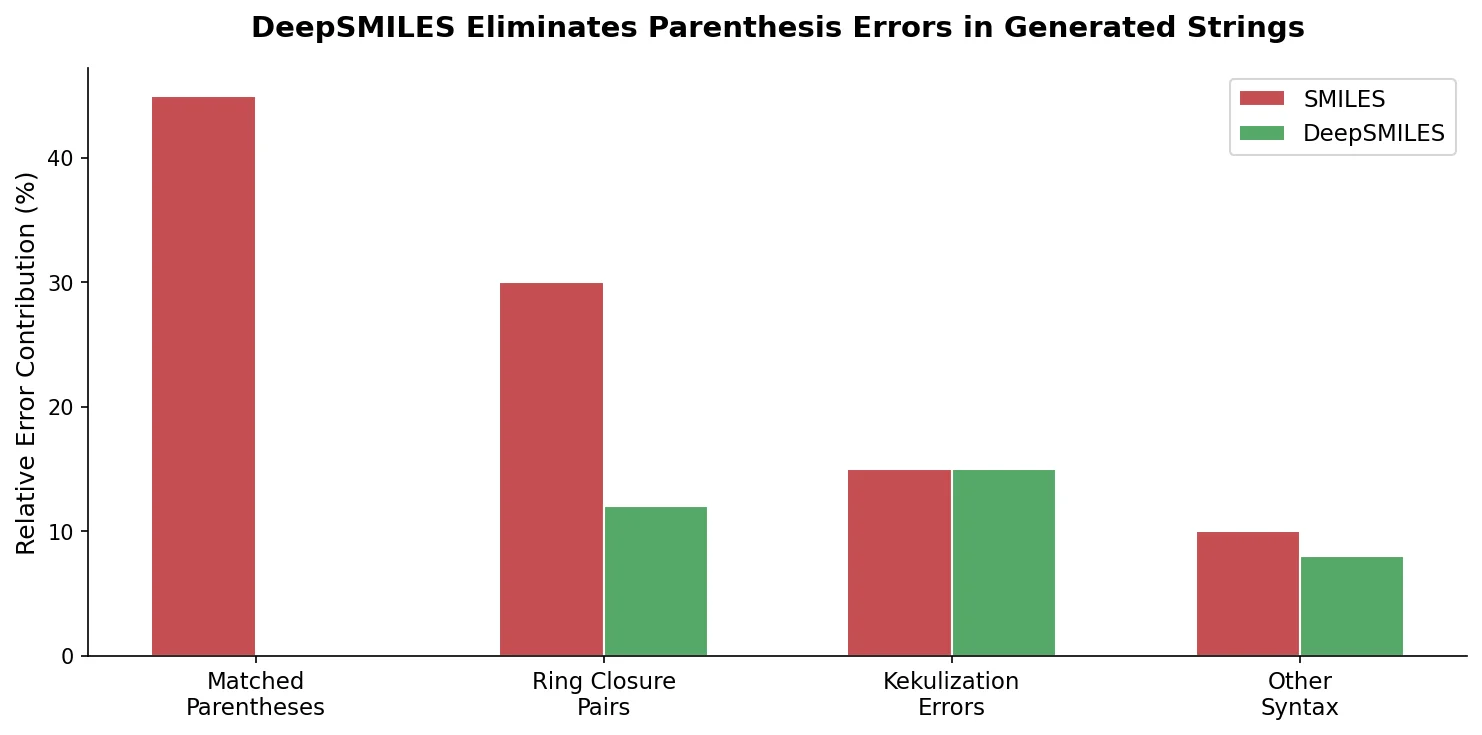
DeepSMILES replaces paired parentheses and ring closure symbols in SMILES with a postfix notation and single ring-size digits, making it easier for generative models to produce syntactically valid molecular strings.