MICER: Molecular Image Captioning with Transfer Learning
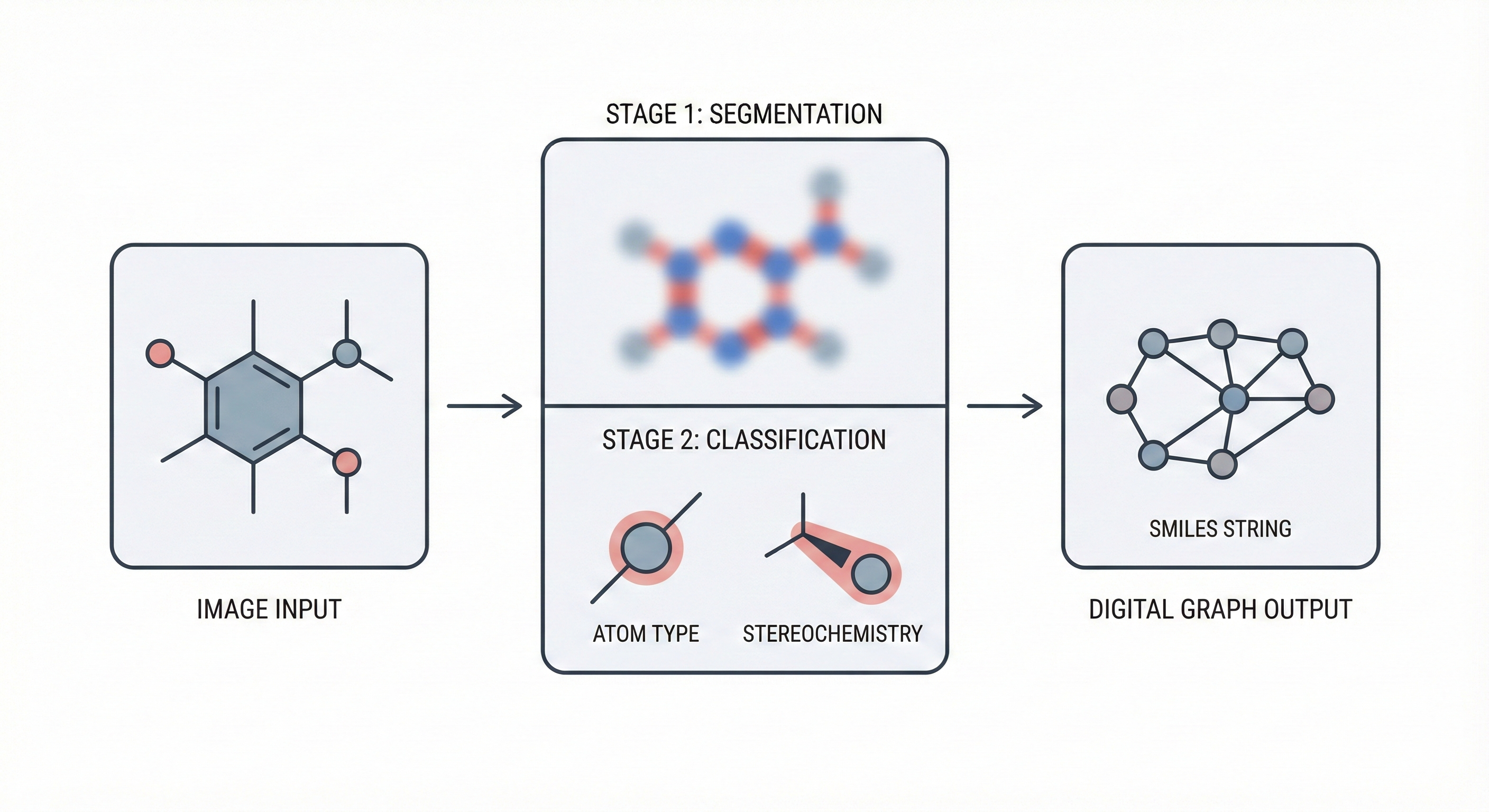
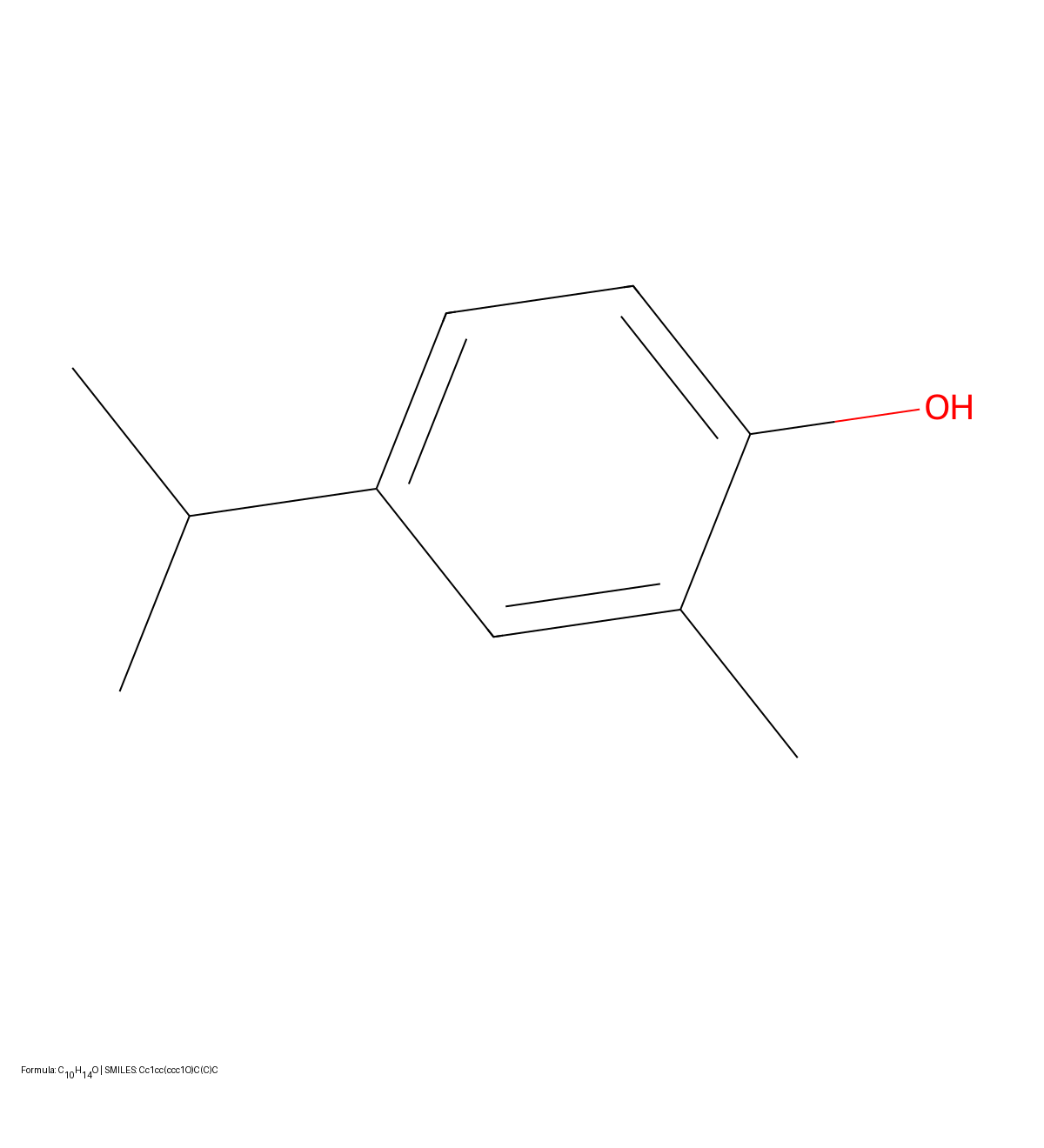
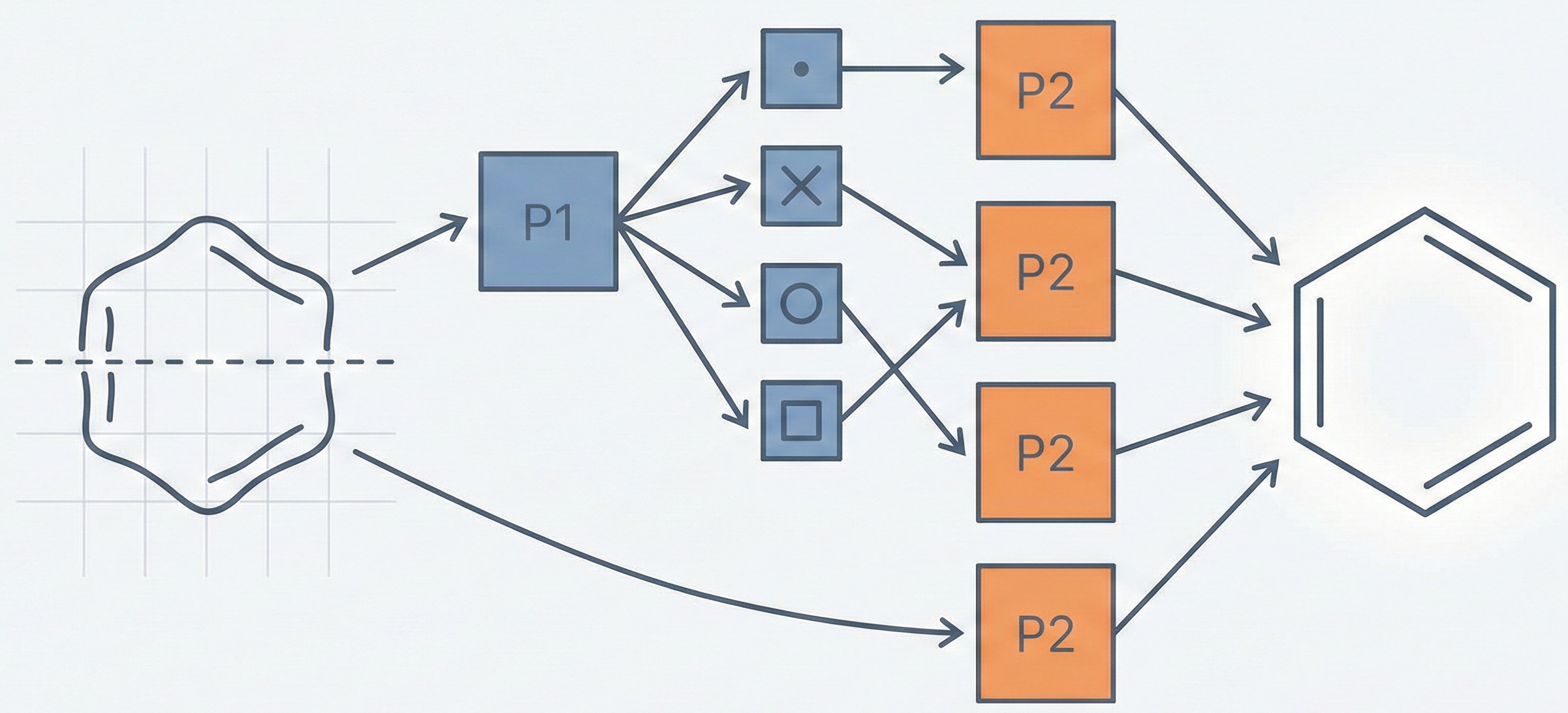
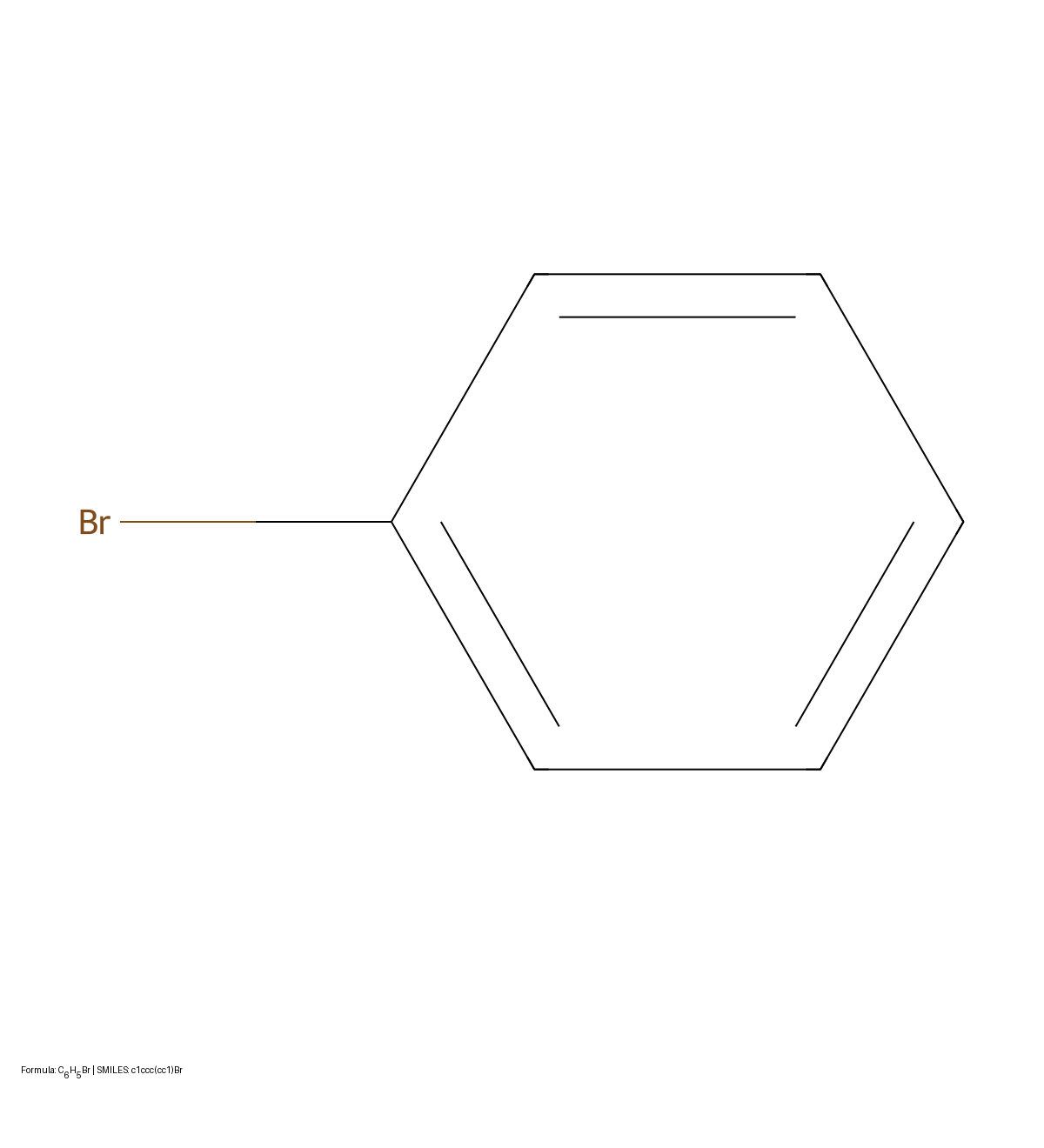
MICER treats optical chemical structure recognition as an image captioning task, using transfer learning with a fine-tuned ResNet encoder and attention-based LSTM decoder to convert molecular images into SMILES strings, reaching 97.54% sequence accuracy on synthetic data and 82.33% on real-world images.