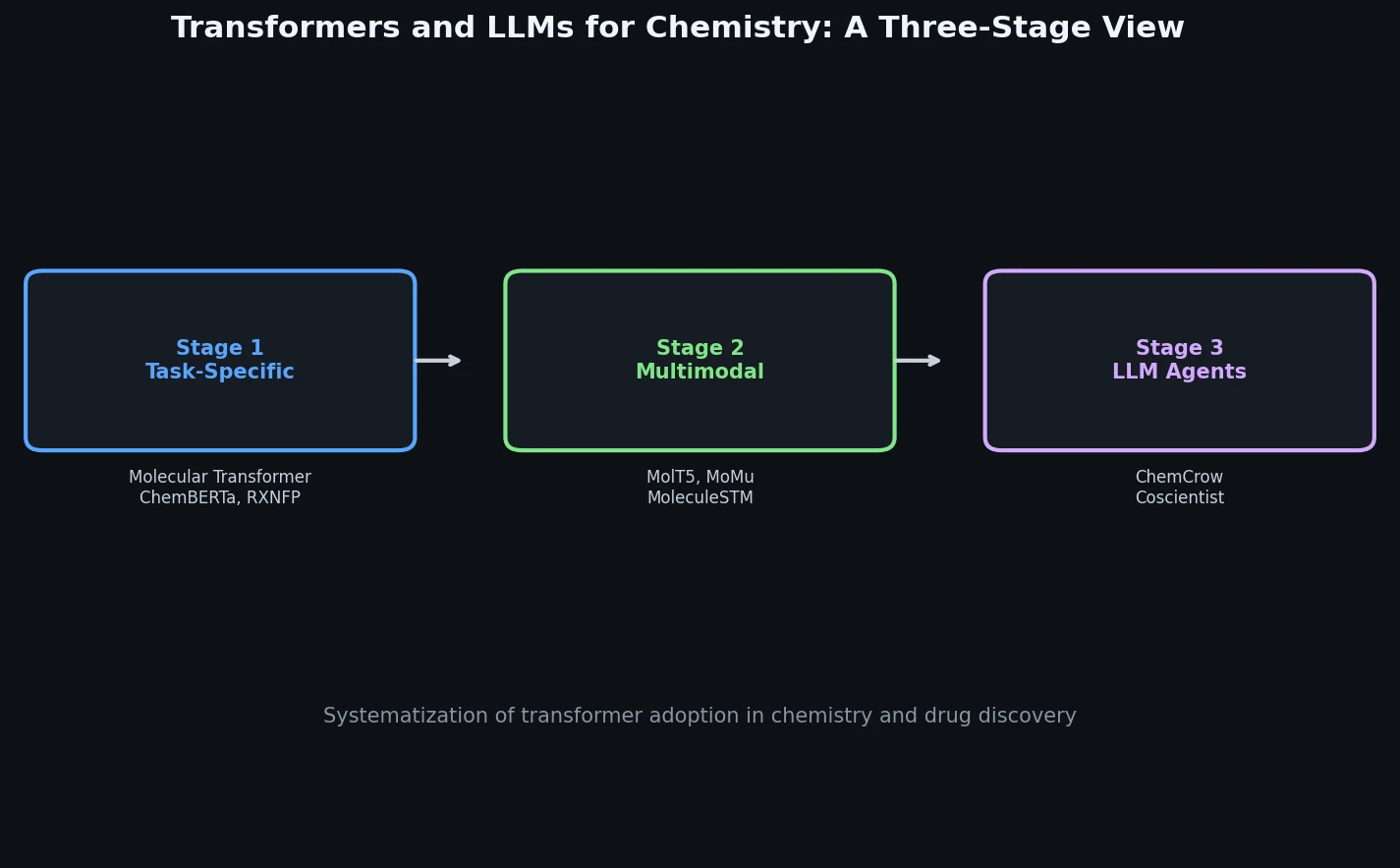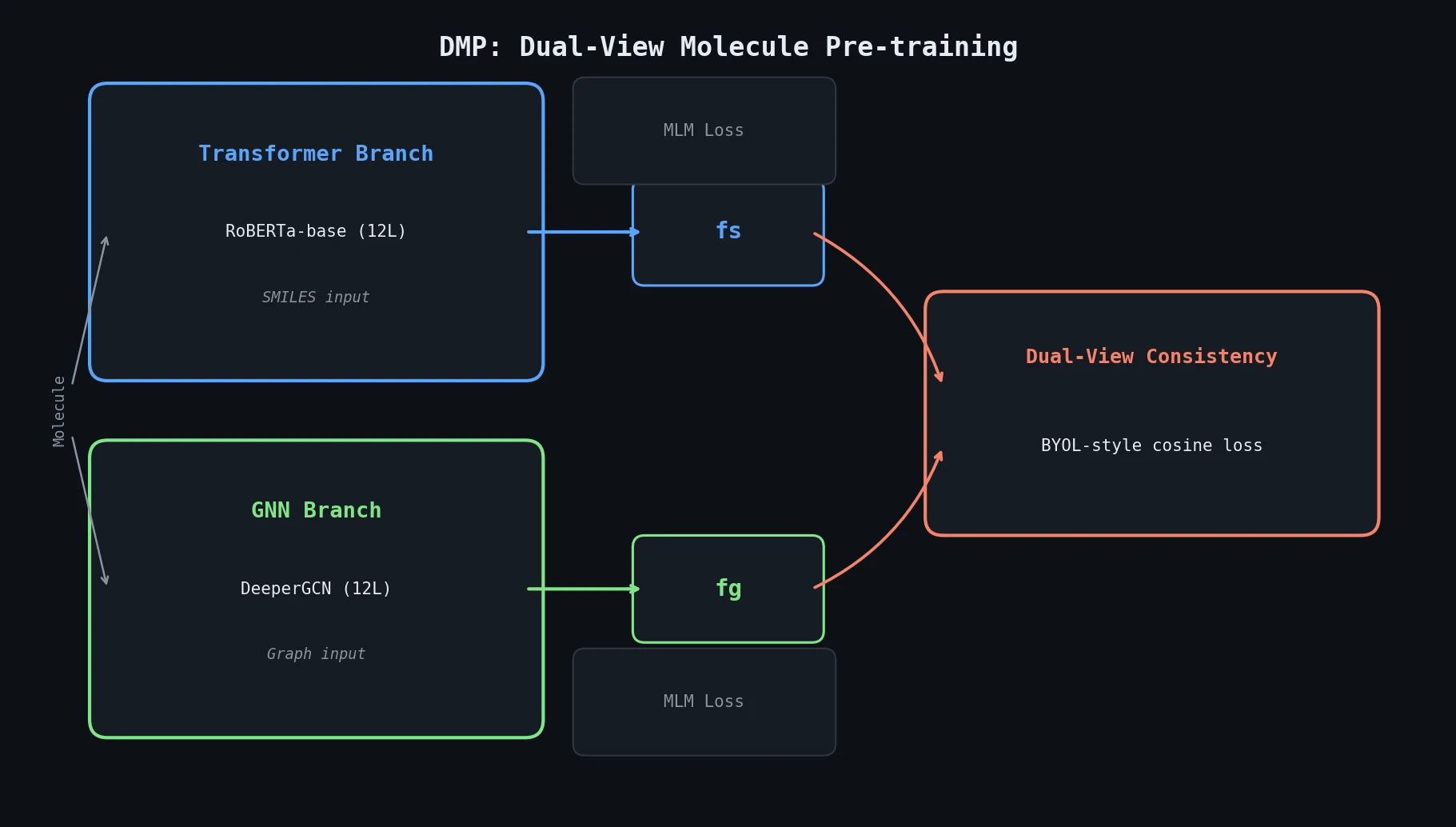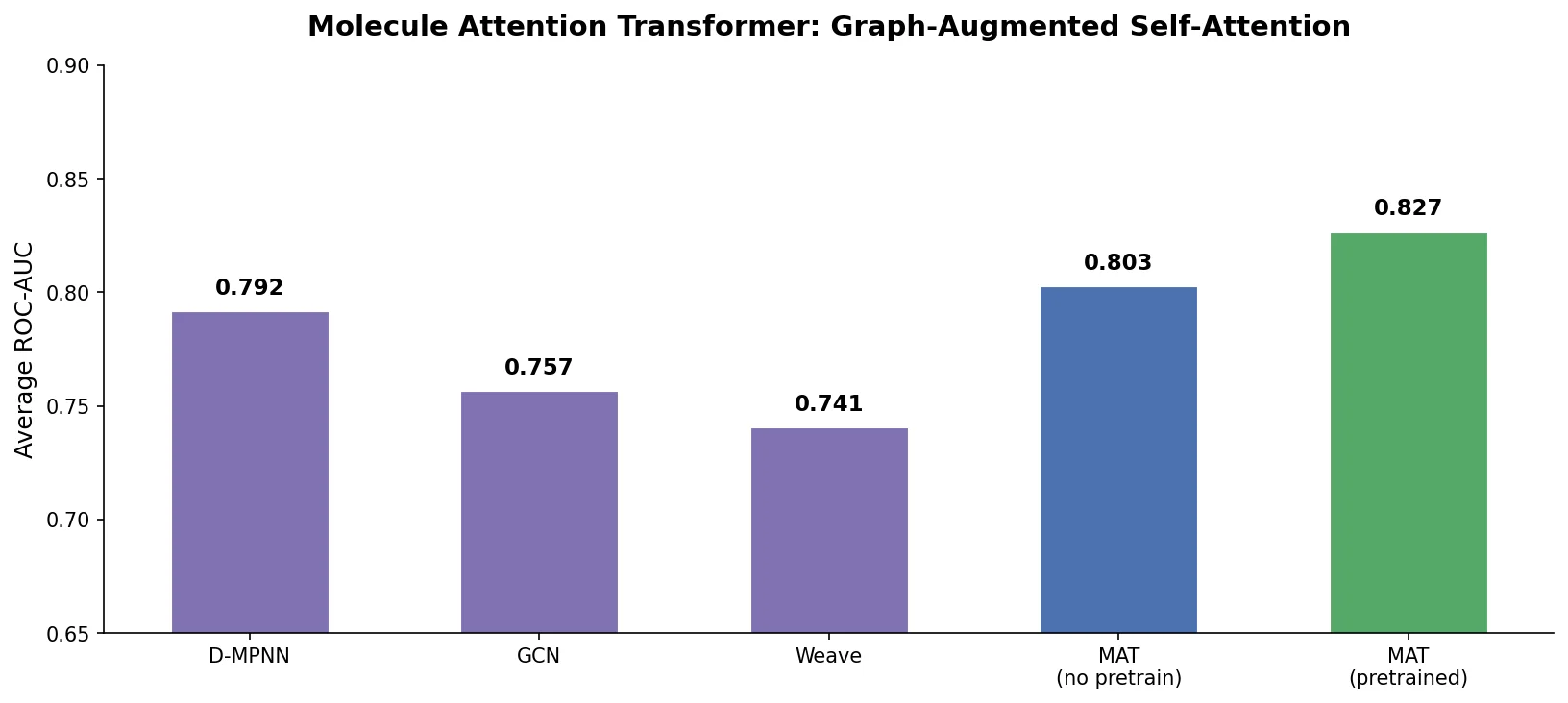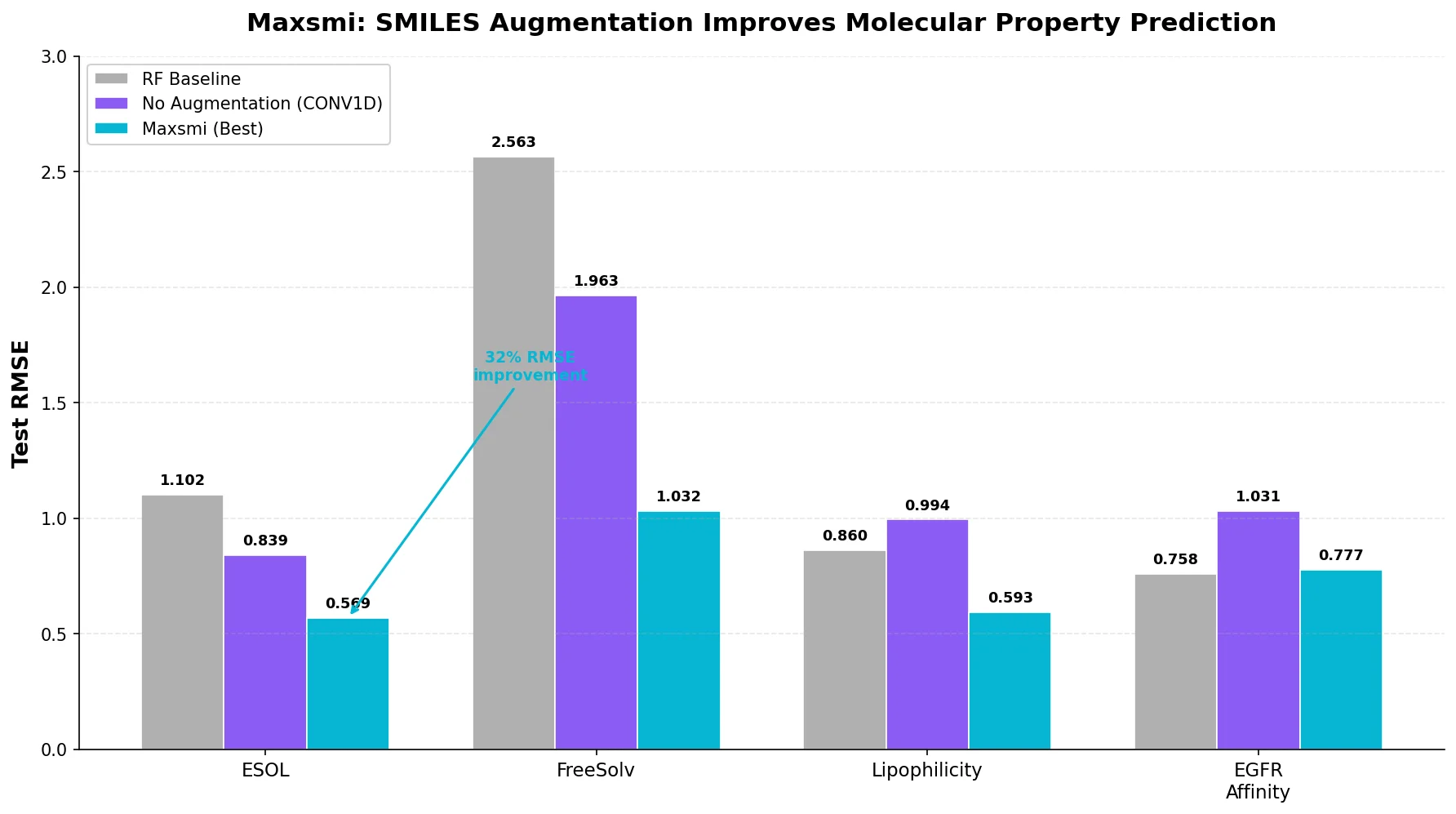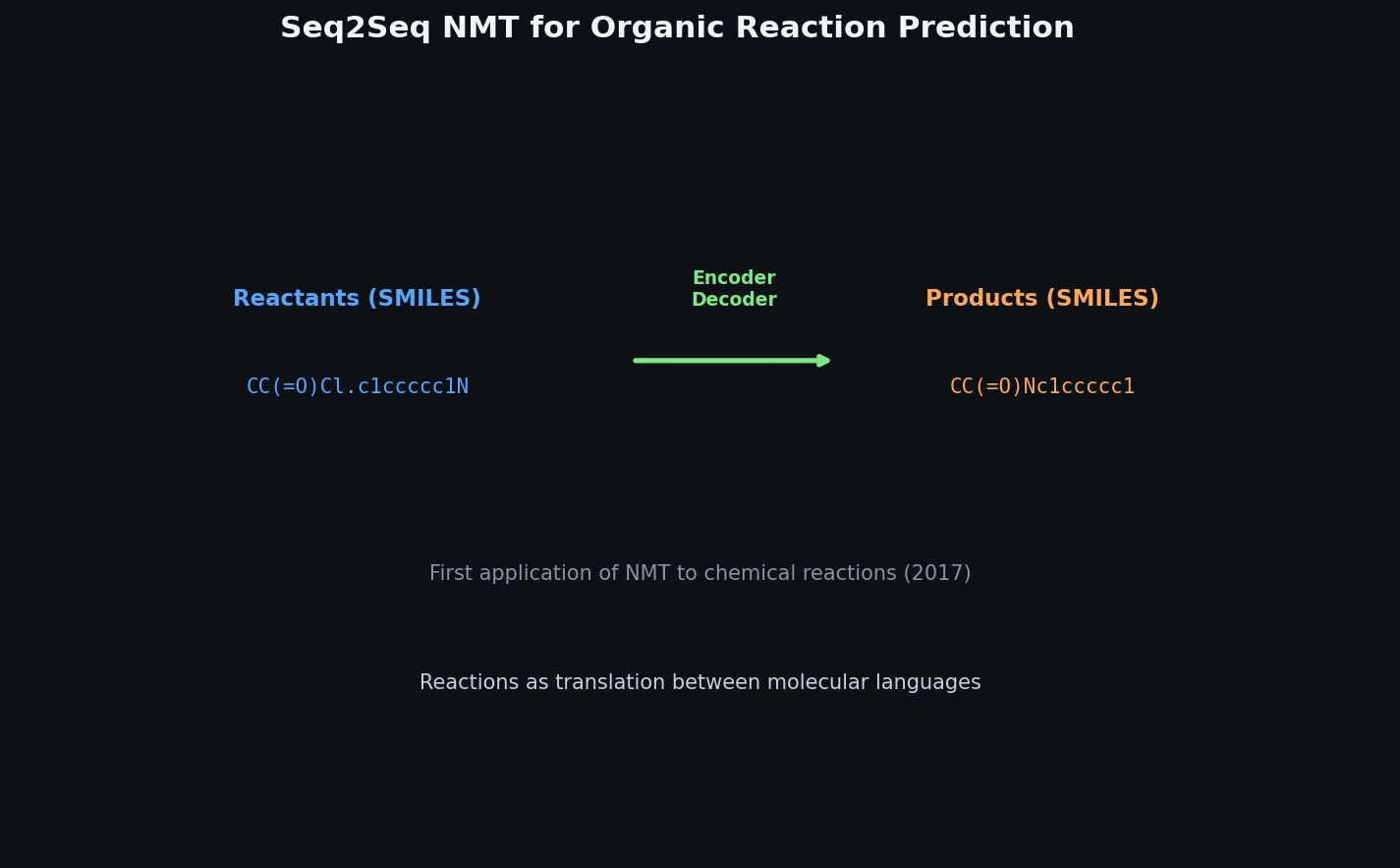
Neural Machine Translation for Reaction Prediction
This 2016 paper first proposed treating organic reaction prediction as a neural machine translation problem, using a GRU-based sequence-to-sequence model with attention to translate reactant SMILES strings into product SMILES strings.