MolGrapher: Graph-based Chemical Structure Recognition
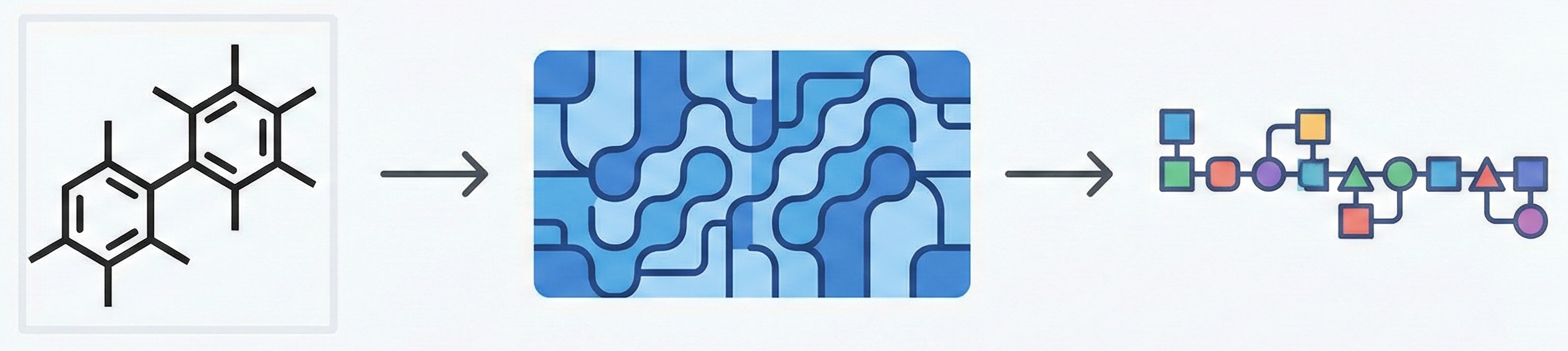
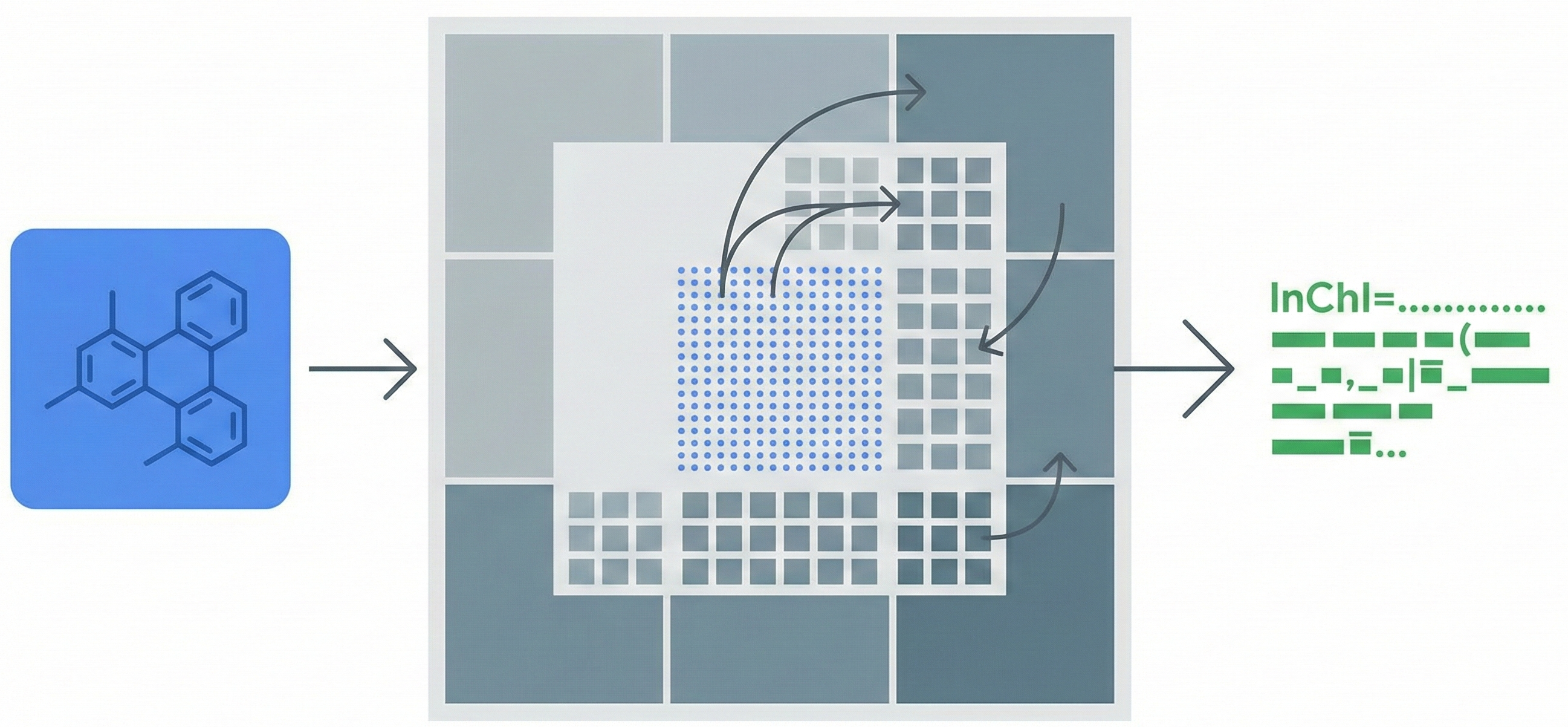
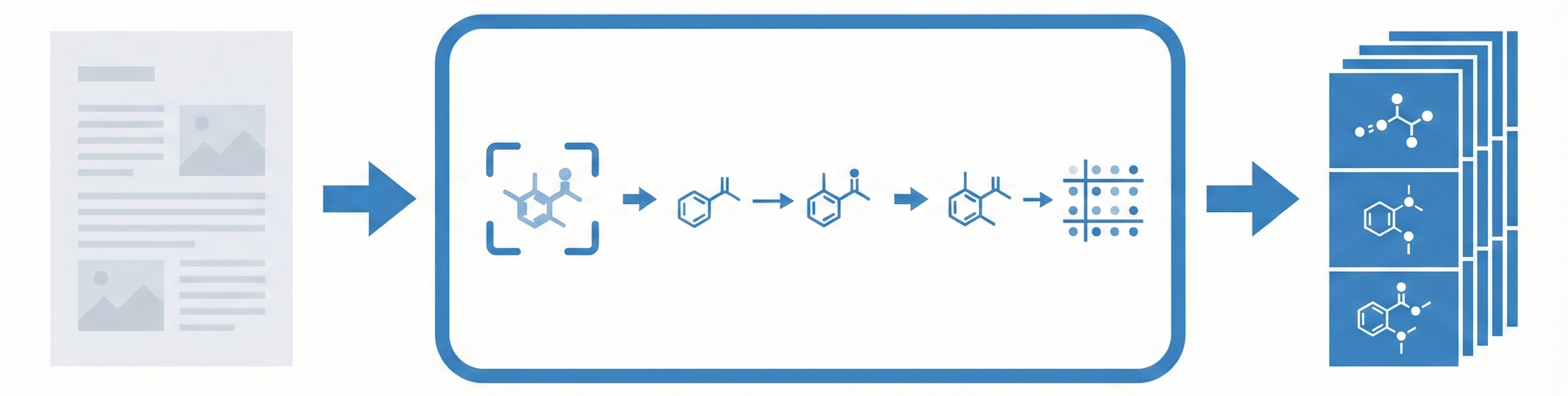
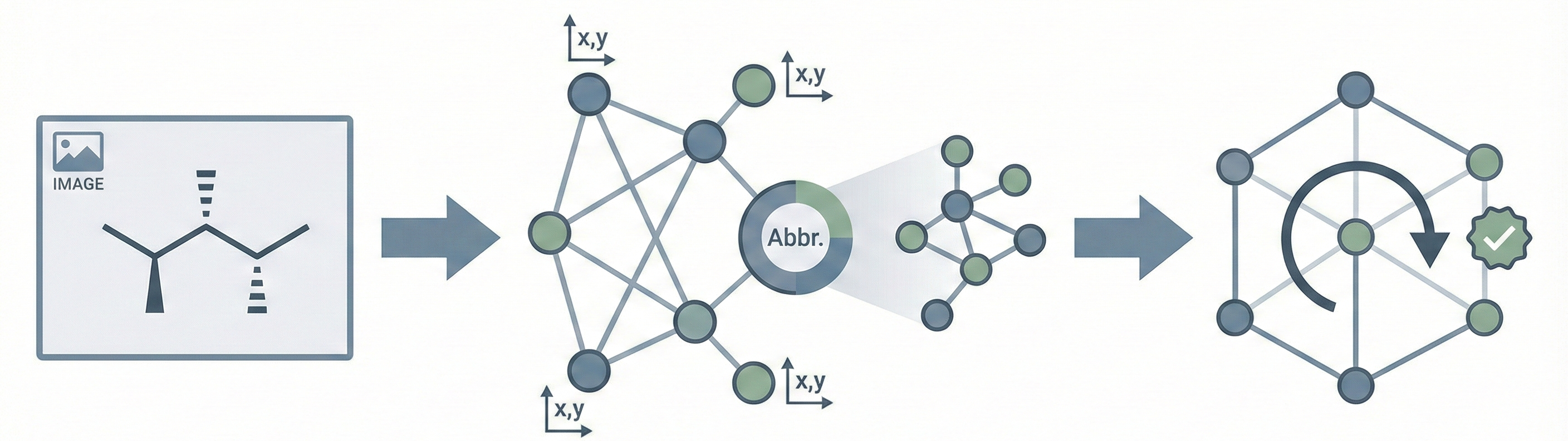
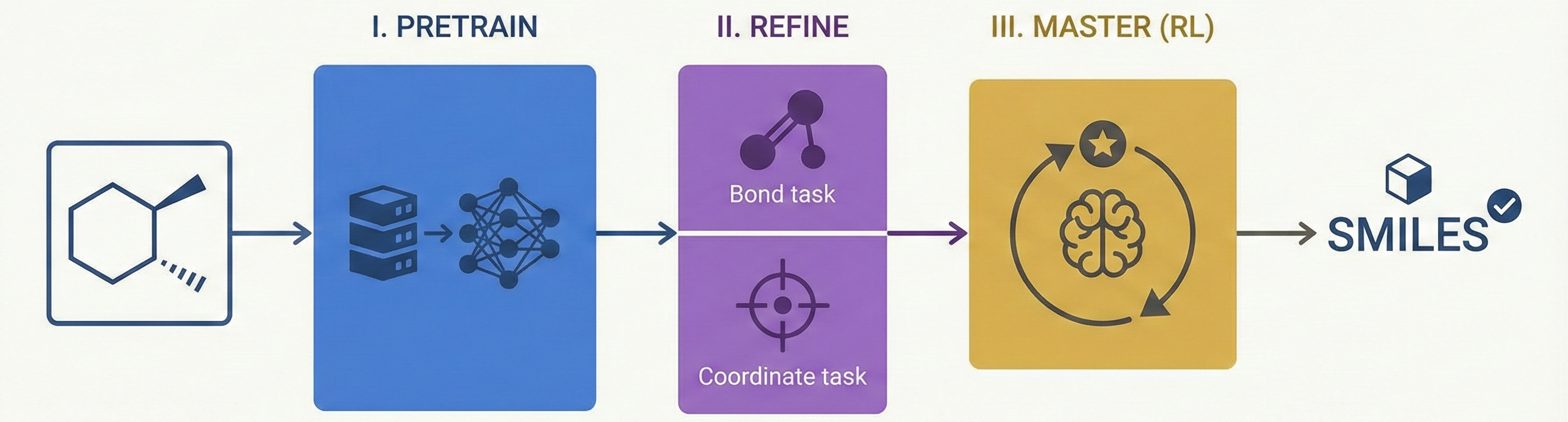
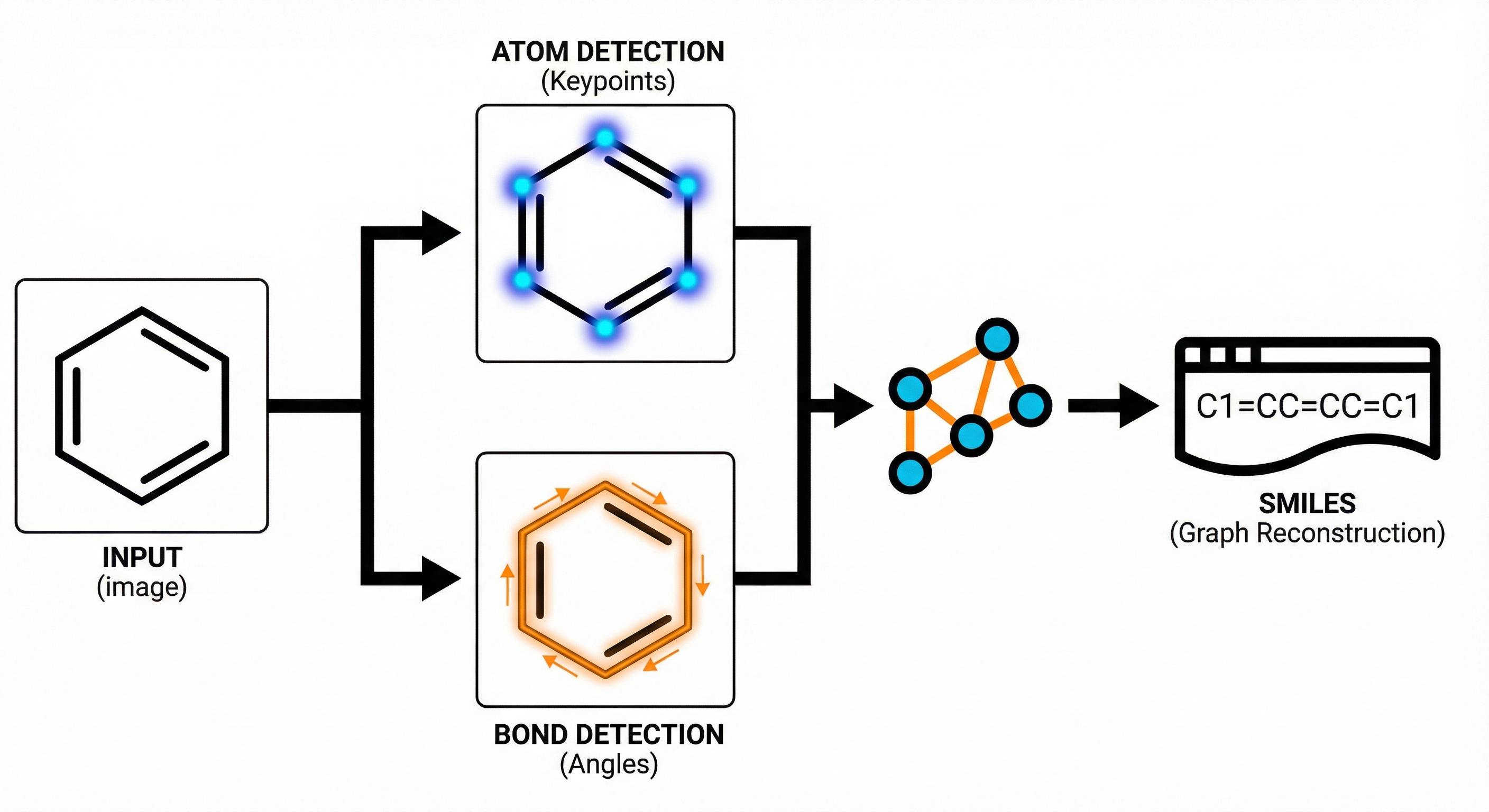
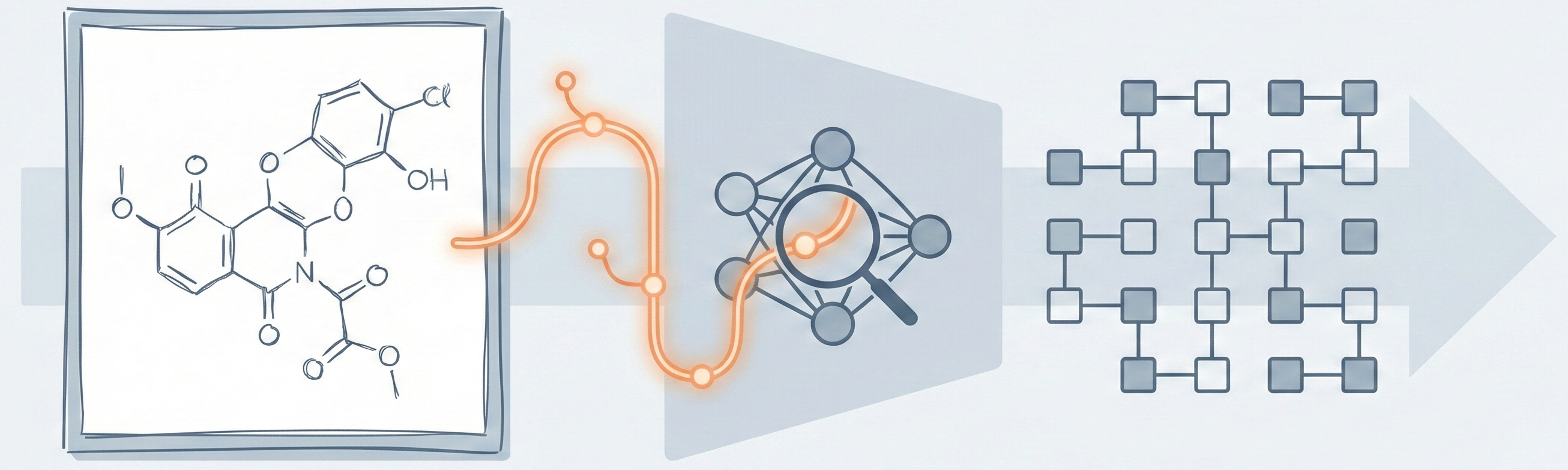
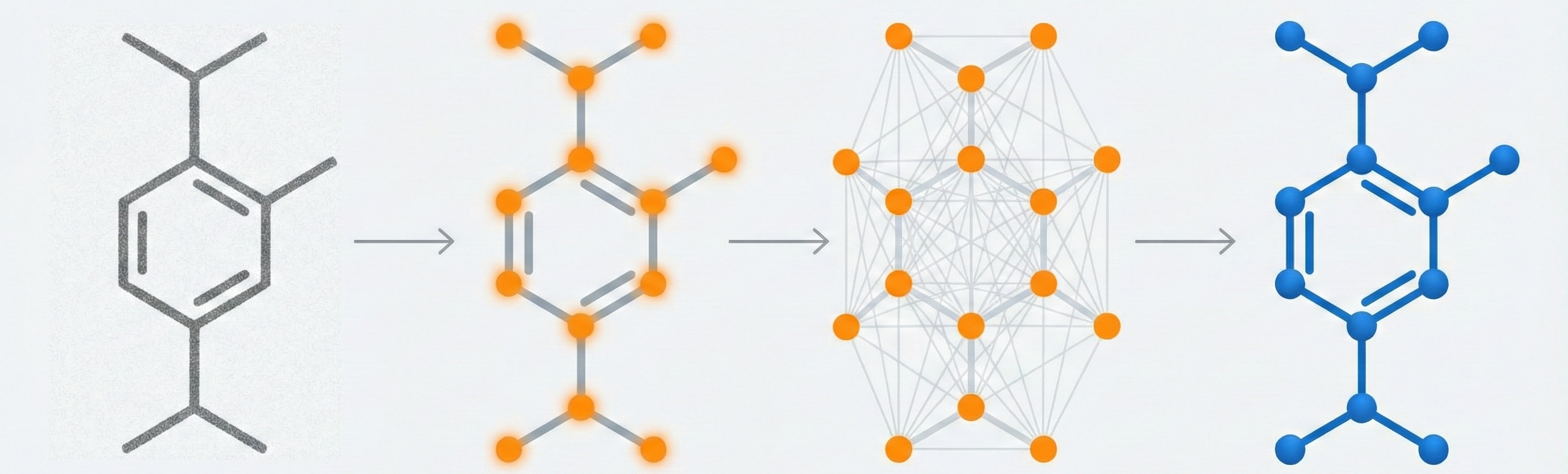
MolGrapher introduces a three-stage pipeline (keypoint detection, supergraph construction, GNN classification) for recognizing chemical structures from images. It achieves 91.5% accuracy on USPTO by treating molecules as graphs, and introduces the USPTO-30K benchmark.