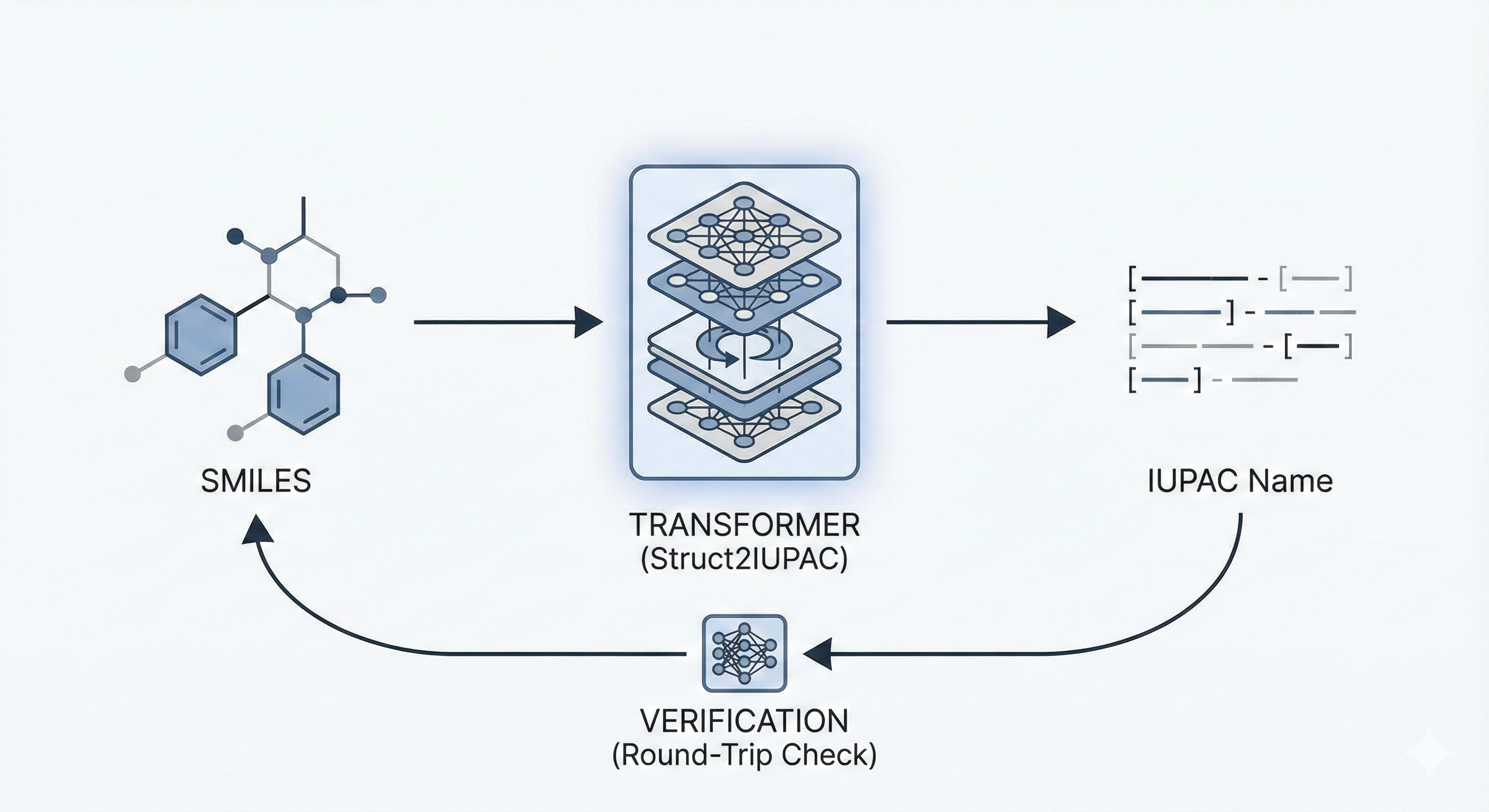
Struct2IUPAC: Translating SMILES to IUPAC via Transformers
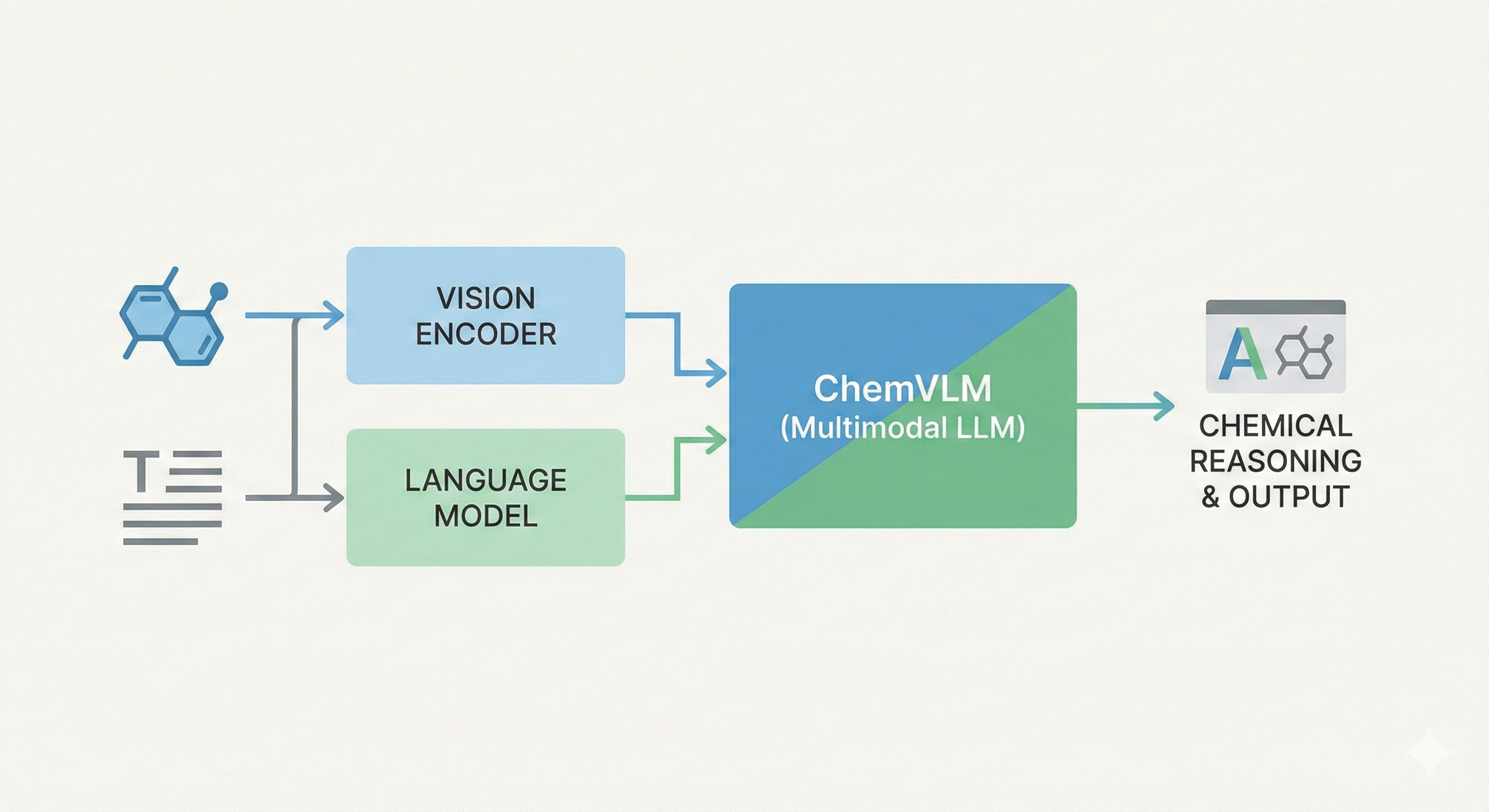
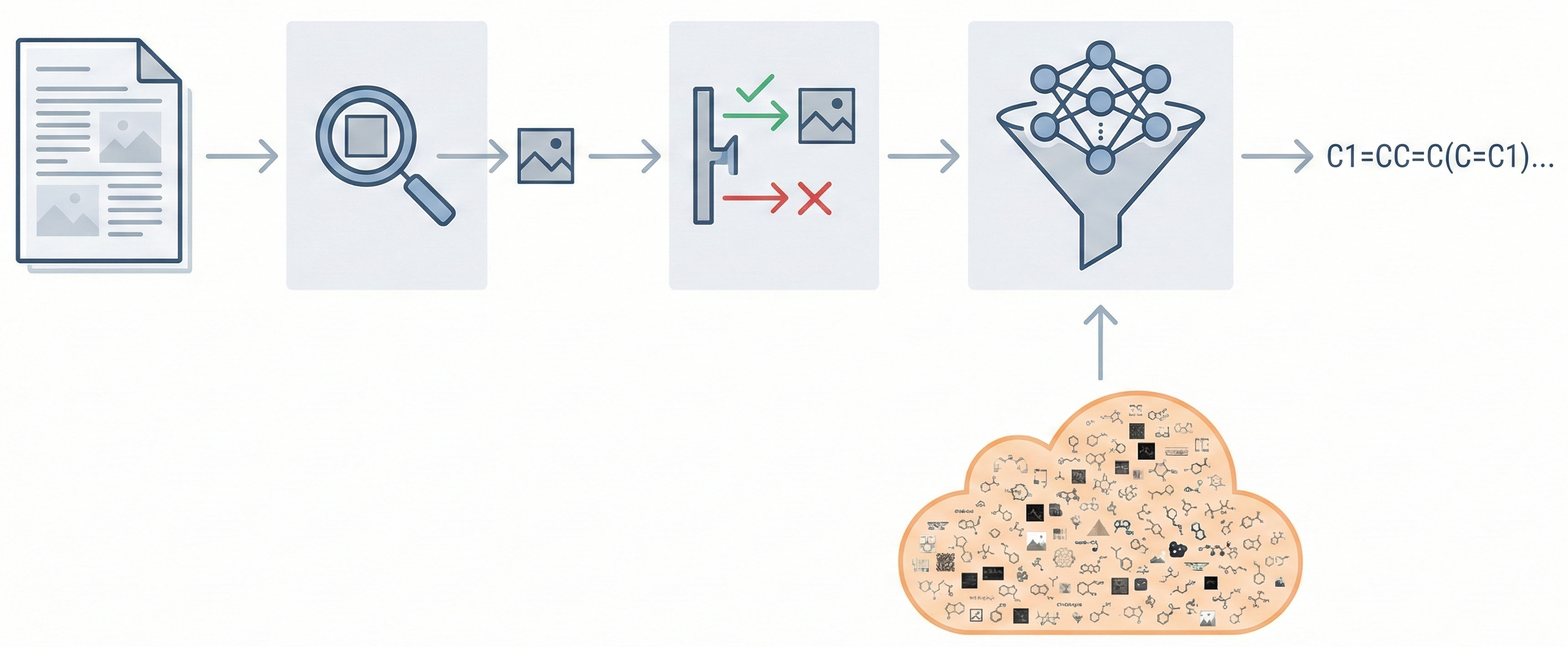
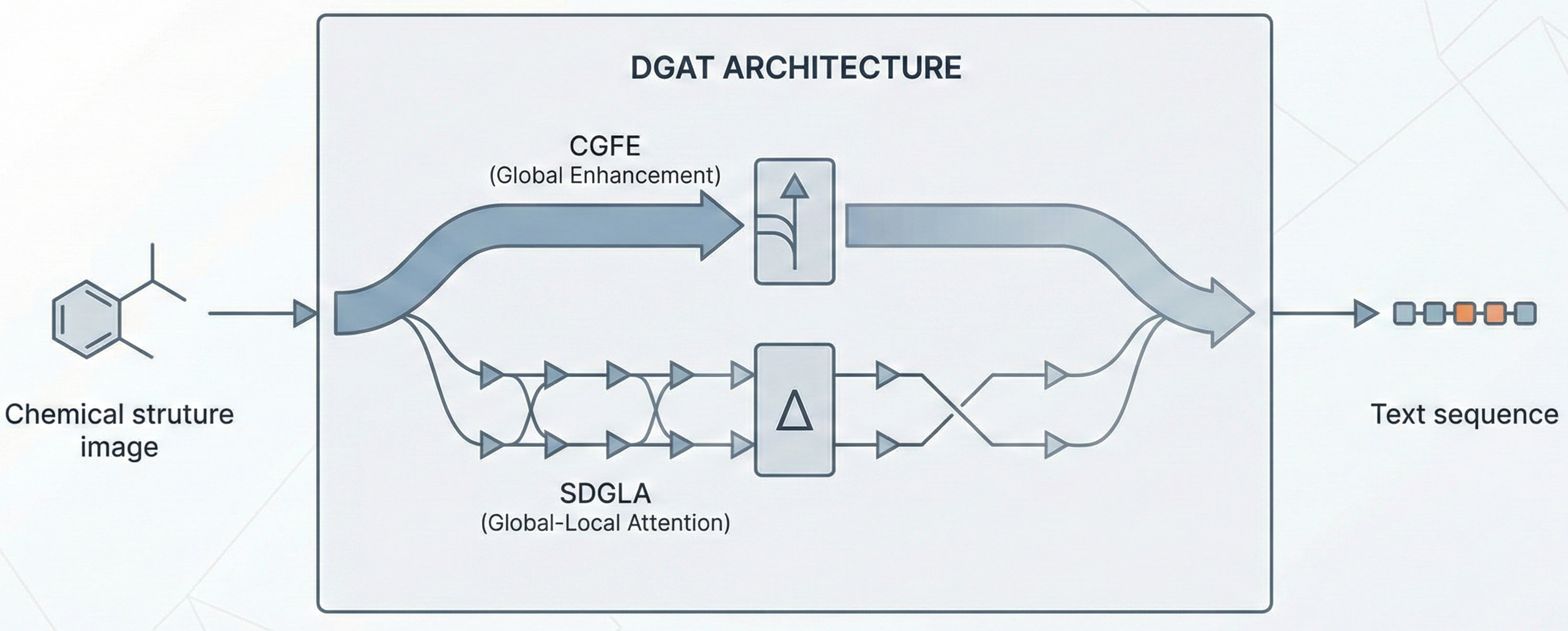
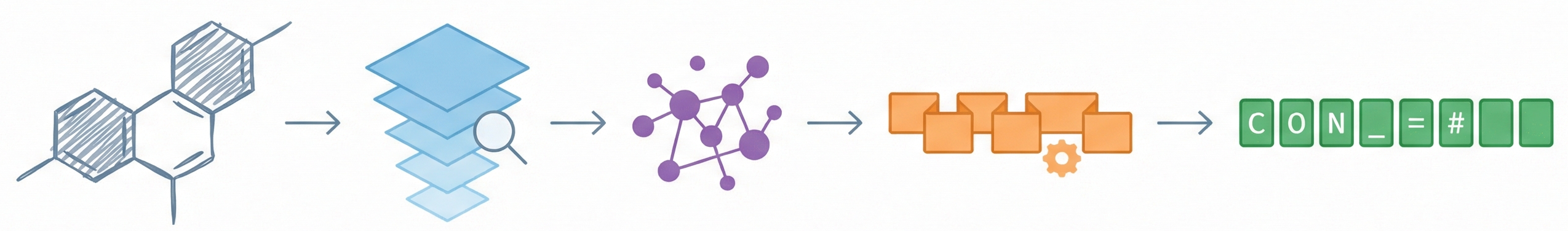
This paper proposes a Transformer-based approach (Struct2IUPAC) to convert chemical structures to IUPAC names, challenging the dominance of rule-based systems. Trained on ~47M PubChem examples, it achieves near-perfect accuracy using a round-trip verification step with OPSIN.