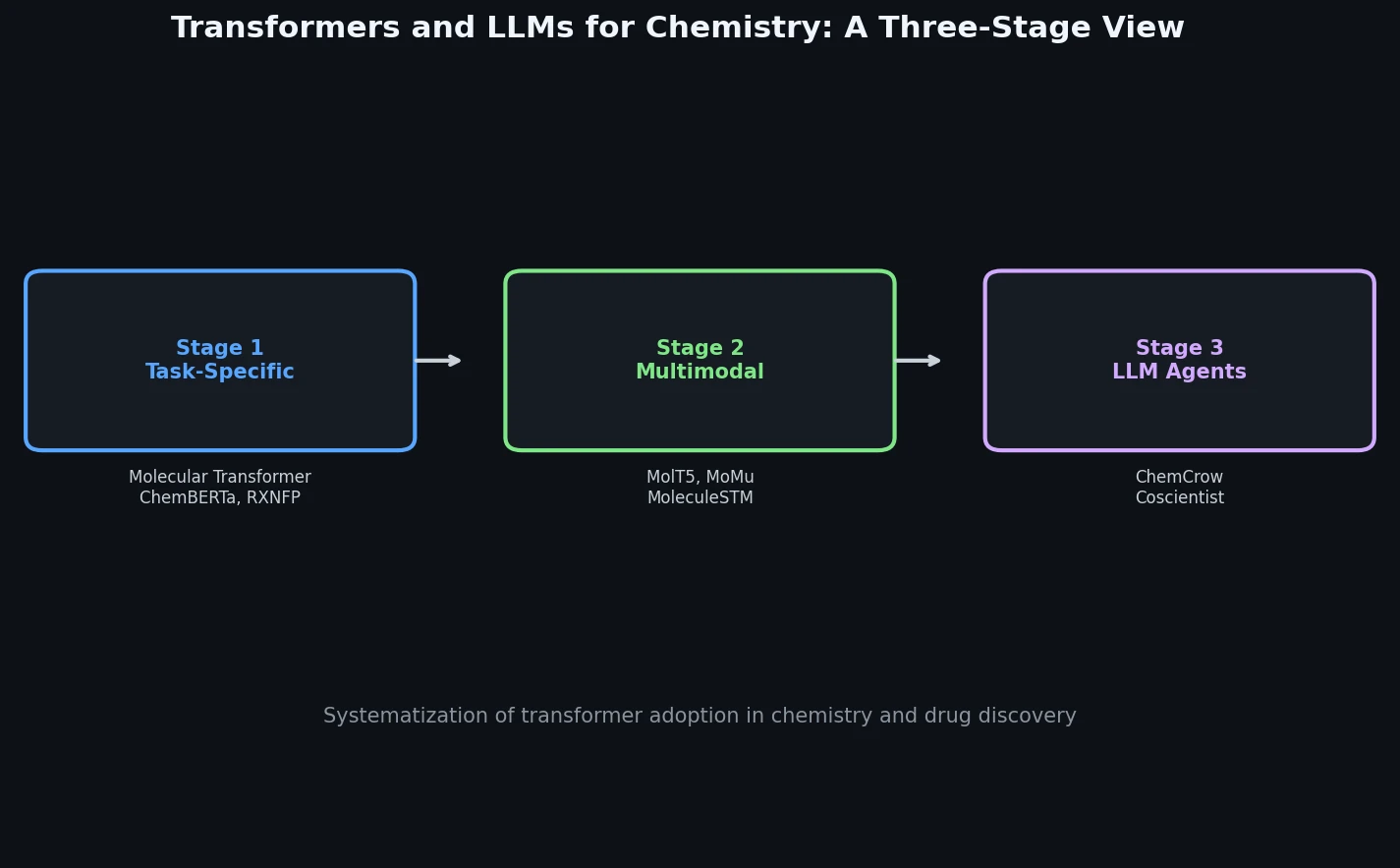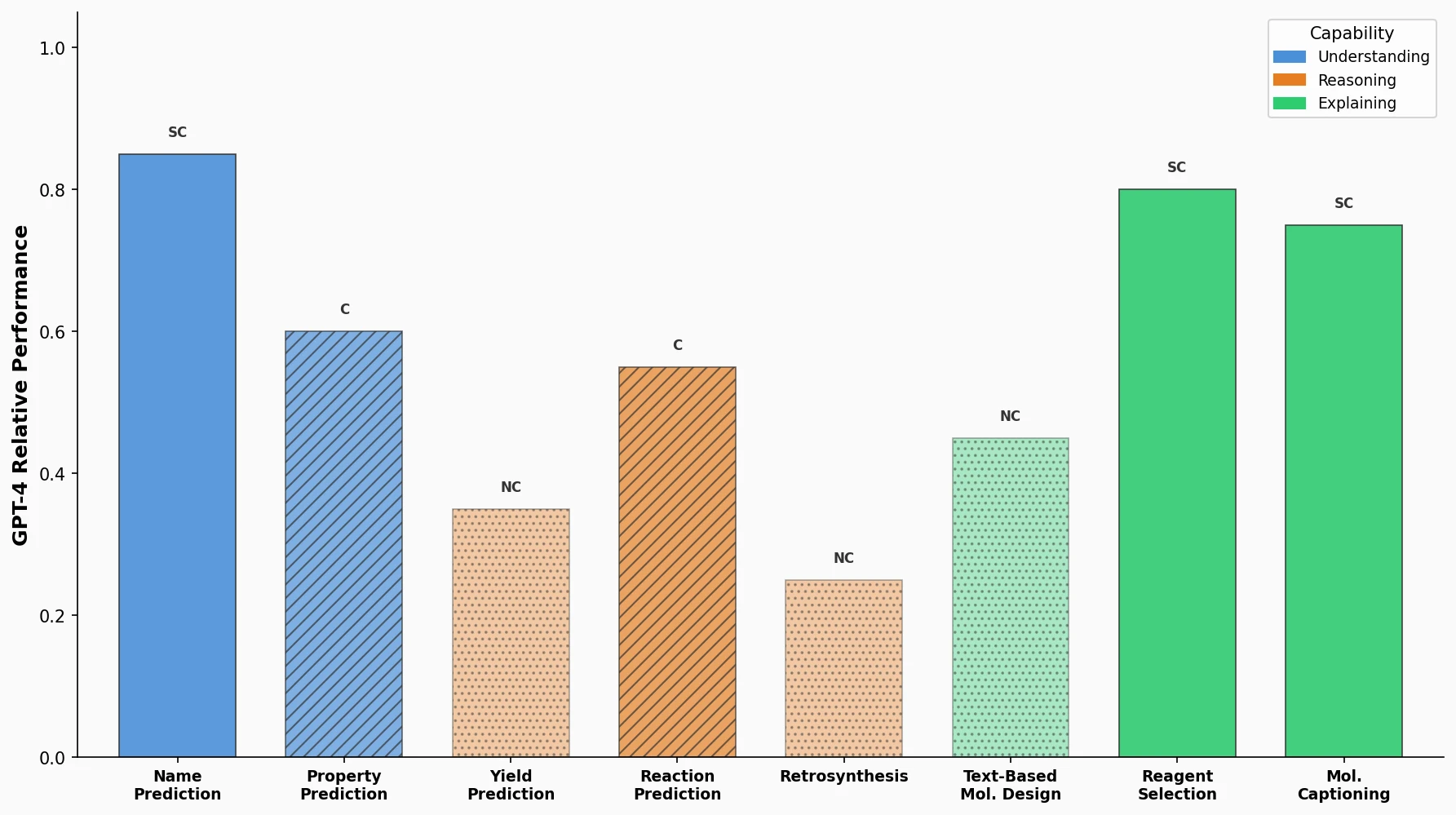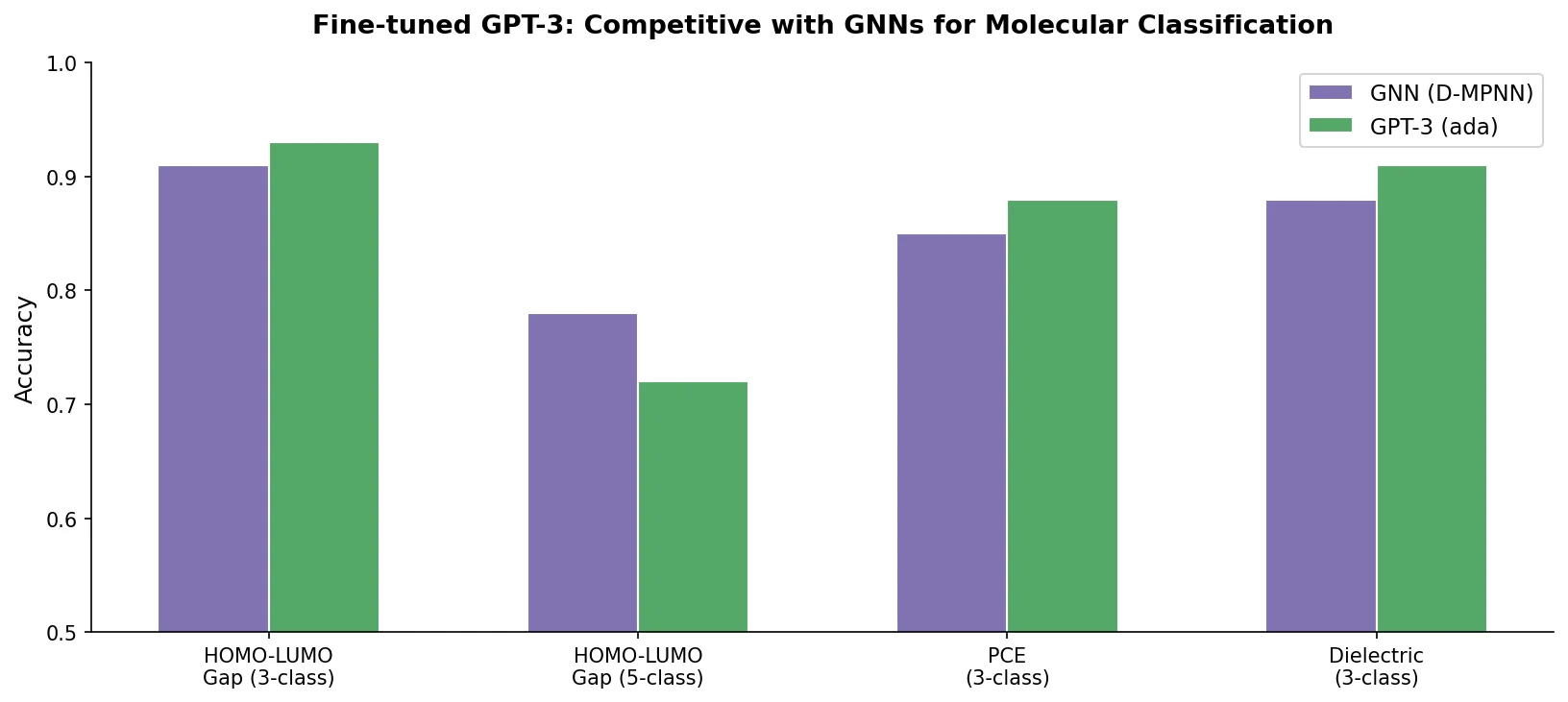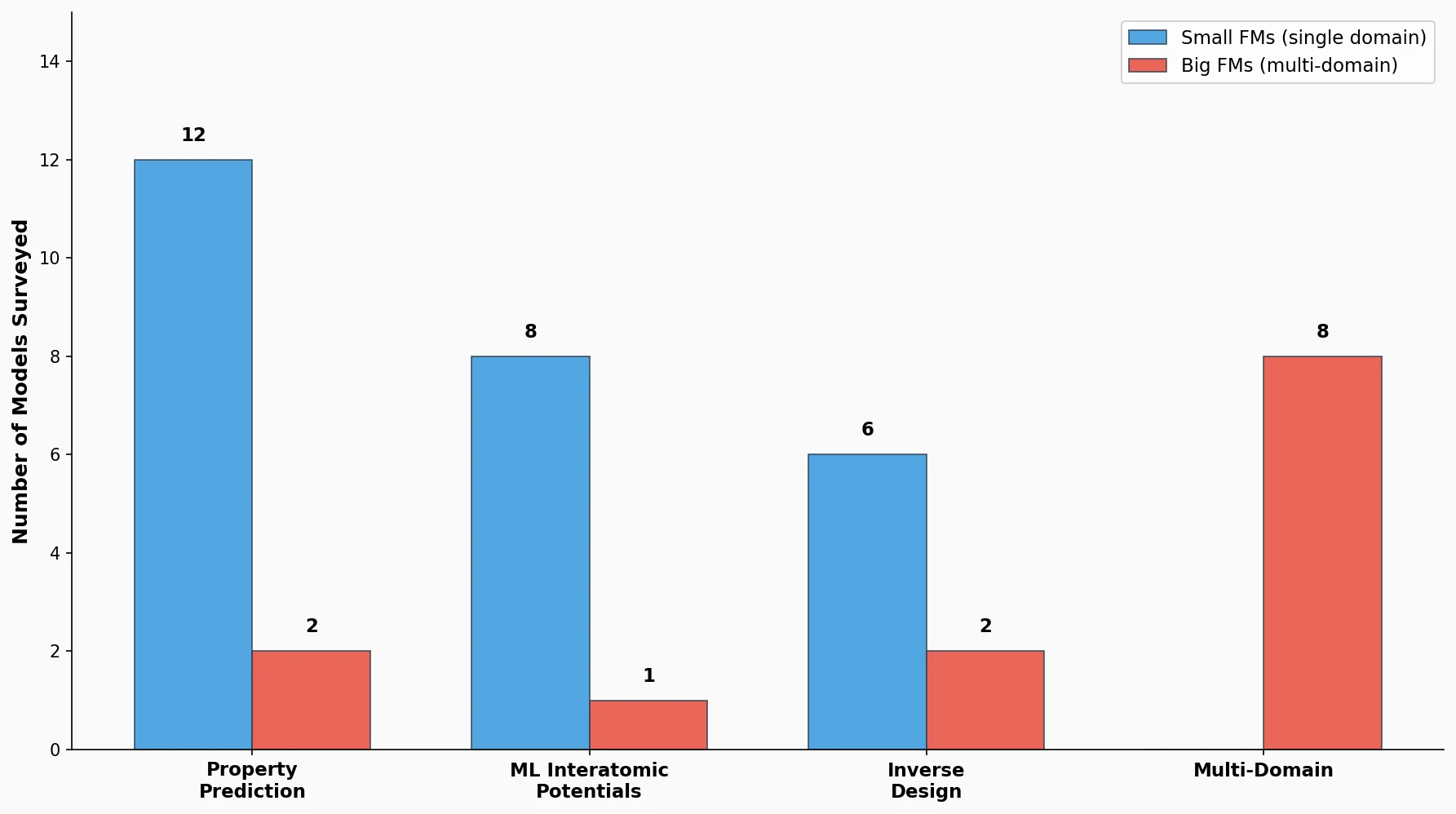
LlaSMol: Instruction-Tuned LLMs for Chemistry Tasks
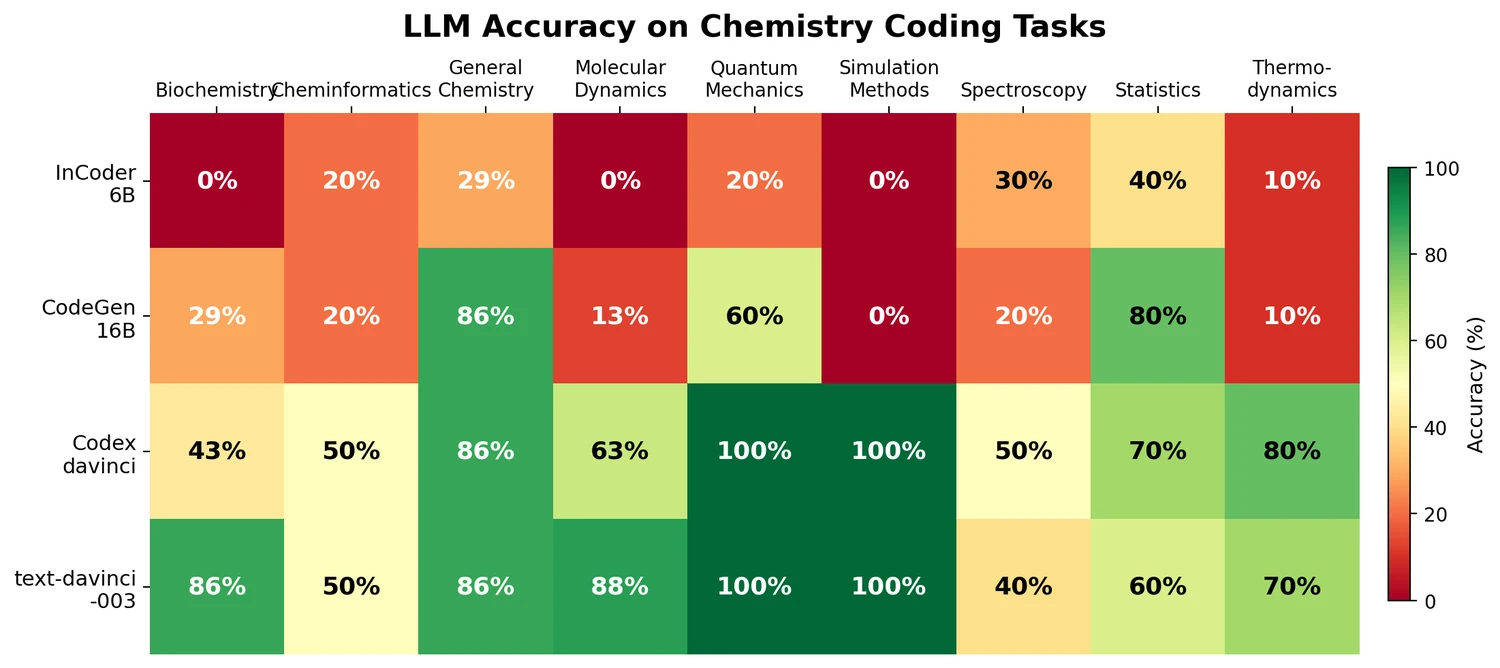
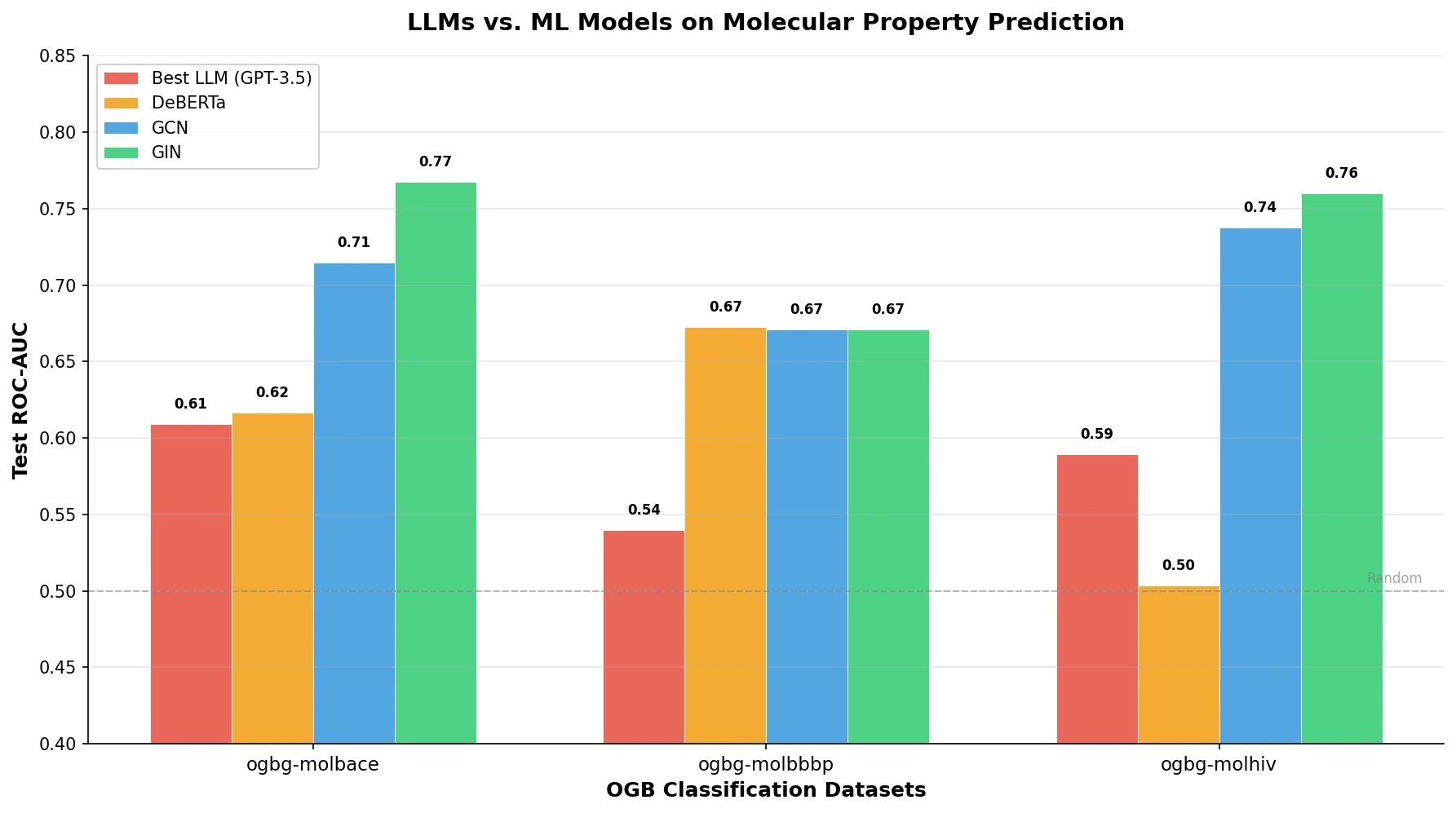
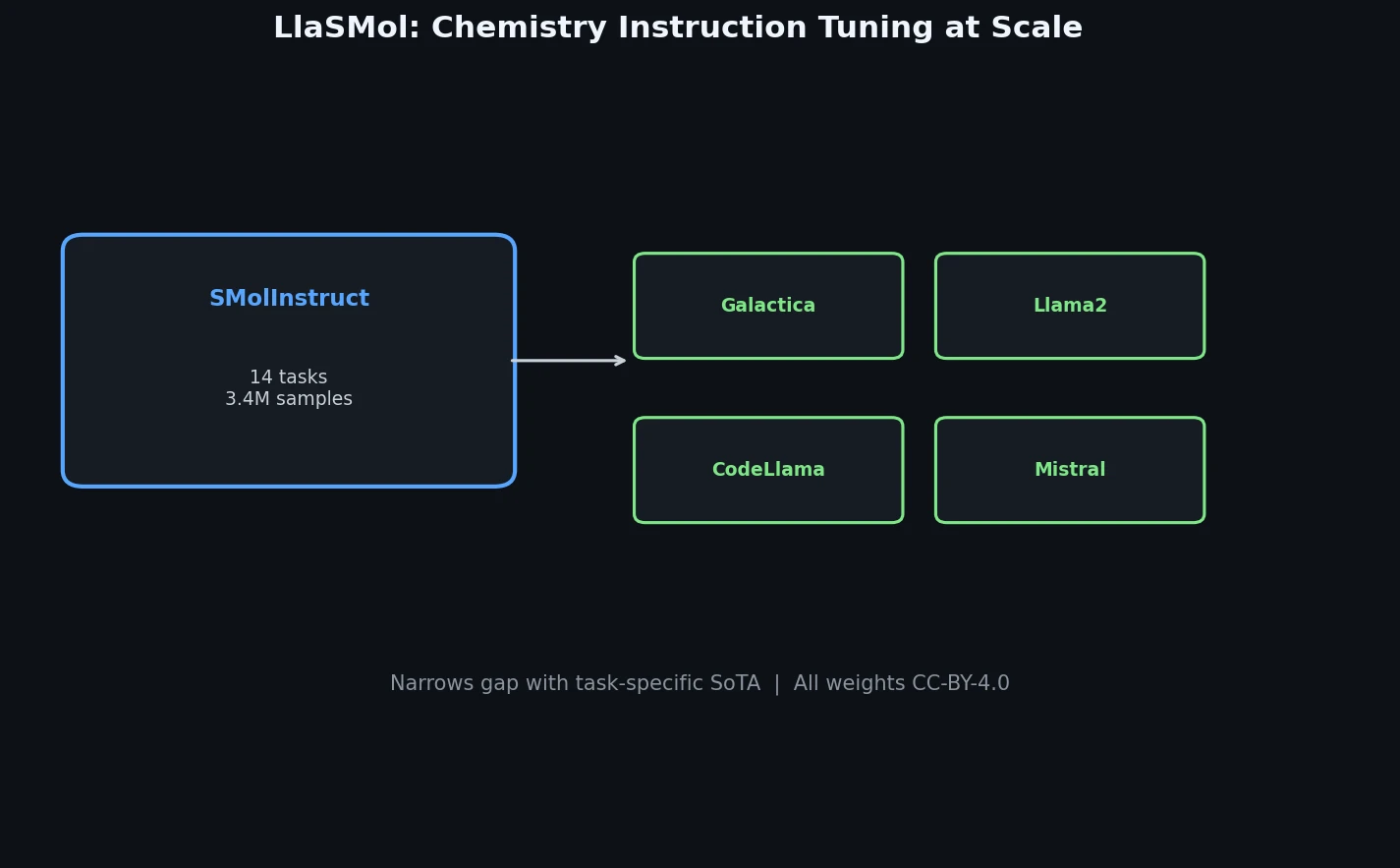
LlaSMol fine-tunes Mistral, Llama 2, and other open-source LLMs on SMolInstruct, a 3.3M-sample instruction tuning dataset covering 14 chemistry tasks. The Mistral-based model outperforms GPT-4 and Claude 3 Opus across all tasks.