DrugChat: Conversational QA on Drug Molecule Graphs
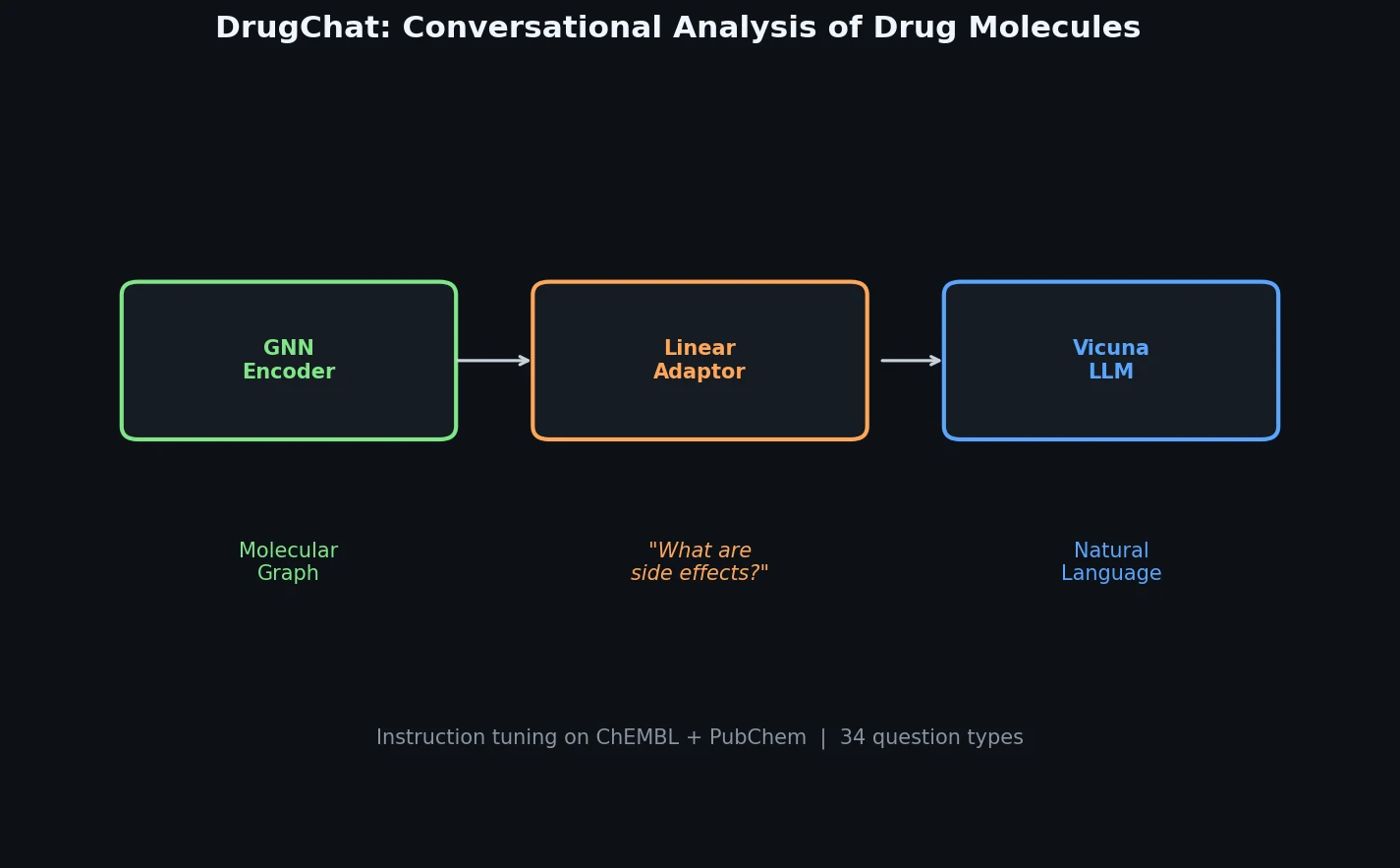
DrugChat is a prototype system that bridges molecular graph neural networks with large language models for interactive, multi-turn question answering about drug compounds. It trains only a lightweight linear adaptor between a frozen GNN encoder and Vicuna-13B using 143K curated QA pairs from ChEMBL and PubChem.