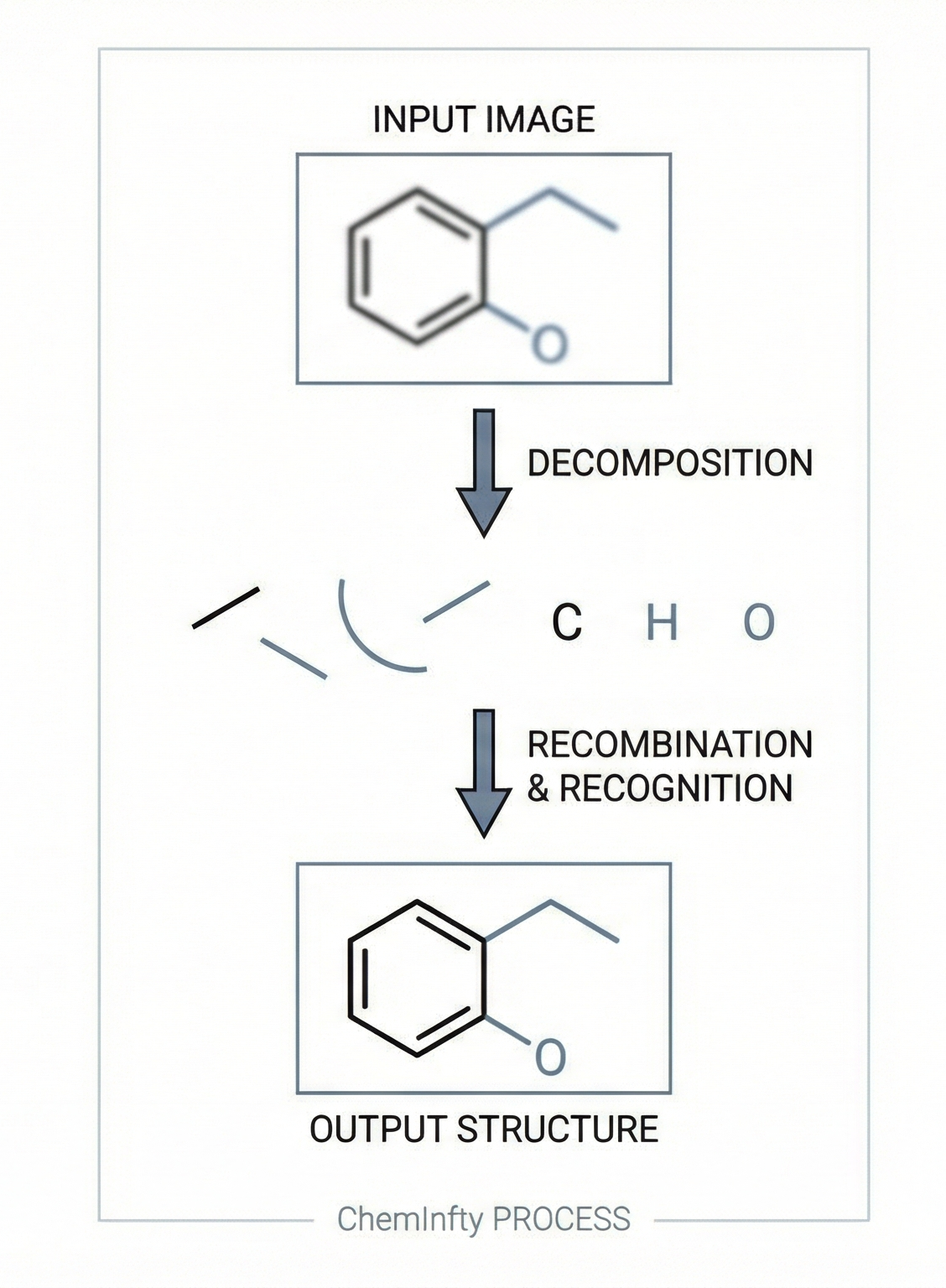
ChemInfty: Chemical Structure Recognition in Patent Images
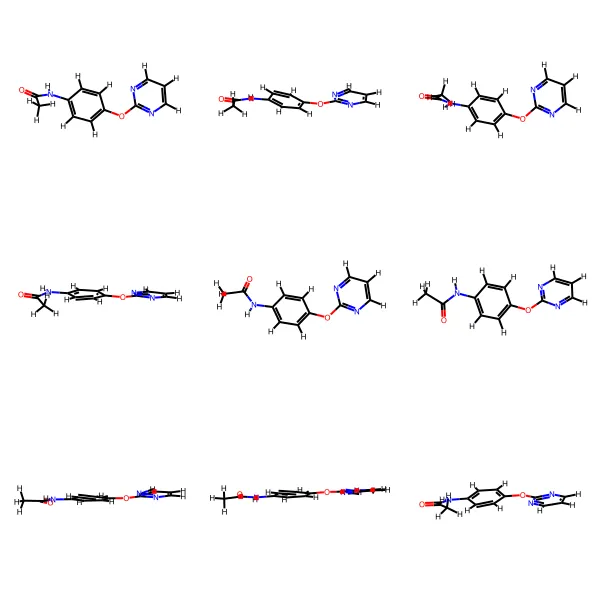
A 2011 rule-based OCSR system designed specifically for the challenging low-quality images in Japanese patent applications, using segment-based methods to handle pervasive problems like touching characters, merged atom labels with bonds, and broken lines.